Declaratively configured pipeline for image classification
Project description
Classification training pipeline
My puny attempt to build reusable training pipeline for image classification
- Motivation
- Installation
- Usage guide
- Analyzing Experiments Results
- What is supported?
- Custom architectures, callbacks, metrics
- Examples
- Faq
Motivation
Idea for this project came from my first attempts to participate in Kaggle competitions. My programmers heart was painfully damaged by looking on my own code as well as on other people kernels. Code was highly repetitive, suffering from numerous reimplementations of same or almost same things through the kernels, model/experiment configuration was often mixed with models code, in other words - from programmer perspective it all looked horrible.
So I decided to extract repetitive things into framework that will work at least for me and will follow these statements:
- experiment configurations should be cleanly separated from model definitions;
- experiment configuration files should be easy to compare and should fully describe experiment that is being performed except for the dataset;
- common blocks like an architecture, callbacks, storing model metrics, visualizing network predictions, should be written once and be a part of common library
Installation
pip install classification_pipeline
Note: this package requires python 3.6
Usage guide
Training a model
Let's start from a simple example of classification. Suppose, your data are structured as follows: a .cvs file with images ids and their labels and a folder with all these images. For training a neural network to classify these images all you need are few lines of python code:
import musket_core
from classification_pipeline import classification
class ProteinDataGenerator:
def __init__(self, paths, labels):
self.paths, self.labels = paths, labels
def __len__(self):
return len(self.paths)
def __getitem__(self, idx):
X,y = self.__load_image(self.paths[idx]),self.labels[idx]
return PredictionItem(self.paths[idx],X, y)
def __load_image(self, path):
R = Image.open(path + '_red.png')
G = Image.open(path + '_green.png')
B = Image.open(path + '_blue.png')
im = np.stack((
np.array(R),
np.array(G),
np.array(B),
), -1)
return im
dataset = ProteinDataGenerator(paths,labels)
cfg = classification.parse("config.yaml")
cfg.fit(dataset)
Looks simple, but there is a config.yaml file in the code, and probably it is the place where everything actually happens.
architecture: DenseNet201 #pre-trained model we are going to use
pooling: avg
augmentation: #define some minimal augmentations on images
Fliplr: 0.5
Flipud: 0.5
classes: 28 #define the number of classes
activation: sigmoid #as we have multilabel classification, the activation for last layer is sigmoid
weights: imagenet #we would like to start from network pretrained on imagenet dataset
shape: [224, 224, 3] #our desired input image size, everything will be resized to fit
optimizer: Adam #Adam optimizer is a good default choice
batch: 16 #our batch size will be 16
lr: 0.005
copyWeights: true
metrics: #we would like to track some metrics
- binary_accuracy
- macro_f1
primary_metric: val_binary_accuracy #the most interesting metric is val_binary_accuracy
primary_metric_mode: max
callbacks: #configure some minimal callbacks
EarlyStopping:
patience: 3
monitor: val_macro_f1
mode: max
verbose: 1
ReduceLROnPlateau:
patience: 2
factor: 0.3
monitor: val_binary_accuracy
mode: max
cooldown: 1
verbose: 1
loss: binary_crossentropy #we use binary_crossentropy loss
stages:
- epochs: 10 #let's go for 100 epochs
So as you see, we have decomposed our task in two parts, code that actually trains the model and experiment configuration, which determines the model and how it should be trained from the set of predefined building blocks.
What does this code actually do behind the scenes?
- it splits your data into 5 folds, and trains one model per fold;
- it takes care of model checkpointing, generates example image/label tuples, collects training metrics. All this data will
be stored in the folders just near your
config.yaml; - All your folds are initialized from fixed default seed, so different experiments will use exactly the same train/validation splits.
Image Augmentations
Framework uses awesome imgaug library for augmentation, so you only need to configure your augmentation process in declarative way like in the following example:
augmentation:
Fliplr: 0.5
Flipud: 0.5
Affine:
scale: [0.8, 1.5] #random scalings
translate_percent:
x: [-0.2,0.2] #random shifts
y: [-0.2,0.2]
rotate: [-16, 16] #random rotations on -16,16 degrees
shear: [-16, 16] #random shears on -16,16 degrees
Freezing/Unfreezing encoder
Freezing encoder is often used with transfer learning. If you want to start with frozen encoder just add
freeze_encoder: true
stages:
- epochs: 10 #Let's go for 10 epochs with frozen encoder
- epochs: 100 #Now let's go for 100 epochs with trainable encoder
unfreeze_encoder: true
in your experiments configuration, then on some stage configuration just add
unfreeze_encoder: true
to stage settings.
Note: This option is not supported for DeeplabV3 architecture.
Custom datasets
You can declare your own dataset class as in this example:
from musket_core.datasets import PredictionItem
import os
import imageio
import pandas as pd
import numpy as np
import cv2
class Classification:
def __init__(self,imgPath):
self.species = ['Black-grass', 'Charlock', 'Cleavers', 'Common Chickweed', 'Common wheat',
'Fat Hen', 'Loose Silky-bent', 'Maize', 'Scentless Mayweed',
'Shepherds Purse', 'Small-flowered Cranesbill', 'Sugar beet']
self.data = []
self.targets = []
self.ids = []
for s_id, s in enumerate(self.species):
s_folder = os.path.join(imgPath,s)
for file in os.listdir(s_folder):
self.data.append(os.path.join(s_folder, file))
self.targets.append(s_id)
self.ids.append(file)
def __len__(self):
return len(self.data)
def __getitem__(self, item):
item_file = self.data[item]
target = self.targets[item]
t = np.zeros(len(self.species))
t[target] = 1.0
image = self.read_image(item_file, (224,224))
return PredictionItem(self.ids[item], image, t)
def read_image(self, filepath, target_size=None):
img = cv2.imread(filepath, cv2.IMREAD_COLOR)
img = cv2.resize(img.copy(), target_size, interpolation = cv2.INTER_AREA)
return img
Multi output classification
Sometimes you need to create network that performs several classification tasks at the same moment, in this situation
you need to declare classes , activation and loss as the lists of class counts, activation functions and losses
like in the following snippet:
classes: [ 4, 4 ] #define the number of classes
activation: [sigmoid,sigmoid]
loss:
- binary_crossentropy
- binary_crossentropy
primary_metric: val_loss #the most interesting metric is val_binary_accuracy
primary_metric_mode: min
it is also very likely that you need to change primary metric to val_loss
Balancing your data
One common case is the situation when part of your images does not contain any objects of interest, like in Airbus ship detection challenge. More over your data may be to heavily inbalanced, so you may want to rebalance it. Alternatively you may want to inject some additional images that do not contain objects of interest to decrease amount of false positives that will be produced by the framework.
These scenarios are supported by negatives and validation_negatives settings of training stage configuration,
these settings accept following values:
- none - exclude negative examples from the data
- real - include all negative examples
- integer number(1 or 2 or anything), how many negative examples should be included per one positive example
if you are using this setting your dataset class must support isPositive method which returns true for indexes
which contain positive examples:
def isPositive(self, item):
pixels=self.ddd.get_group(self.ids[item])["EncodedPixels"]
for mask in pixels:
if isinstance(mask, str):
return True;
return False
Multistage training
Sometimes you need to split your training into several stages. You can easily do it by adding several stage entries in your experiment configuration file like in the following example:
stages:
- epochs: 6 #Train for 6 epochs
negatives: none #do not include negative examples in your training set
validation_negatives: real #validation should contain all negative examples
- lr: 0.0001 #let's use different starting learning rate
epochs: 6
negatives: real
validation_negatives: real
- loss: lovasz_loss #let's override loss function
lr: 0.00001
epochs: 6
initial_weights: ./fpn-resnext2/weights/best-0.1.weights #let's load weights from this file
Stage entries allow you to configure custom learning rate, balance of negative examples, callbacks, loss function and even initial weights which should be used on a particular stage.
Composite losses
Framework supports composing loss as a weighted sum of predefined loss functions. For example, following construction
loss: binary_crossentropy+0.1*dice_loss
will result in loss function which is composed from binary_crossentropy and dice_loss functions.
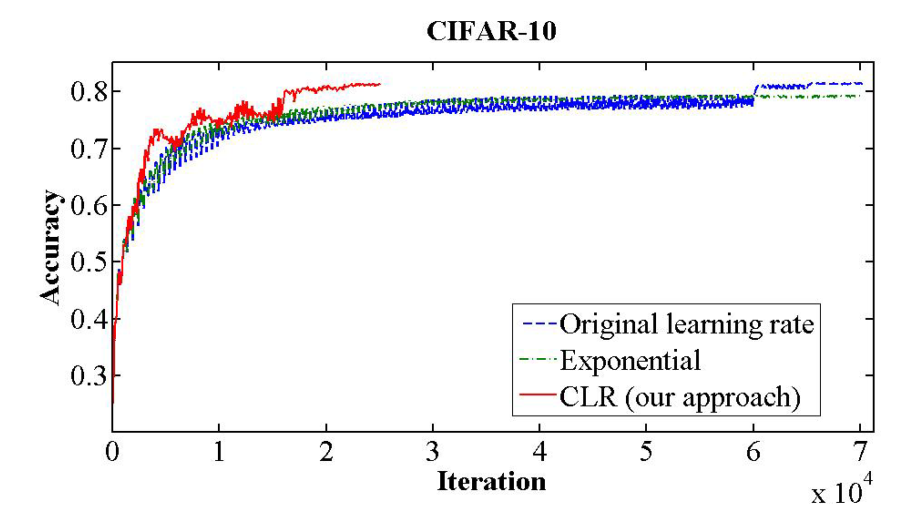
Cyclical learning rates
As told in Cyclical learning rates for training neural networks CLR policies can provide quicker converge for some neural network tasks and architectures.
We support them by adopting Brad Kenstler CLR callback for Keras.
If you want to use them, just add CyclicLR in your experiment configuration file as shown below:
callbacks:
EarlyStopping:
patience: 40
monitor: val_binary_accuracy
verbose: 1
CyclicLR:
base_lr: 0.0001
max_lr: 0.01
mode: triangular2
step_size: 300
LR Finder
Estimating optimal learning rate for your model is an important thing, we support this by using slightly changed version of Pavel Surmenok - Keras LR Finder
cfg = classification.parse(config.yaml)
ds = SimplePNGMaskDataSet("./train","./train_mask") - ???????????????????
finder=cfg.lr_find(ds,start_lr=0.00001,end_lr=1,epochs=5)
finder.plot_loss(n_skip_beginning=20, n_skip_end=5)
plt.show()
finder.plot_loss_change(sma=20, n_skip_beginning=20, n_skip_end=5, y_lim=(-0.01, 0.01))
plt.show()
will result in this couple of helpful images:
Training on crops
Your images can be too large to train model on them. In this case you probably want to train model on crops. All that you need to do is to specify number of splits per axis. For example, following lines in config
shape: [768, 768, 3]
crops: 3
will lead to splitting each image into 9 cells (3 horizontal splits and 3 vertical splits) and training model on these splits. Augmentations will be run separately on each cell.
During prediction time, your images will be split into these cells, prediction will be executed on each cell, and then results will be assembled in single final mask. Thus the whole process of cropping will be invisible from a consumer perspective.
Using trained model
Okey, our model is trained, now we need to actually do image classification. Let's say, we need to run image classification on images in the directory and store results in csv file:
predictions = []
images = []
#Now let's use best model from fold 0 to do image segmentation on images from images_to_segment
preds = cfg.predict_all_to_array(dataset_test, 0, 0)
for i, item in enumerate(dataset_test):
images.append(dataset_test.get_id(i))
p = np.argmax(preds[i])
predictions.append(dataset_test.get_label(p))
#Let's store results in csv
df = pd.DataFrame.from_dict({'file': images, 'species': predictions})
df.to_csv('submission.csv', index=False)
Ensembling predictions
And what if you want to ensemble models from several folds? Just pass a list of fold numbers to
predict_all_to_array like in the following example:
cfg.predict_all_to_array(dataset_test, [0,1,2,3,4], 0)
Another supported option is to ensemble results from extra test time augmentation (flips) by adding keyword arg ttflips=True.
Custom evaluation code
Sometimes you need to run custom evaluation code. In such case you may use: evaluateAll method, which provides an iterator
on the batches containing original images, training masks and predicted masks
for batch in cfg.evaluateAll(ds,2):
for i in range(len(batch.predicted_maps_aug)):
masks = ds.get_masks(batch.data[i])
for d in range(1,20):
cur_seg = binary_opening(batch.predicted_maps_aug[i].arr > d/20, np.expand_dims(disk(2), -1))
cm = rle.masks_as_images(rle.multi_rle_encode(cur_seg))
pr = f2(masks, cm);
total[d]=total[d]+pr
Accessing model
You may get trained keras model by calling: cfg.load_model(fold, stage).
Analyzing experiments results
Okey, we have done a lot of experiments and now we need to compare the results and understand what works better. This repository contains script which may be used to analyze folder containing sub folders with experiment configurations and results. This script gathers all configurations, diffs them by doing structural diff, then for each configuration it averages metrics for all folds and generates csv file containing metrics and parameters that was actually changed in your experiment like in the following example
This script accepts following arguments:
- inputFolder - root folder to search for experiments configurations and results
- output - file to store aggregated metrics
- onlyMetric - if you specify this option all other metrics will not be written in the report file
- sortBy - metric that should be used to sort results
Example:
python analize.py --inputFolder ./experiments --output ./result.py
What is supported?
At this moment classification pipeline supports following pre-trained models:
- Resnet
- ResNet18
- ResNet34
- ResNet50
- ResNet101
- ResNet152
- ResNeXt50
- ResNeXt101
- VGG:
- VGG16
- VGG19
- InceptionV3
- InceptionResNetV2
- Xception
- MobileNet
- MobileNetV2
- DenseNet:
- DenseNet121
- DenseNet169
- DenseNet201
- NasNet:
- NASNetMobile
- NASNetLarge
Each architecture also supports some specific options, list of options is documented in segmentation RAML library.
Supported augmentations are documented in augmentation RAML library.
Callbacks are documented in callbacks RAML library.
Custom architectures, callbacks, metrics
Classification pipeline uses keras custom objects registry to find entities, so if you need to use custom loss function, activation or metric all that you need to do is to register it in Keras as:
keras.utils.get_custom_objects()["my_loss"]= my_loss
If you want to inject new architecture, you should register it in classification.custom_models dictionary.
For example:
classification.custom.models['MyUnet']=MyUnet
where MyUnet is a function that accepts architecture parameters as arguments and returns an instance
of keras model.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file classification_pipeline-0.432.tar.gz.
File metadata
- Download URL: classification_pipeline-0.432.tar.gz
- Upload date:
- Size: 12.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.12.1 pkginfo/1.4.2 requests/2.22.0 setuptools/41.4.0 requests-toolbelt/0.8.0 tqdm/4.36.1 CPython/3.6.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
395a1f38c4829cdc1173cfb7ca545505f36d72523e6948f63ad7f6a7f54f6292
|
|
| MD5 |
ef41aba91910be627a81813443df79d0
|
|
| BLAKE2b-256 |
bf0cec1ea1d0d61bcb3ca4423d73a8b98ac9ec2de0310e5986eb183c9d810bd0
|
File details
Details for the file classification_pipeline-0.432-py2.py3-none-any.whl.
File metadata
- Download URL: classification_pipeline-0.432-py2.py3-none-any.whl
- Upload date:
- Size: 11.7 kB
- Tags: Python 2, Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.12.1 pkginfo/1.4.2 requests/2.22.0 setuptools/41.4.0 requests-toolbelt/0.8.0 tqdm/4.36.1 CPython/3.6.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5268ad396f12883ce6e5ee3d78483361d45c5b008a192f8798ecad6a2358ec70
|
|
| MD5 |
2dda3356c0017ed7eef93d7117aab414
|
|
| BLAKE2b-256 |
8c6f9b4663244ca69717c31885fab34ea28701a51fdf2cf6441e03025647df02
|