Constitutive Relation Inference Toolkit
Project description
CRIKit: The Constitutive Relation Inference Toolkit
Quick Start | Documentation | Installation Guide
Overview
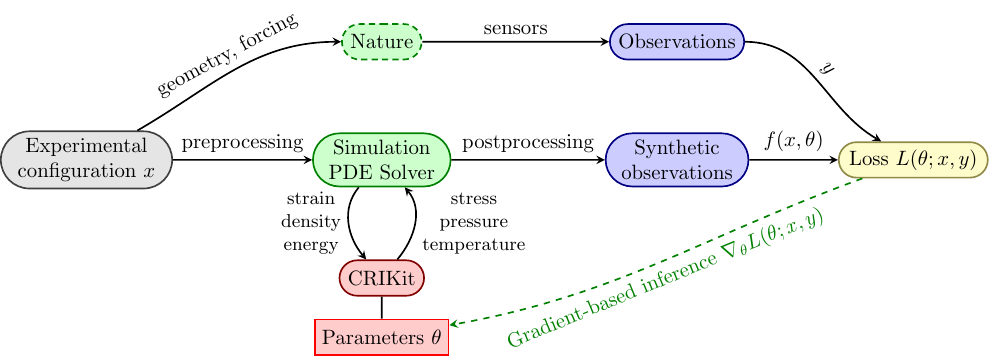
CRIKit integrates FEniCS and Pyadjoint with machine learning libraries
like JAX and TensorFlow, and provides tools to infer physically-compatible constitutive relations from sparse, noisy observations of a system modeled by partial differential equations. CRIKit bridges the FEniCS world with those of
JAX and TensorFlow by storing covering maps between abstract Space classes that
represent spaces like a FEniCS FunctionSpace or a space of JAX arrays of a
particular shape, or a direct sum of multiple Spaces.
CRIKit also provides tools to help perform post-processing, such as observation operators, as well as a collection of loss functions.
Quick Start
Constructing And Optimizing a CR
This guide will show you the basics of constructing and optimizing a simple CR that represents linear elasticity, assuming that you're already familiar with the basics of FEniCS. You can compare the mechanics of CRIKit to that of FEniCS directly by comparing this example to the 2D linear elasticity example from Numerical tours of Computational Mechanics using FEniCS. The primary difference between the model shown here and the linked example in the previous sentence is that here we use a geometrically nonlinear model, as described in the documentation for the libCEED hyperelasticity example.
from crikit import *
import jax
from jax import numpy as jnp
import numpy as np
from functools import partial
# set up mesh, FunctionSpace, etc
fe_order = 2
dims = 2
Nx, Ny = 50, 5
L = 20.
H = 1.
mesh = RectangleMesh(Point(0., 0.), Point(L, H), Nx, Ny)
V = VectorFunctionSpace(mesh, "CG", fe_order)
quad_params = {'quadrature_degree' : fe_order + 1}
set_default_covering_params(domain=mesh.ufl_domain(),
quad_params=quad_params)
u = Function(V)
def left_boundary(x, on_boundary):
return near(x[0], 0.)
bcs = [DirichletBC(V, Constant((0., 0.)), left_boundary)]
# these will tell CRIKit what the inputs and ouputs to the CR
# are so that we can automatically generate the scalar and form-invariants
# Let's suppose you want the Cauchy stress tensor as a function of the
# strain sym(grad(u))
input_types = (TensorType.make_symmetric(2, dims, 'strain'),)
output_type = TensorType.make_symmetric(2, dims, 'stress')
# initial guess of parameters
Youngs = 1.0e5
Poisson = 0.3
lmbda = (Youngs * Poisson) / ((1 + Poisson) * (1 - 2 * Poisson))
mu = Youngs / (2 * (1 + Poisson))
# since this is 2-d, we need to use a modified version of lambda
# to make our initial guesses physical
lmbda = 2 * lmbda * mu / (lmbda + 2 * mu)
theta = array([lmbda, mu])
def cr_func(invariants, theta):
lmbda, mu = theta
return jnp.array([lmbda * jnp.log1p(invariants[0]), 2 * mu])
cr = CR(output_type, input_types, cr_func, params=[theta])
# If you're in a Jupyter notebook, run this at the bottom of a cell instead of
# calling `print()` on it to get neatly-rendered HTML output.
# This function shows you a description of the scalar and form invariants of `cr`
# in the order they are placed in the arrays
print(cr.invariant_descriptions())
# set the default covering params for crikit.covering so we can automatically
# generate covering maps between spaces of FEniCS Functions and JAX arrays
# Let's just pretend that degree 3 is sufficient quadrature for whatever problem
# we're solving
quad_params = {'quadrature_degree' : 3}
set_default_covering_params(domain=mesh.ufl_domain(), quad_params=quad_params)
# create_ufl_standins() returns a tuple of objects that can act as standins
# for the output of a CR. You can't directly call the CR on the inputs because
# the CR expects JAX arrays as an input, not a FEniCS Function. You'll instead have
# to assembly the variational form F using assemble_with_cr(), which will generate
# a covering map from the space of FEniCS Functions to the space of JAX arrays
# using crikit.covering (and likewise from the output JAX array space to a space of
# `crikit.fe.Function`s), use it to get appropriate arguments, call the CR, and project the result
# back into a Function
target_shape = tuple(i for i in cr.target.shape() if i != -1)
standin_sigma, = create_ufl_standins((target_shape,))
# create your form as if standin_sigma were (cr(sym(grad(u)))
v = TestFunction(V)
# external force
f = Constant((0,-1e-3), name='force')
F = inner(standin_sigma, sym(grad(v))) * dx - inner(f, v) * dx
# define a new sub-tape that records the actions of this equation
with push_tape():
# a function that we can assemble the variational form into
# using the `tensor` kwarg of `crikit.assemble()`, which
# is directly passed on to `crikit.fe.backend.assemble()`
# (e.g. `fenics.assemble()`)
residual = Function(V)
# input to the CR is sym(grad(u))
assemble_with_cr(F, cr, sym(grad(u)), standin_sigma, tensor=residual,
quad_params=quad_params)
ucontrol = Control(u)
# a ReducedFunction to represent the residual as a function of `u`
res_rf = ReducedFunction(residual, ucontrol)
# an object to represent the equation defined above
red_eq = ReducedEquation(res_rf, bcs, homogenize_bcs(bcs))
# and an object to solve it. Make sure your .petscrc is set appropriately!
# if you want to pass an assembled Jacobian, use 'jmat_type' : 'assembled',
# but if you want the solver to instead use the matrix-free Jacobian action,
# pass 'jmat_type' : 'action'
solver = SNESSolver(red_eq, {'jmat_type' : 'assembled'})
pred_u = solver.solve(ucontrol)
# define a loss function and an observer
num_slices = 100
seed = 0
# sliced quadratic Wasserstein distance
loss = SlicedWassersteinDistance(V, num_slices, jax.random.PRNGKey(seed), p=2)
class ObservedSubDomain(SubDomain):
def inside(self, x, on_boundary):
... # return appropriate True/False if x is in the observed subdomain or not
# observe only on a given SubDomain
observer = SubdomainObserver(mesh, ObservedSubDomain())
# get your observations from somewhere as a Function in V
obs = ...
err = loss(observer(obs), observer(pred_u))
Jhat = ReducedFunctional(err, Control(theta))
#check the derivative
h = np.random.randn(*theta.shape)
v = array(1.0) # test the adjoint
assert taylor_test(Jhat, theta, h, v=v) >= 1.9
# choose an optimization method
opt_method = 'L-BFGS-B'
optimal_params = minimize(Jhat, method=opt_method)
Installation
First, install FEniCS. Then you can install the latest development version of this package by running
pip install .
or, to install in editable mode (useful for devs),
pip install -e .
There are four optional sets of dependencies that can be installed by listing
any of them (with no spaces) in square brackets. Or if you want to install all
of them, you can use the key all, so the two lines below are equivalent.
pip install -e .[test,visualization,tensorflow,doc]
pip install -e .[all]
If you would rather install the latest release version of CRIKit, you can instead run
pip install crikit
Make sure you have install CRIKit into an environment with a working FEniCS
installation, or the build might fail. In particular, if you encounter
errors building petsc4py, ensure that the FEniCS installation in your
environment works.
Setting Up a Conda Environment
FEniCS provides a conda package, so installation into a conda environment is simple.
conda create --name fenics2019 python=3.7 --no-default-packages -y -q
conda activate fenics2019
conda install -c conda-forge fenics=2019.1.0 -y -q
pip install -e .[all]
Whenever you want to enter this environment, run conda activate fenics2019.
Documentation
Documentation is done with Sphinx.
Documentation can be built by running make in the docs folder. By default,
that will create HTML documentation in docs/build/html, and you can view it by
opening up docs/build/html/index.html in your favorite browser.
Run make todo=true to include Todo blocks in the output. Otherwise, they won't show up.
You can also run make help to see a list of other formats that can be built. For instance,
make coveragecreates a file showing what classes/functions are missing documentation.make doctesttests the example code in the documentation.
Examples
Example programs are in the examples folder. Run with the --help argument to
see command-line parameters. For example,
cd examples/p-stokes
python p-stokes.py --help
Tests
You can run the main tests with the command below.
python3 -m pytest tests
You can run a specific test by running python3 -m pytest tests -k test_name,
where test_name is all or part of the test name. For example,
python3 -m pytest tests/crikit/test_assemble_with_cr.py -k test_assemble_with_cr_scalar
or
python3 -m pytest tests/crikit/test_assemble_with_cr.py -k scalar
Some implementation files and documentation files have doctests. Those can be run like so:
python -m pytest --doctest-modules --rootdir tests crikit
cd docs && make doctest
Developer style guidelines
CRIKit uses the auto-formatter black to ensure the code has a consistent style. To automatically run it before each git commit, use the following commands.
pip install -e .[dev]
pre-commit install
Support
This material is based upon work supported by the National Science Foundation under Grant No. 1835825 and 1835792. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file crikit-0.1.5.tar.gz.
File metadata
- Download URL: crikit-0.1.5.tar.gz
- Upload date:
- Size: 198.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.6.0 importlib_metadata/4.2.0 pkginfo/1.8.1 requests/2.26.0 requests-toolbelt/0.9.1 tqdm/4.62.3 CPython/3.7.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
02434dd7b9c810daa02bbbc824cc3a4c078f7d50ad76a643dd550d862d5df33d
|
|
| MD5 |
f582a5489355144f7d1bbe39f718a78b
|
|
| BLAKE2b-256 |
ac607f305d70fa9b6f59fcaf19e5bf9ab9b838c706fb195ea26852ecf57f56af
|
File details
Details for the file crikit-0.1.5-py3-none-any.whl.
File metadata
- Download URL: crikit-0.1.5-py3-none-any.whl
- Upload date:
- Size: 144.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.6.0 importlib_metadata/4.2.0 pkginfo/1.8.1 requests/2.26.0 requests-toolbelt/0.9.1 tqdm/4.62.3 CPython/3.7.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
419229b64504ec6e0471252600cff3701921d5eabea63866551655fe51886195
|
|
| MD5 |
dfe67aa92024c5308bd50339f88c7761
|
|
| BLAKE2b-256 |
4f0ca1d5e9ed8042e43c18aaff05280ba024f1c79e2d9b71722136792dc32d24
|