No project description provided
Project description
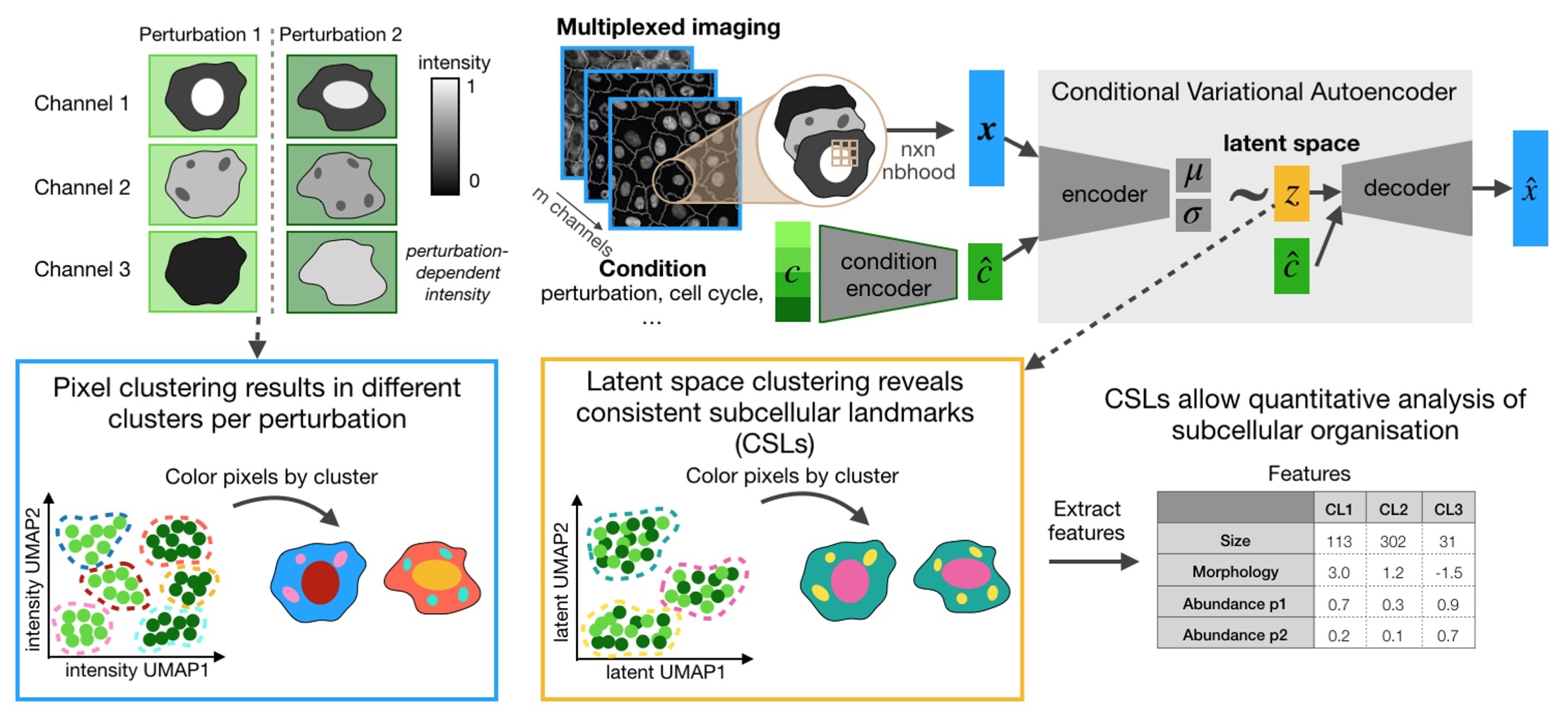
CAMPA is a framework for quantiative analysis of subcellular multi-channel imaging data. It consists of a workflow that generates consistent subcellular landmarks (CSLs) using conditional Variational Autoencoders (cVAE). The output of the CAMPA workflow is an anndata object that contains interpretable per-cell features summarizing the molecular composition and spatial arrangement of CSLs inside each cell.
Visit our documentation for installation and usage examples.
Manuscript
Please see our preprint “Learning consistent subcellular landmarks to quantify changes in multiplexed protein maps” (Spitzer, Berry et al. (2022)) to learn more.
Installation
CAMPA was developed for Python 3.9 and can be installed by running:
pip install campa
Contributing
We are happy about any contributions! Before you start, check out our contributing guide.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file campa-1.1.0-py3-none-any.whl.
File metadata
- Download URL: campa-1.1.0-py3-none-any.whl
- Upload date:
- Size: 101.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.0 CPython/3.9.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8622d0fd7c46989d089b9f49c495c2d7db8dc75d962d2d355ccf7e15e4b019b3
|
|
| MD5 |
a5b9bc5e97f677422e32da997518c166
|
|
| BLAKE2b-256 |
f2b828100a507c5e9eeb61fac08a65424cb41de2b8a526929e33f6945712adb7
|











