Deep Learning methods for the segmentation of Tumour Spheroids
Project description
Deep-Tumour-Spheroid
This package contains several commands and utilities to easily use Semantic Segmentation models in tumor spheroids detection, specifically Glioblastoma Multiforme Tumors (GBM).
🚀 Getting Started
To start using this package, install it using pip:
For example, for installing it in Ubuntu use:
pip3 install Deep-Tumour-Spheroid
It is recommended to install it globally and not inside virtual environments. Have been tested in Windows, Linux and MacOS.
👩💻 Usage
This package makes easier the use of the best trained model. For that purpose you have available 2 commands:
deep-tumour-spheroid image <inputImagePath> <outputFolder>This method predict over an image. Supported types are:.jpg,.png,.nd2,.tify.tiff.deep-tumour-spheroid folder <inputFolder> <outputFolder>This method predict in all the images of a folder.
You can use deep-tumour-spheroid or it's two abbreviations dts or deep-tumour.
In addition, you can use the GUI developed for preparing the dataset. For that purpose run: deep-tumour-spheroid gui. More information of the utilities in the next section.
You can also execute deep-tumour-spheroid --help, deep-tumour-spheroid gui --help, deep-tumour-spheroid image --help, deep-tumour-spheroid folder --help for a detailed help.
💻 GUI
This GUI contains 4 different utilities: predict, convert ".nd2" and ".tiff" 8 bits unsigned to ".png", transform ".roi" into a ".png" Mask and generating the Dataset.
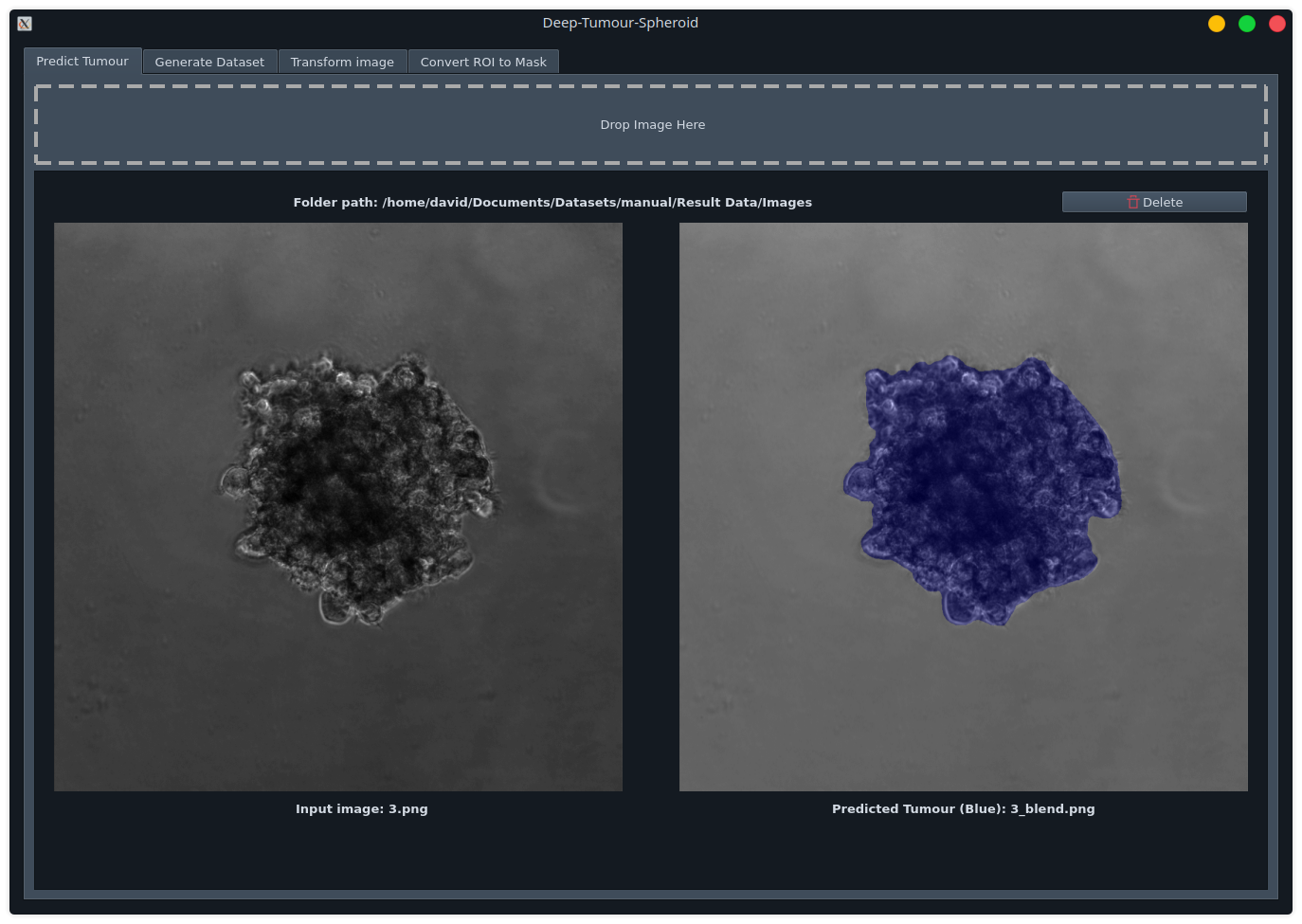
Predict
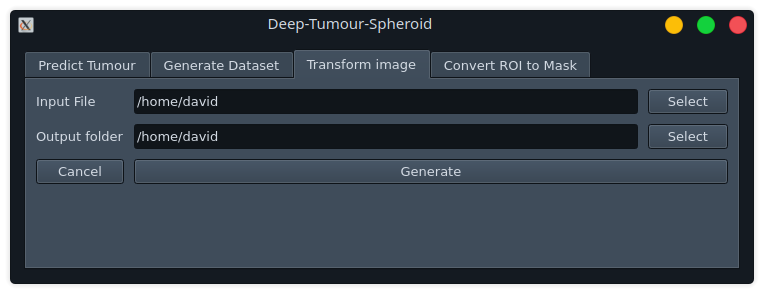
Transform Image
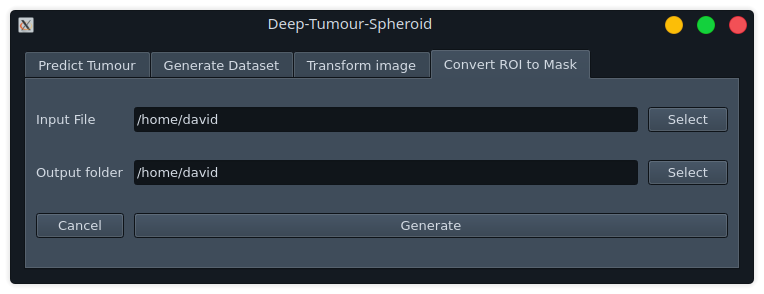
Convert ROI to Mask
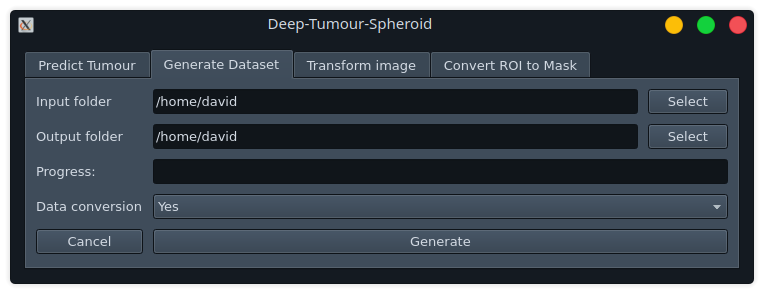
Generate Dataset
📩 Contact
📧 dvdlacallecastillo@gmail.com
💼 Linkedin David Lacalle Castillo
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file Deep-Tumour-Spheroid-1.0.0rc1.tar.gz.
File metadata
- Download URL: Deep-Tumour-Spheroid-1.0.0rc1.tar.gz
- Upload date:
- Size: 26.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.21.0 setuptools/49.6.0 requests-toolbelt/0.9.1 tqdm/4.31.1 CPython/3.7.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
248a152cc7a4fe111bb078fe3cdc9f091e62bb48cae8cc83fa93d84f99609a4b
|
|
| MD5 |
f5f2373e95c7cc1f45ea5a15a538e2b4
|
|
| BLAKE2b-256 |
49f9d9d0d8ccef66c0a0bd46f0638e22a4d440caf756b95167d6cb6db823634a
|