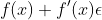Package for Forward/Reverse Autodifferentiation
Project description
Dotua Documentation
Nick Stern, Vincent Viego, Summer Yuan, Zach Wehrwein
Introduction
Calculus, according to the American mathematician Michael Spivak in his noted textbook, is fundamentally the study of "infinitesimal change." An infinitesimal change, according to Johann Bernoulli as Spivak quotes, is so tiny that "if a quantity is increased or decreased by an infinitesimal, then that quantity is neither increased nor decreased." The study of these infinitesimal changes is the study of relationships of change, not the computation of change itself. The derivative is canonically found as function of a limit of a point as it approaches 0 -- we care about knowing the relationship of change, not the computation of change itself.
One incredibly important application of the derivative is varieties of optimization problems. Machines are able to traverse gradients iteratively through calculations of derivatives. However, in machine learning applications, it is possible to have millions of parameters for a given neural net and this would imply a combinatorially onerous number of derivatives to compute analytically. A numerical Newton's method approach (iteratively calculating through guesses of a limit) is likewise not a wise alternative because even for "very small"
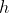
So, one might think that a career in ML thus requires an extensive calculus background, but, Ryan P Adams, formerly of Twitter (and Harvard IACS), now of Princeton CS, describes automatic differentiation as "getting rid of the math that gets in the way of solving a [ML] problem." What we ultimately care about is tuning the hyperparameters of a machine learning algorithm, so if we can get a machine to do this for us, that is ultimately what we care about. What is implemented in this package is automatic differentiation which allows us to calculate derivatives of complex functions to machine precision 'without the math getting in the way.'
One of the most crucial applications of auto-differentiation is backpropagation in neural networks. Backpropagation is the process by which weights are optimized relative to a loss function. These steps are illustrated through a simple neural network example.
Background
The most important calculus derivative rule for automatic differentiation is the multivariate chain rule.
The basic chain rule states that the derivative of a composition of functions is:

That is, the derivative is a function of the incremental change in the outer function applied to the inner function, multiplied by the change in the inner function.
In the multivariate case, we can apply the chain rule as well as the rule of total differentiation. For instance, if we have a simple equation:
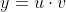
Then,
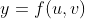
The partial derivatives:
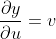
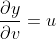
The total variation of y depends on both the variations in u, v and thus,
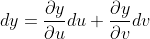
What this trivial example illustrates is that the derivative of a multivariate function is ultimately the addition of the partial derivatives and computations of its component variables. If a machine can compute any given sub-function as well as the partial derivative between any sub-functions, then the machine need only add-up the product of a function and its derivatives to calculate the total derivative.
An intuitive way of understanding automatic differentiation is to think of any complicated function as ultimately a a graph of composite functions. Each node is a primitive operation -- one in which the derivative is readily known -- and the edges on this graph -- the relationship of change between any two variables -- are partial derivatives. The sum of the paths between any two nodes is thus the partial derivative between those two functions (this a graph restatement of the total derivative via the chain rule).
Forward Mode
Forward mode automatic differentiation thus begins at an input to a graph and sums the source paths. The below diagrams (from Christopher Olah's blog) provide an intuition for this process. The relationship between three variables (X, Y, Z) is defined by a number of paths (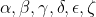
Consequently, provided that within each node there is an elementary function, the machine can track the derivative through the computational graph.
There is one last piece of the puzzle: dual numbers which extend the reals by restating each real as 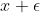
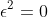
In our chain rule equation, there are two pieces to the computation: the derivative of the outer function applied to the inner function and that value multiplied by the derivative of the inner function. This means that the full symbolic representation of an incredibly complicated function can grow to exponentially many terms. However, dual numbers allow us to parse that symbolic representation in bitesized pieces that can be analytically computed. The reason for this is the Taylor series expansion of a function:
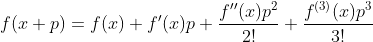
When one evaluates 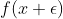
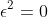
Reverse Mode
One intuition for reverse mode auto differentiation is to consider again our chain-rule-as-graph notion.
One intuitive motivation for this (h/t Rufflewind) is to think of reverse mode as an inversion of the chain-rule.
In this notation, the derivative for some output variable w to some variable t is a linear combination of derivatives for each u_i that w is connected to.
By flipping the numerator and the denominator, this is the notation of the partial derivative of a new parameter s with respect to the input variable u.
The graph theory intuition is perhaps the most straightforward: just as a machine computes an equation in a series of steps, reverse mode is the traversal of that computational graph in reverse. In the context of a neural network in which we wish to minimize weights relative to some loss function, this allows us to efficiently retrace our path without attempting to reevaluate the weights of what could be an incredibly complex network.
To recap: automatic differentiation is an algorithmic means of computing complicated derivatives by parsing those functions as a graph structures to be traversed. Dual numbers are used as a sort of mathematical data structure which allows the machine to analytically compute the derivative at any given node. It is superior to analytic or symbolic differentiation because it is actually computationally feasible on modern machines! And it is superior to numerical methods because automatic differentiation is far more accurate (it achieves machine precision). It is therefore extremely useful for applications like backpropagation on neutral networks.
How to Use Dotua
How to Install
To install our package, one can simply use pip install like so:
$ pip install Dotua
Import and Usage Examples
Forward Mode
In order to instantiate a forward mode auto-differentiation object from our package, the user shall first import the AutoDiff function from the Dotua library as such:
from Dotua.autodiff import AutoDiff as ad
The general workflow for the user is as follows:
- Instantiate all variables as AutoDiff objects.
- Input these variables into operators from the Operator class within the Dotua library to create more complex expressions that propagate the derivative using forward mode automatic differentiation.
The AutoDiff class is the core constructor for all variables in the function that are to be differentiated. There are two options for instantiating variables: Scalar and Vector, generated with create_scalar() and create_vector() respectively. Scalar variables have a single value per variable, while Vector variables can have multiple associated values. The general schematic for how the user shall instantiate AutoDiff objects is outlined below:
- Create either a Scalar or Vector AutoDiff object to generate seed variables to later build the function to be differentiated. The initialization works as follows:
x, y = ad.create_scalar(vals = [1, 2])
z = ad.create_vector(vals = [1, 2, 3])
- Next, the user shall import the Operator class and pass in these variables into elementary functions as follows:
from Dotua.operator import Operator as op
result = op.sin(x * y)
results = op.sin(z)
Simple operators, such as sums and products, can be used normally:
result = 6 * x
results = z + 4
- Finally, (as a continuation of the previous example), the user may access the value and derivative of a function using the eval() and partial() methods:
print(result.eval()) # 6
print(result.partial(x)) # 6
print(result.partial(y)) # 0
For Scalar variables, result.eval() will return the value of the function, while result.partial(v) will return the partial derivative with respect to any variable, v. For Vector variables, results.eval() returns a list of tuples (value, jacobian), with one tuple for each function in the vector. The jacobian is a dictionary that represents the derivative of that element with respect to all of the elements in the vector.
Reverse Mode
The initialization for reverse mode variables is very similar to forward mode. The only difference is that there is an "r" in front of the module names. Additionally, for the initialization of reverse mode variables, the user must instantiate an initializer object. This differs from the forward mode variables which can be initialized using static methods. One can initialize a reverse mode scalar object as follows:
from Dotua.rautodiff import rAutoDiff
rad = rAutoDiff()
x, y, z = rad.create_rscalar([1, 2, 3])
In reverse mode, when the user calls the gradient function, they must specify the variable they would like to differentiate with respect to. This time, the gradient function simply returns a numeric constant. An example of this is shown below:
f = x + y + z
f_gradx = rad.partial(f, x) # f_gradx = 1
The following code shows how the user may interact with rVector. Note that rVector operates differently in reverse mode, as it is mainly an extension to allow one to compute rScalar functions for a vector of values.
v = rad.create_rvector([1, 2 ,3])
g = 2*v
g_grad = rad.partial(g, v) # g_grad = [2, 2, 2]
Examples
There are several files in the top level directory of the Dotua package that demonstrate the usage of the package.
The first file is an interactive jupyter notebook which contains an example use case where the Dotua package performs well, namely, the Newton-Raphson method for approximating roots of functions. This notebook is titled "newton_demo.ipynb" and resides in "examples" folder in the top level directory of the package. The output of this demo is reproduced here for convenience:
A second file is an example of how our reverse mode auto differentiation package can be used to do backpropagation in a neural network. The file is called "neuralnet_demo.ipynb"
For some output y_hat and set of inputs X, the task of a neural network is to find a function which minimizes a loss function, like MSE:
Where N is the number of data points, f_i the value returned by the model and y_i the true value for y at a given observation i. A 'dense' neural network will involve connections between every parameter and these edges each contain a weight value. The task is to find the weights which minimize the distance between y_hat and y. Because each node is connected to every other, this captures non-linearities as certain parameters may be more expressive than others. However, this also carries computational burdens.
Backwards auto-differentiation is particularly useful means of traversing the graph and refitting those weights.
Here we implement a toy neural network that has only one hidden layer. Using R.A. Fischer's well-known Iris dataset -- a collection of measurements of flowers and their respective species -- we predict species based on their floral measurements. As the first scatter plot indicates, there is a clear separation boundary between the setosa plants and the other two species, versicolor and virginica. To capture the boundary between each species would require more complexity, and so our simple neural network can only capture the biggest boundary -- a consequence of our package using reverse auto differentiation to fit the appropriate gradient. This is not a neural networks package, but an auto-differentiation package and this is just an illustration of one very useful application.
Software Organization
Directory Structure
Our project will adhere to the directory structure outlined in the python-packaging documentation. At a high level, the project has the following structure:
Dotua/
__init__.py
autodiff.py
operator.py
rautodiff.py
roperator.py
nodes/
__init__.py
node.py
rscalar.py
rvector.py
scalar.py
vector.py
tests/
__init__.py
test_initializer.py
test_operator.py
test_rautodiff.py
test_roperator.py
test_rscalar.py
test_scalar.py
test_vector.py
docs/
documentation.md
milestone1.md
milestone2.md
examples/
__init__.py
newton_demo.py
neural_network_demo.py
...
LICENSE
MANIFEST.in
README.md
requirements.txt
setup.py
.gitignore
Modules
Dotua/
The Dotua module contains the codes for forward mode implementation and reverse mode implementation.
It contains AutoDiff (autodiff.py), which is the driver of the forward mode autodifferentiation. The driver helps the users with getting access to the Node superclass (node.py) and associated subclasses (i.e., Vector (vector.py) and Scalar (scalar.py)) in the nodes file, and the Operator class (operator.py).
It also contains rAutoDiff (rautodiff.py), which is the driver of the reverse mode autodifferentiation. The driver helps the users with getting access to the rScalar class (rscalar.py) and rVector class (rvector.py) in the nodes file and the rOperator class (roperator.py).
examples/
The Examples module has Python files with documented use cases of the library. Examples include an implementation of Newton’s Method for approximating the roots of a non-linear function and a module which computes local extrema and an implementation of Neural Network for prediction problems.
tests/
The Tests module contains the project’s testing suite and is formatted according to the pytest requirements for automatic test discovery.
Testing
Overview
The majority of the testing in Dotua's test suite consists of unit testing. The aim is to verify the correctness of the application with thorough unit testing of all simple usages of the forward and reverse modes of automatic differentiation. Essentially, this involves validating that our application produces correct calculations (evaluations and derivatives) for all elementary functions. Additionally, a range of more complex unit testing covers advanced scenarios such as functions with multidimensional domains and codomains (for forward mode) as well as functions with inherent complexity generated from the composition of elementary functions.
Test Automation
Dotua uses continuous integration testing through Travis CI to perform automated, machine independent testing. Additionally, Dotua uses Coveralls to validate the high code coverage of our testing suite (currently 100%). Travis CI and Coveralls badges are embedded into the project README to provide transparency for users interacting with our project through GitHub.
Installation
To install our package, one can simply use pip install like so:
$ pip install Dotua
User Verification
The entire Dotua test suite is included in the project distribution. Thus, users are able to verify correctness for themselves using pytest after installing the Dotua package.
Distribution
Licensing
Dotua is distributed under the GNU GPLv3 license to allow free “as is” usage while requiring all extensions to remain open source.
Implementation
The purpose of the Dotua library is to perform automatic differentation on user defined functions, where the domain and codomain may be single- or multi-dimensional (n.b. this library provides support for both the forward and reverse modes of automatic differentation, but for the reverse mode only functions with single-dimensional codomains are supported). At a high level, Dotua serves as a partial replacement for NumPy in the sense that Dotua provides methods for many of the mathematical functions (e.g., trigonometric, inverse trigonometric, hyperbolic, etc.) that NumPy implements; however, while the NumPy versions of these methods can only provide function evaluation, the Dotua equivalents provide both evaluation and differentiation.
To achieve this, the Dotua library implements the following abstract ideas:
- Allowing users to be as expressive as they would like to be by providing our own versions of binary and unary operators.
- Forward AD: keeping track of the value and derivative of user defined expressions and functions.
- Reverse AD: constructing a computational graph from user defined functions that can be used to quickly compute gradients.
With these goals in mind, the Dotua forward mode implementation relies on the Nodes modules and the Operator class and allows user interface through the AutoDiff class which serves as a Node factory for initializing instances of Scalar and Vector. Analogously, the Dotua reverse mode implementation relies on the Nodes module and the rOperator class and facilitates user interface through the rAutoDiff class which serves as a factory for initializing instances of rScalar.
Nodes
The Nodes module contains a Node superclass with the following basic design:
class Node():
def eval(self):
'''
For the Scalar and Vector subclasses, this function returns the node's
value as well as its derivative, both of which are guaranteed to be
up to date by the class' operator overloads.
Returns (self._val, self._jacobian)
'''
raise NotImplementedError
def __add__(self, other):
raise NotImplementedError
def __radd__(self, other):
raise NotImplementedError
def __mul__(self, other):
raise NotImplementedError
def __rmul__(self, other):
raise NotImplementedError
... # Additional operator overloads
Essentially, the role of the Node class (which in abstract terms is meant to represent a node in the computational graph underlying forward mode automatic differentiation of user defined expressions) is to serve as an interface for two other classes in the Nodes package: Scalar, Vector, rScalar, rVector. Each of these subclasses implements the required operator overloading as necessary for scalar and vector functions respectively (i.e., addition, multiplication, subtraction, division, power, etc.). This logic is separated into four separate classes to provide increased organization for higher dimensional functions and to allow class methods to use assumptions of specific properties of scalars and vectors to reduce implementation complexity.
Both the Scalar class and the Vector class have _val and _jacobian class attributes which allow for forward automatic differentiation by keeping track of each node's value and derivative. Both the rScalar class and the rVector class have _roots and _grad_val class attributes which allow for reverse auto differentiation by storing the computational graph and intermediate gradient value.
Scalar
The Scalar class is used for user defined one-dimensional variables. Specifically, users can define functions of scalar variables (i.e., functions defined over multiple scalar variables with a one-dimensional codomain) using instances of Scalar in order to simultaneously calculate the function value and first derivative at a pre-chosen point of evaluation using forward mode automatic differentiation. Objects of the Scalar class are initialized with a value (i.e., the point of evaluation) which is stored in the class attribute self._val (n.b., as the single underscore suggests, this attribute should not be directly accessed or modified by the user). Additionally, Scalar objects – which could be either individual scalar variables or expressions of scalar variables – keep track of their own derivatives in the class attribute self._jacobian. This derivative is implemented as a dictionary with Scalar objects serving as the keys and real numbers as values. Note that each Scalar object's _jacobian attribute has an entry for all scalar variables which the object might interact with (see AutoDiff Initializer section for more information).
Users interact with Scalar objects in two ways:
- eval(self): This method allows users to obtain the value for a Scalar object at the point of evaluation defined when the user first initialized their Scalar objects. Specifically, this method returns the float self._val.
- partial(self, var): This method allows users to obtain a partial derivative of the given Scalar object with respect to var. If self is one of the Scalar objects directly initialized by the user (see AutoDiff Initializer section), then partial() returns 1 if var == self and 0 otherwise. If self is a Scalar object formed by an expression of other Scalar objects (e.g., self = Scalar(1) + Scalar(2)), then this method returns the correct partial derivative of self with respect to var.
Note that these are the only methods that users should be calling for Scalar objects and that users should not be directly accessing any of the object's class attributes.
Scalar objects support left- and right-sided addition, subtraction, multiplication, division, exponentiation, and negation.
rScalar
The rScalar class is the reverse mode automatic differentiation analog of the Scalar class. That is to say, rScalar is used for user defined one-dimensional variables with which users can define scalar functions (i.e., functions defined over multiple scalar variables with a one-dimensional codomain) in order to calculate the function value and later easily determine the function's gradient reverse mode automatic differentiation (see the rAutoDiff Initializer section for more details). Objects of the rScalar class are initialized with a value (i.e., the point of evaluation) which is stored in the class attribute self._val (n.b., as the single underscore suggests, this attribute should not be directly accessed or modified by the user). Additionally, rScalar objects – which could be either individual scalar variables or expressions of scalar variables – explicitly construct the computational graph used in automatic differentiation in the class attribute self.roots(). This attribute represents the computational graph as a dictionary where keys represent the children and the values represent the derivatives of the children with respect to rScalar self. This dictionary is constructed explicitly through operator overloading whenever the user defines functions using rScalar objects.
Users interact with rScalar objects in one way:
- eval(self): This method allows users to obtain the value for an rScalar object at the point of evaluation defined when the user first initialized the rScalar object. Specifically, this method returns the value of the attribute self._val.
Note that this is the only method that users should be calling for rScalar objects and that users should not be directly accessing any of the object's class attributes. While the rScalar class contains a gradient() method, this method is for internal use only. Users should only be obtaining the derivatives of functions represented by rScalar objects through the partial() method provided in the rAutoDiff initializer class (see rAutoDiff Initializer section for more details).
rScalar objects support left- and right-sided addition, subtraction, multiplication, division, exponentiation, and negation.
Vector
Vector is a subclass of Node. Every vector variable consists of a 1-d numpy array to store the values and a 2-d numpy array to store the jacobian matrix. User can use index to acess specific element in a Vector instance. And operations between elements in the same vector instance and operations between vectors are implemented by overloading the operators of the class.
AutoDiff Initializer
The AutoDiff class functions as a Node factory, allowing the user to initialize variables for the sake of constructing arbitrary functions. Because the Node class serves only as an interface for the Scalar and Vector classes, users should not instantiate objects of the Node class directly. Thus, we define the AutoDiff class in the following way to allow users to initialize Scalar and Vector variables:
from Dotua.nodes.scalar import Scalar
from Dotua.nodes.vector import Vector
class AutoDiff():
@staticmethod
def create_scalar(vals):
'''
@vals denotes the evaluation points of variables for which the user
would like to create Scalar variables. If @vals is a list,
the function returns a list of Scalar variables with @vals
values. If @vals is a single value, the user receives a single Scalar
variable (not as a list). This function also initializes the jacobians
of all variables allocated.
'''
pass
@staticmethod
def create_vector(vals):
'''
The idea is similar to create_scalar.
This will allow the user to create vectors and specify initial
values for the elements of the vectors.
'''
pass
Using the create_scalar and create_vector methods, users are able to initialize variables for use in constructing arbitrary functions. Additionally, users are able to specify initial values for these variables. Creating variables in this way will ensure that users are able to use the Dotua defined operators to both evaluate functions and compute their derivatives.
Variable Universes
The implementaiton of the AutoDiff library makes the following assumption: for each environment in which the user uses autodifferentiable variables (i.e., Scalar and Vector objects), the user initializes all such variables with a single call to create_scalar or create_vector. This assumption allows Scalar and Vector objects to fully initialize their jacobians before being used by the user. This greatly reduces implementation complexity.
This design choice should not restrict users in their construction of arbitrary functions for the reason that in order to define a function, the user must know how many primitive scalar variables they need to use in advance. Realize that this does not mean that a user is prevented from defining new Python variables as functions of previously created Scalar objects, but only that a user, in defining a mathematical function f(x, y, z) must initialize x, y, and z with a single call to create_scalar. It is perfectly acceptable that in the definition of f(x, y, z) a Python variable such as a = x + y is created. The user is guaranteed that a.eval() and a.partial(x), a.partial(y), and a.partial(z) are all well defined and correct because a in this case is an instance of Scalar; however, it is not a "primitive" scalar variable and thus the user could not take a partial derivative with respect to a.
rAutoDiff Initializer
The rAutoDiff class functions as an rScalar factory, allowing the user to initialize variables for the sake of constructing arbitrary functions of which they want to later determine the derivative using reverse mode automatic differentiation. Because the same rScalar variables can be used to define multiple functions, users must instantiate an rAutoDiff object to manage the rScalar objects they create and calcuate the gradients of different functions of the same variables. Thus, we define the rAutoDiff class in the following ways:
from Dotua.nodes.rscalar import rScalar
class rAutoDiff():
def __init__(self):
self.func = None
def create_rscalar(vals):
'''
@vals denotes the evaluation points of variables for which the user
would like to create rScalar variables. If @vals is a list,
the function returns a list of rScalar variables with @vals
values. If @vals is a single value, the user receives a single rScalar
variable (not as a list). This function also adds the new rScalar
object(s) to the _universe of the rAutoDiff object.
'''
pass
def partial(self, func, var):
'''
This method allows users to calculate the derivative of @func the
function of rScalar objects with respect to the variable represented
by the rScalar @var. This method also sets the self.func attribute
of the rAutoDiff object to the given @func.
'''
pass
def _reset_universe(self, func, var):
'''
This method is for internal use only. When a user calls partial(),
the rAutoDiff object will first call _reset_universe() to reset then
grad_val variables of the necessary rScalar objects before
calculating the desired derivative.
'''
pass
By instantiating an rAutoDiff object and using the create_rscalar method, users are able to initialize variables for use in constructing arbitrary functions. Additionally, users are able to specify initial values for these variables. Creating variables in this way will ensure that users are able to use the Dotua defined operators to both evaluate functions and compute their derivatives. Furthermore, using the partial method, users are able to determine the derivative of their constructed function with respect to a specified rScalar variable.
Operator
The Operator class defines static methods for elementary mathematical functions and operators (specifically those that cannot be overloaded in the Scalar and Vector classes) that can be called by users in constructing arbitrary functions. The Operator class will import the Nodes module in order to return new Scalar or Vector variables as appropriate. The design of the Operator class is as follows:
import numpy as np
from Dotua.nodes.scalar import Scalar
from Dotua.nodes.vector import Vector
class Operator():
@staticmethod
def sin(x):
pass
@staticmethod
def cos(x):
pass
... # Other elementary functions
For each method defined in the Operator class, our implementation uses ducktyping to return the necessary object. If user passes a Scalar object to one of the methods then a new Scalar object is returned to the user with the correct value and jacobian. If user passes a Vector object to one of the methods then a new Vector object is returned to the user with the correct value and jacobian. On the other hand, if the user passes a Python numeric type, then the method returns the evaluation of the corresponding NumPy method on the given argument (e.g., op.sin(1) = np.sin(1)).
rOperator
Similarly, the rOperator class defines static methods for elementary mathematical functions and operators (specifically those that cannot be overloaded in the rScalar class) that can be called by users in constructing arbitrary functions. The design of the rOperator class is as follows:
import numpy as np
from Dotua.nodes.rscalar import rScalar
from Dotua.nodes.rvector import rVector
class rOperator():
@staticmethod
def sin(x):
pass
@staticmethod
def cos(x):
pass
... # Other elementary functions
Once again, for each method defined in the rOperator class, our implementation uses ducktyping to return the necessary object. If user passes an rScalar object to one of the methods, then a new rScalar object is returned to the user with the correct value and self/child link. If user passes an rVector object to one of the methods, then a new rVector object is returned to the user with the correct value and parent/child links. On the other hand, if the user passes a Python numeric type, then the method returns the evaluation of the corresponding NumPy method on the given argument (e.g., rop.sin(1) = np.sin(1)).
A Note on Comparisons
It is important to note that the Dotua library intentionally does not overload comparison operators for its variables class (i.e., Scalar, rScalar, Vector, and rVector). Users should only use the the equality and inequality operators == and != to determine object equivalence. For users wishing to perform comparisons on the values of functions or variables composed of Scalar, rScalar, Vector, or rVector variables with the values of functions or variables of the same type, they can do so by accessing the values with the eval() function.
External Depencies
Dotua restricts dependencies on third-party libraries to the necessary minimum. Thus, the only external dependencies are NumPy, and SciPy NumPy is used as necessary within the library for mathematical computation (e.g., trigonometric functions). SciPy is used within the Newton-Raphson Demo as a comparison.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file Dotua-1.0.0.tar.gz.
File metadata
- Download URL: Dotua-1.0.0.tar.gz
- Upload date:
- Size: 43.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.11.0 pkginfo/1.4.2 requests/2.18.4 setuptools/40.6.2 requests-toolbelt/0.8.0 tqdm/4.28.1 CPython/3.6.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1eb5f4f1ef6ab3981931730dccf2f89aa9864c0653fbbac6fe5b773a0b22efc0
|
|
| MD5 |
b7ff504ef14bad86f5894918c61caa68
|
|
| BLAKE2b-256 |
f8993d0a65a496a91eef5f609ea3d7c63f9d900908d36f03f445a4ff19aba3ef
|
File details
Details for the file Dotua-1.0.0-py3-none-any.whl.
File metadata
- Download URL: Dotua-1.0.0-py3-none-any.whl
- Upload date:
- Size: 58.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.11.0 pkginfo/1.4.2 requests/2.18.4 setuptools/40.6.2 requests-toolbelt/0.8.0 tqdm/4.28.1 CPython/3.6.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1cf60b31ee45892c26f460938ee4cea80d842932edebee096c2626fd8fd7dda6
|
|
| MD5 |
d6240bcef99d91a0cbf8334014f00eb9
|
|
| BLAKE2b-256 |
94161a7ae36a1c894c02e12ddf780fe036abc47420a4878a6ef66ede4668c624
|