Compute single-cell cell-type expression specificity
Project description
CELLEX
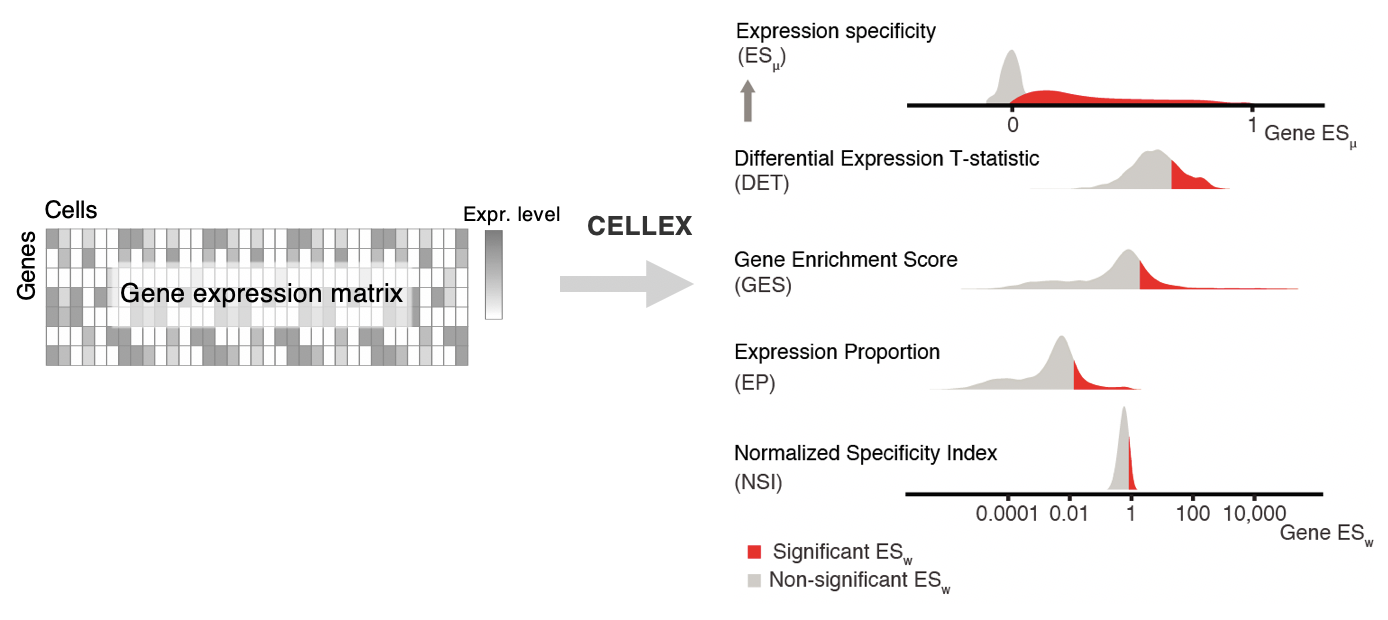
CELLEX (CELL-type EXpression-specificity) is a tool for computing cell-type Expression Specificity (ES) profiles. It employs a "wisdom of the crowd"-approach by integrating multiple ES metrics, thus combining complementary cell-type ES profiles, to capture multiple aspects of ES and obtain improved robustness.
Contents
Documentation
The documentation for CELLEX can be accessed in the following ways:
- CELLEX Wiki : main documentation on the usage of CELLEX
- CELLEX API docs: documentation of CELLEX API/functions
- Publication: technical details on the CELLEX method. Genetic mapping of etiologic brain cell types for obesity (Timshel eLife, 2020, Appendix)
We are continually updating the documentation for CELLEX. If some information is missing, please submit your request or question via our issue tracker.
Quick start
This brief tutorial showcases the core features of CELLEX.
TL;DR
import numpy as np
import pandas as pd
import cellex
data = pd.read_csv("./data.csv", index_col=0)
metadata = pd.read_csv("./metadata.csv", index_col=0)
eso = cellex.ESObject(data=data, annotation=metadata, verbose=True)
eso.compute(verbose=True)
eso.results["esmu"].to_csv("mydataset.esmu.csv.gz")
Walkthrough
Setup
Option A: Install the latest release from PyPi
pip install cellex
Option B: Install the development version from this repo
Clone the development repo and install from source using pip. The development version may contain bug fixes that have not been released, as well as experimental features.
git clone https://github.com/perslab/CELLEX.git --branch develop --single-branch
cd CELLEX
pip install -e .
Import modules
import numpy as np # needed for formatting data for this tutorial
import pandas as pd # needed for formatting data for this tutorial
import cellex
Load input data and metadata
data = pd.read_csv("./data.csv", index_col=0)
metadata = pd.read_csv("./metadata.csv", index_col=0)
Data format
Data may consist of UMI counts (integer) for each gene and cell.
| cell_1 | ... | cell_9 | |
|---|---|---|---|
| gene_x | 0 | ... | 4 |
| ... | ... | ... | ... |
| gene_z | 3 | ... | 1 |
Shape: m genes by n cells.
Metadata format
Metadata should consist of unique cell id's and matching annotation (string).
| cell_id | cell_type |
|---|---|
| cell_1 | type_A |
| ... | ... |
| cell_9 | type_C |
Shape: n cells by 2.
Create ESObject and compute ESmu
eso = cellex.ESObject(data=data, annotation=metadata, verbose=True)
eso.compute(verbose=True)
View Expression Specificity scores
All results are accessible via the results attribute of the ESObject.
eso.results["esmu"]
Save result(s)
Pro-tip: Using CELLEX with CELLECT
The ESmu scores may be used with CELLECT. CELLECT requires that genes are in the Human Ensembl Gene ID format. CELLEX provides a simple renaming utility for this purpose:
cellex.utils.mapping.mouse_ens_to_human_ens(eso.results["esmu"], drop_unmapped=True, verbose=True)
Save ESmu
eso.results["esmu"].to_csv("mydataset.esmu.csv.gz")
Save all or specific results
eso.save_as_csv(keys=["all"], verbose=True)
Output format
Output consist of Expression Specificity Weights (float) for each gene and cell-type. ESmu values lie in the range [0,1].
| type_A | ... | type_C | |
|---|---|---|---|
| gene_x | 0.0 | ... | 0.9 |
| ... | ... | ... | ... |
| gene_z | 0.1 | ... | 0.2 |
Shape: m genes by x unique annotations. N.B. a number of genes may be removed during preprocessing.
Tutorials
Various tutorials and walkthroughs will be made available here, while the Wiki is in the making. These Jupyter Notebooks cover everything from downloading CELLEX and data to analysis and plotting.
- Demo: Downloading and running CELLEX with sample Mousebrain Atlas data
- Demo: Downloading and running CELLEX with sample MOCA data
About
Developers
- Tobias Overlund Stannius (University of Copenhagen) @TobiasStannius
- Pascal Nordgren Timshel (University of Copenhagen) @ptimshel
Contact
Please create an issue in this repo, if you encounter any problems using CELLEX. Alternatively, you may write an email to timshel(at)sund.ku.dk
References
If you find CELLEX useful for your research, please consider citing: Timshel (eLife, 2020): Genetic mapping of etiologic brain cell types for obesity
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cellex-1.2.2.tar.gz.
File metadata
- Download URL: cellex-1.2.2.tar.gz
- Upload date:
- Size: 7.5 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.8.0 pkginfo/1.8.2 readme-renderer/34.0 requests/2.27.1 requests-toolbelt/0.9.1 urllib3/1.26.9 tqdm/4.63.0 importlib-metadata/4.11.3 keyring/23.5.0 rfc3986/2.0.0 colorama/0.4.4 CPython/3.10.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3966eb164fe49b9d5c40450929084b799288d6d8bc061ece758c028087a8d6d1
|
|
| MD5 |
1d53d5ceb7f5f3d21263a983d862f394
|
|
| BLAKE2b-256 |
c94ff7f7c650a47237f95eb6c278cad845806009df62530210dee238370e4cd7
|
File details
Details for the file cellex-1.2.2-py3-none-any.whl.
File metadata
- Download URL: cellex-1.2.2-py3-none-any.whl
- Upload date:
- Size: 2.6 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.8.0 pkginfo/1.8.2 readme-renderer/34.0 requests/2.27.1 requests-toolbelt/0.9.1 urllib3/1.26.9 tqdm/4.63.0 importlib-metadata/4.11.3 keyring/23.5.0 rfc3986/2.0.0 colorama/0.4.4 CPython/3.10.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d3f7d920c038e547b07e25e524e64f14541178a5699b5da43f6cd76d253bf3da
|
|
| MD5 |
2463b7fcb6f38463fb503e8a81f84d1a
|
|
| BLAKE2b-256 |
694b1a4129468ef41d85eb00de019327e673083bd0159f1e632f0063003fa261
|