A package for generating synthetic images of bacteria in phase contrast or fluorescence. Used for creating training data for machine learning segmentation and tracking algorithms.
Project description
SyMBac: Synthetic Micrographs of Bacteria
Update 2025 Please see the SyMBac_2 repo. There is currently a big update coming which will feature lineage tracking ability, full texture generation within cells, spatial fluorescence (membrane, nucleoid, ribosome, etc), and more advanced image simulation models, and BENDY cells! It will also (hopefully) bring about full modularity of SyMBac so you can plug in individual parts of the pipeline to your own workflows.
Read the paper: Synthetic Micrographs of Bacteria (SyMBac) allows accurate segmentation of bacterial cells using deep neural networks , Georgeos Hardo, Maximillian Noka, Somenath Bakshi
[New! Try the Colab version] (please let me know if you run into issues)
- What is it?
- Why would I want to generate synthetic images?
- How do I use these synthetic images?
- Prerequisites
- Installation
- Usage
- FAQs
What is it?
SyMBac is a tool to generate synthetic phase contrast or fluorescence images of bacteria. Currently the tool only supports bacteria growing in the mother machine, however support for bacteria growing in monolayers (and maybe even biofilms!) is coming.
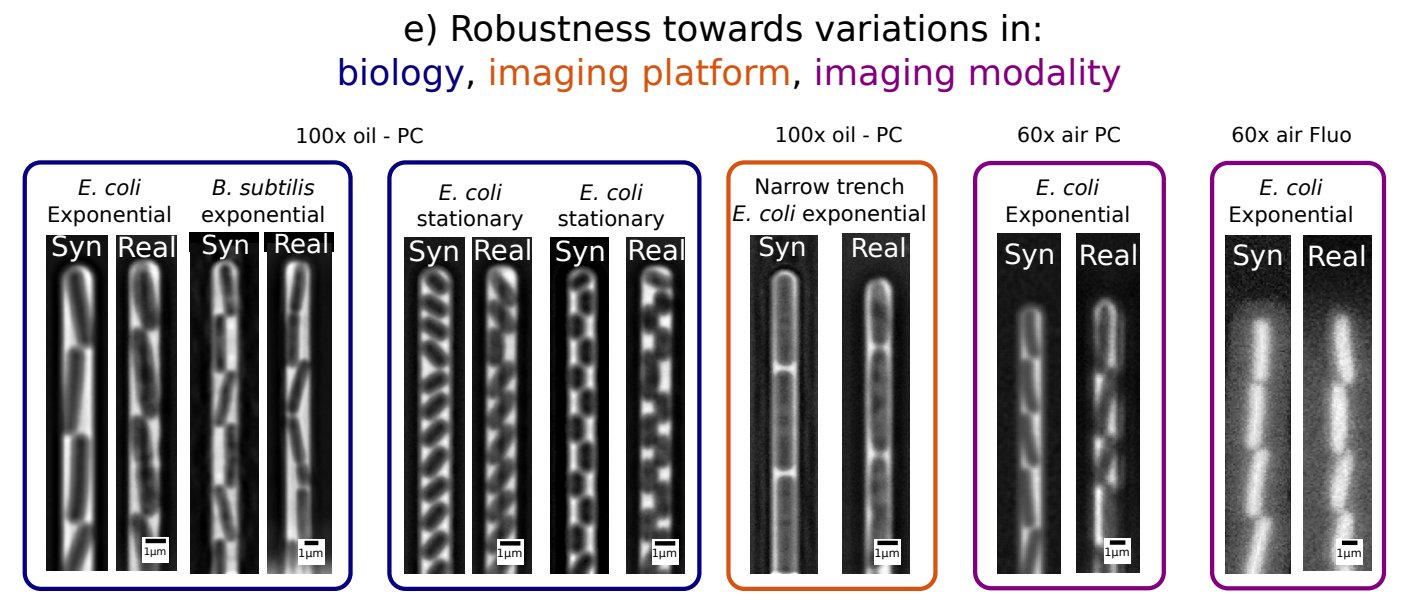
Why would I want to generate synthetic images?
Because you're sick of generating your own training data by hand! Synthetic images provide an instant source of high quality and unlimited training data for machine learning image segmentation algorithms!
The images are tuned to perfectly replicate your experimental setup, no matter what your microscope's objective is (we have tested 20x air all the way to 100x oil), no matter your imaging modality (phase contrast/fluorescence), and no matter the geometry of your microfluidic device.
Additionally,
- SyMBac is very fast compared to humans:
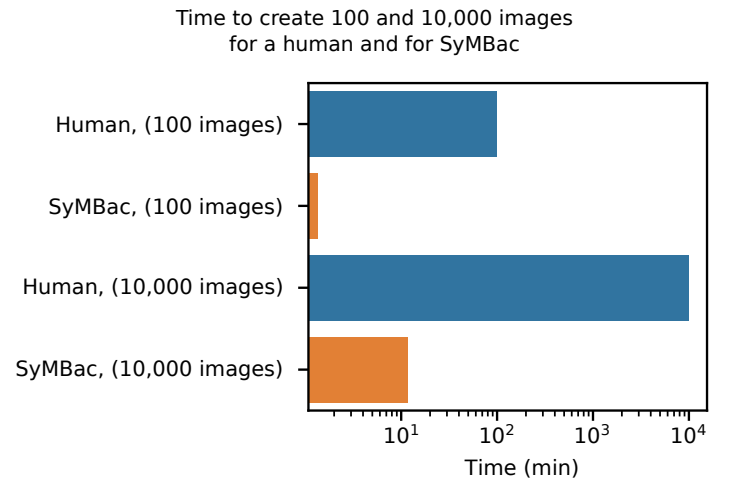

- The image generation process uses a rigid body physics model to simulate bacterial growth, 3D cell geometry to calculate the light's optical path, and a model of the phase contrast/fluorescence optics (point spread function), with some post-rendering optimisation to match image similarity:
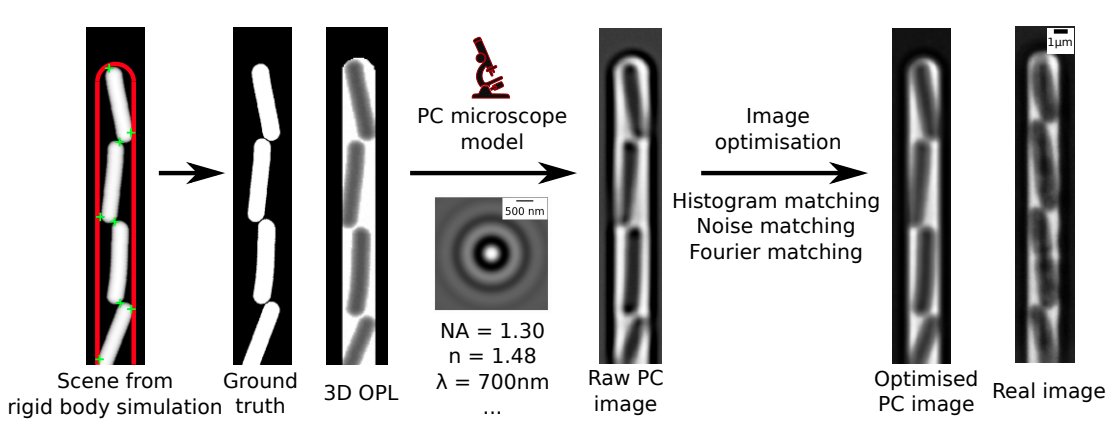
How do I use these synthetic images?
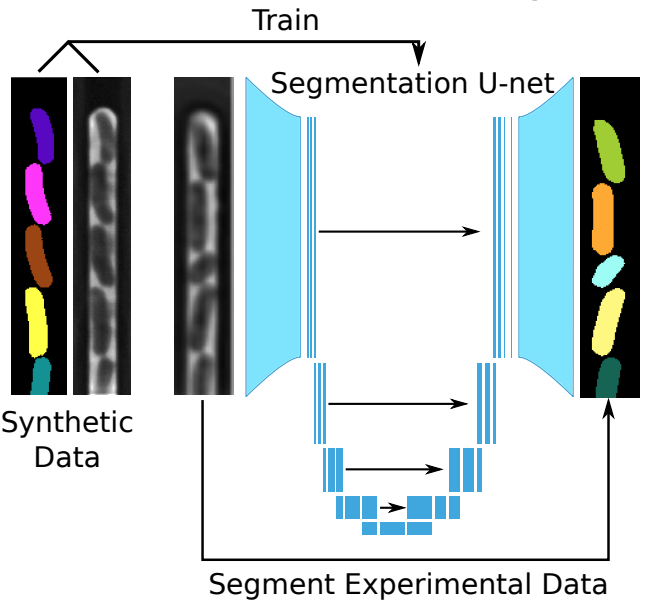
That is up to you. SyMBac is not a machine learning tool. It is a tool to generate unlimited free training data which accurately represents your experiment. It is up to you to train a machine learning network on these synthetic images. We do however provide example notebooks for how to train a U-net (as implemented by DeLTA).
Prerequisites
SyMBac is meant to be run interactively (in a notebook + with a small Qt/GTK interface), so make sure that you are running this on a local machine (you should have access to the machine's display).
A GPU is strongly recommended for convolution performance but is not required — SyMBac will fall back to CPU FFT convolution if no GPU backend is available.
Installation
Base install (CPU only, no GPU acceleration)
pip install SyMBac
This works on any machine. Convolution will use scipy's FFT backend, which is significantly slower than GPU-accelerated convolution.
NVIDIA GPU (recommended)
Install SyMBac with CuPy for CUDA-accelerated convolution:
pip install "SyMBac[cupy]"
This requires a working NVIDIA driver and CUDA 12.x installation. Check with nvcc --version or nvidia-smi.
Apple Silicon (MPS)
Install SyMBac with PyTorch for Metal Performance Shaders acceleration:
pip install "SyMBac[torch]"
PyTorch will use the MPS backend on Apple Silicon Macs automatically.
Development version
pip install "git+https://github.com/georgeoshardo/SyMBac"
Jupyter widgets
Activate the Jupyter widgets extension. This is needed to interact with sliders in the notebooks to optimise images.
jupyter nbextension enable --py widgetsnbextension
Usage
FAQs
- Do I need to have a GPU?
- No, although image synthesis will be around 40x slower on the CPU. SyMBac will detect that you do not have CuPy installed and default to using CPU convolution.
- Interactive image optimisation will be very painful on the CPU. By default I turn off slider interactivity if you are using the CPU, so that you can move a slider without the CPU being maxed out. This means that every time you move a slider you must click the button to update the image (do a convolution).
- Can I generate fluorescence images as well?
- Yes, you can do fluorescence image generation, just make sure that in the interactive image generation part of the code, you select fluorescence.
- Since our fluorescence kernel is defined to be a subset of the phase contrast kernel, you can choose any condenser, and your fluorescence kernel should be correct. Just ensure that the imaging wavelength, numerical aperture, refractive index, and pixel size are set correctly.
- What format do my images need to be in?
- The real images you are trying to replicate should be in the format of single-trench timeseries images. If you are unsure what this is, you can call
get_sample_images()["E. coli 100x"]fromSyMBac.miscfor an example image.
- The real images you are trying to replicate should be in the format of single-trench timeseries images. If you are unsure what this is, you can call
- I'm getting libGL MESA-LOADER errors
- I'm getting
ContextException: Could not create GL contextwhen trying to visualise my simulation in real time- Try
conda install -c conda-forge libstdcxx-ng, and if you're on Ubuntu/Debiansudo apt install libgl1-mesa-dri
- Try
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file symbac-0.5.2.tar.gz.
File metadata
- Download URL: symbac-0.5.2.tar.gz
- Upload date:
- Size: 206.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
16e44bcba249a6e95160e27f8379d244b2fd981c07249530790c114dba9e6925
|
|
| MD5 |
c45049c8e290ed09e6a23e8008f84ac1
|
|
| BLAKE2b-256 |
c148df82b3f3c4bc187147eba79c83392d3c79a40efd54ba81f6e8d8b9ea8c35
|
File details
Details for the file symbac-0.5.2-py3-none-any.whl.
File metadata
- Download URL: symbac-0.5.2-py3-none-any.whl
- Upload date:
- Size: 213.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
68228445d17788934b0c3cbf279dfb70e6e91edc52195ce66644a93292f32271
|
|
| MD5 |
86545dd41a27691d453d18b8acd06461
|
|
| BLAKE2b-256 |
748f1d2c163ab4e7c9fb1032256124813081356aad67889694253e513741f63c
|












