BrainWeb-based multimodal models of 20 normal brains
Project description
The following example may be launched interactively via any of the following:
BrainWeb-based multimodal models of 20 normal brains
This project was initially inspired by “BrainWeb: 20 Anatomical Models of 20 Normal Brains.”
However there are a number of generally useful tools, image processing & display functions included in this project. For example, this includes volshow() for interactive comparison of multiple 3D volumes, get_file() for caching data URLs, and register() for image coregistration.
Download and Preprocessing for PET-MR Simulations
This notebook will not re-download/re-process files if they already exist.
Output data
~/.brainweb/subject_*.npz: dtype(shape): float32(127, 344, 344)
-
~/.brainweb/subject_*.bin.gz: dtype(shape): uint16(362, 434, 362)
Install
pip install brainweb
Author: Casper da Costa-Luis <casper.dcl@physics.org>
Date: 2017-2020
Licence: MPLv2.0
from __future__ import print_function, division
%matplotlib notebook
import brainweb
from brainweb import volshow
import numpy as np
from os import path
from tqdm.auto import tqdm
import logging
logging.basicConfig(level=logging.INFO)Raw Data
# download
files = brainweb.get_files()
# read last file
data = brainweb.load_file(files[-1])
# show last subject
print(files[-1])
volshow(data, cmaps=['gist_ncar']);~/.brainweb/subject_54.bin.gz
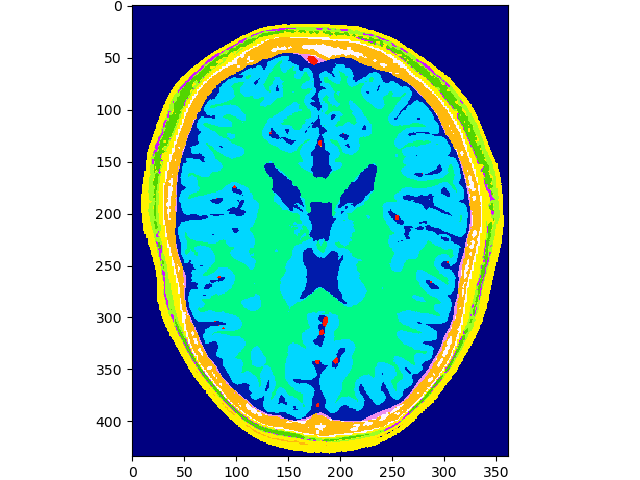
Transform
Convert raw image data:
Siemens Biograph mMR resolution (~2mm) & dimensions (127, 344, 344)
PET/T1/T2/uMap intensities
PET defaults to FDG intensity ratios; could use e.g. Amyloid instead
randomised structure for PET/T1/T2
t (1 + g [2 G_sigma(r) - 1]), where
r = rand(127, 344, 344) in [0, 1),
Gaussian smoothing sigma = 1,
g = 1 for PET; 0.75 for MR, and
t = the PET or MR piecewise constant phantom
# show region probability masks
PetClass = brainweb.FDG
label_probs = brainweb.get_label_probabilities(files[-1], labels=PetClass.all_labels)
volshow(label_probs[brainweb.trim_zeros_ROI(label_probs)], titles=PetClass.all_labels, frameon=False);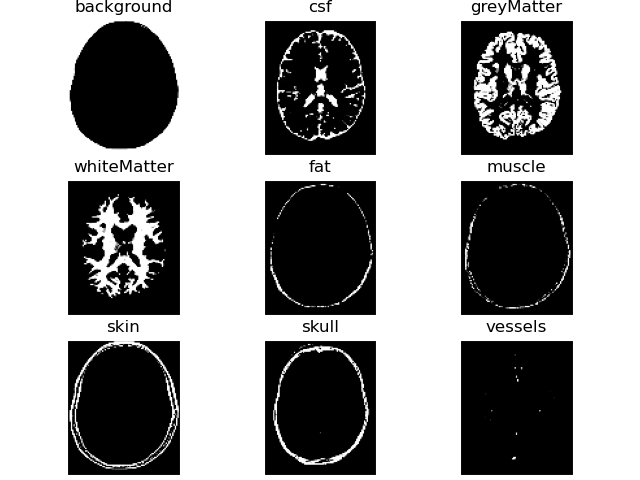
brainweb.seed(1337)
for f in tqdm(files, desc="mMR ground truths", unit="subject"):
vol = brainweb.get_mmr_fromfile(
f,
petNoise=1, t1Noise=0.75, t2Noise=0.75,
petSigma=1, t1Sigma=1, t2Sigma=1,
PetClass=PetClass)# show last subject
print(f)
volshow([vol['PET' ][:, 100:-100, 100:-100],
vol['uMap'][:, 100:-100, 100:-100],
vol['T1' ][:, 100:-100, 100:-100],
vol['T2' ][:, 100:-100, 100:-100]],
cmaps=['hot', 'bone', 'Greys_r', 'Greys_r'],
titles=["PET", "uMap", "T1", "T2"],
frameon=False);~/.brainweb/subject_54.bin.gz
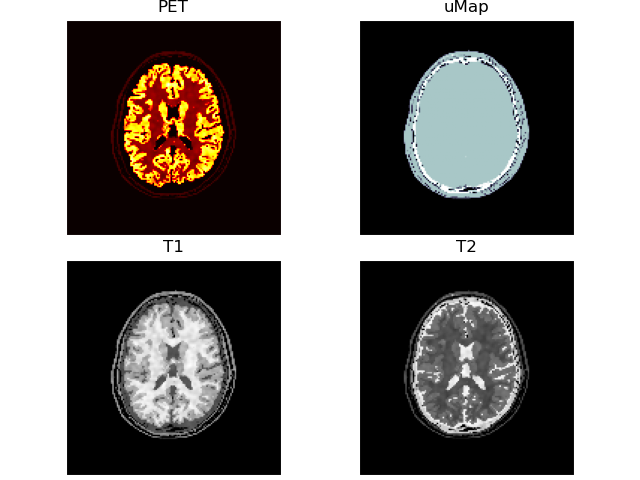
# add some lesions
brainweb.seed(1337)
im3d = brainweb.add_lesions(vol['PET'])
volshow(im3d[:, 100:-100, 100:-100], cmaps=['hot']);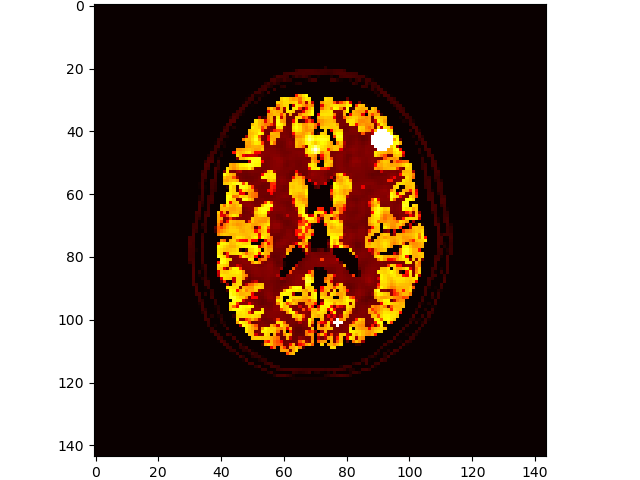
# bonus: use brute-force registration to transform
#!pip install -U 'brainweb[register]'
reg = brainweb.register(
data[:, ::-1], target=vol['PET'],
src_resolution=brainweb.Res.brainweb,
target_resolution=brainweb.Res.mMR)
volshow({
"PET": vol['PET'][:, 100:-100, 100:-100],
"RawReg": reg[ :, 100:-100, 100:-100],
"T1": vol['T1' ][:, 100:-100, 100:-100],
}, cmaps=['hot', 'gist_ncar', 'Greys_r'], ncols=3, tight_layout=5, figsize=(9.5, 3.5), frameon=False);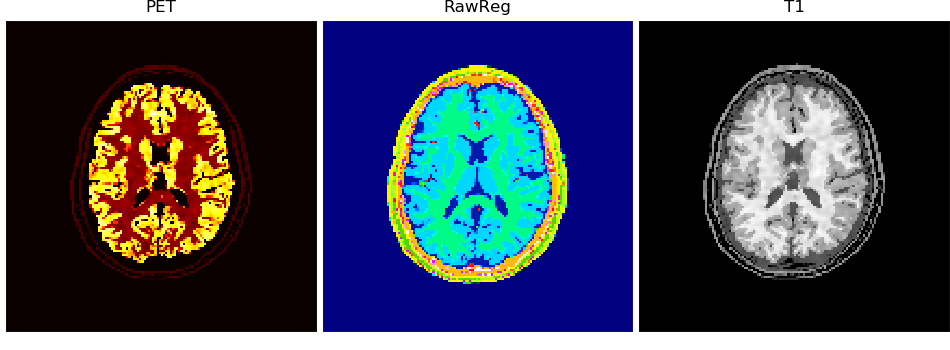
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file brainweb-1.6.2.tar.gz.
File metadata
- Download URL: brainweb-1.6.2.tar.gz
- Upload date:
- Size: 13.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/50.3.0 requests-toolbelt/0.9.1 tqdm/4.50.2 CPython/3.7.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cfca5a08d519326662d4135034f340cee7a664e1e02d85bb348c217baeb74eaf
|
|
| MD5 |
44f8763fb75f9b114b9bf8789cb012c0
|
|
| BLAKE2b-256 |
ee4cc26866bc11ee2d82c3592ee9c67afe6d67e9a566b1490cf1f103a4489e13
|
File details
Details for the file brainweb-1.6.2-py2.py3-none-any.whl.
File metadata
- Download URL: brainweb-1.6.2-py2.py3-none-any.whl
- Upload date:
- Size: 11.5 kB
- Tags: Python 2, Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/50.3.0 requests-toolbelt/0.9.1 tqdm/4.50.2 CPython/3.7.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
db7d16ccc7d5f1dd02094a5eeb991b1554a47fceac34bfab3d6cfb634a7752f1
|
|
| MD5 |
2aad724a3b75c666f7d83b751e4795ef
|
|
| BLAKE2b-256 |
d882e9a43fb3468158dfe6d7a1b6e642bc243abc16fc02175f7491abae990394
|

















