CStreet is a python script (python 3.6, 3.7 or 3.8) for cell state trajectory construction by using k-nearest neighbors graph algorithm for time-series single-cell RNA-seq data.
Project description
CStreet: a computed Cell State trajectory inference method for time-series single-cell RNA-seq data

| Overview | Installation | Quick Start | Parameter Details | Run CStreet in python interface | Data&Code | Citation |
Overview
CStreet is a cell state trajectory inference method for time-series single-cell RNA-seq data. It is written in Python (3.6, 3.7 or 3.8) and is available as a command line tool and a Python library to meet the needs of different users.
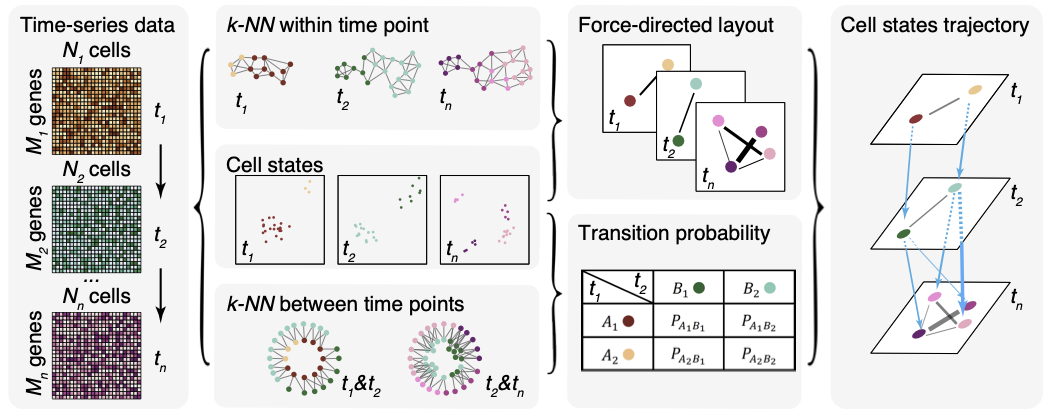
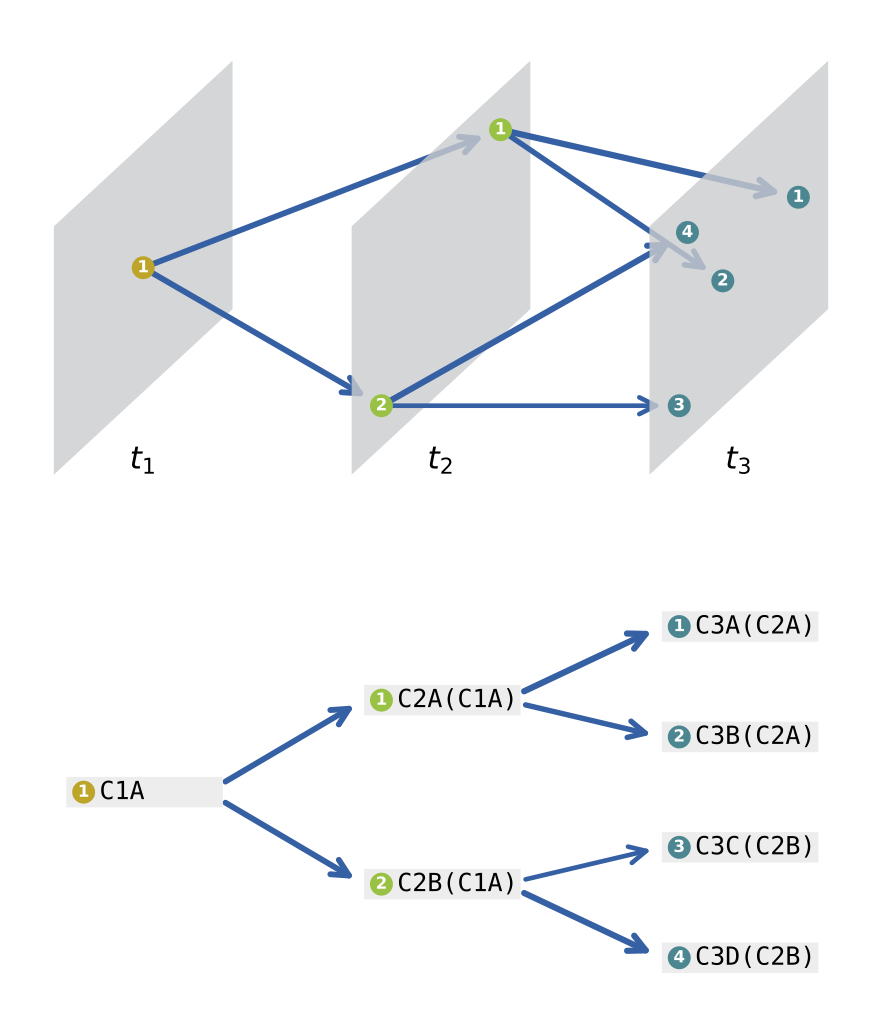
CStreet uses time-series information to construct the k-nearest neighbors connections within and between time points. Then, CStreet calculates the connection probabilities of cell states and visualizes the trajectory, which may include multiple starting points and paths, using a force-directed layout method.
Installation
CStreet has been packaged and uploaded to PyPI. Before your installation, ensure that you have pip available. pip3 is the package installer for Python. If you do not have pip3 on your machine, click here to install it. Then, CStreet and its relevant packages can be installed using a single command.
$ pip3 install cstreet
You may experience errors when installing or updating packages. This is because pip3 will change the way that it resolves dependency conflicts. We recommend you use the command below.
$ pip3 install cstreet --use-feature=2020-resolver
You may need to use the command below to add the default installation path of pip3 to your system path,
$ export PATH=~/.local/bin:$PATH
Then, type the command below to check whether CStreet has been installed successfully.(For the first time running, it may takes about one minute for CStreet’s initial configuration.)
$ CStreet -h
Quick Start
Step 1. CStreet installation following the above tutorial.
Step 2. Input preparation
CStreet utilizes time-series expression levels in tab-delimited format or AnnData format as input.
The cell state information can be generated using the built-in clustering function of CStreet or input by the user.
We provided a small test dataset containing normalized expression levels and the state information at three time points.
Or type the command below to download.
$ wget https://github.com/yw-Hua/CStreet/raw/master/test/test_data.zip
$ unzip test_data.zip
Step 3. Operation of CStreet
Type the command below to run CStreet.
$ CStreet -i test_data/ExpressionMatrix_t1.txt test_data/ExpressionMatrix_t2.txt test_data/ExpressionMatrix_t3.txt -s test_data/CellStates_t1.txt test_data/CellStates_t2.txt test_data/CellStates_t3.txt -n test
Step 4. Output
The contents of the output directory in tree format will be displayed as described below, including the clustered cell state information if it is not provided by users, the connection probabilities of the cell states and a visualization of the inferred cell state trajectories.
PATH/ProjectName
├── ProjectName_CStreetTopology.pdf
├── ProjectName_CellStatesConnCytoscape.txt
├── SupplementaryFigures
│ ├── ProjectName_t1_ForceDirectedLayout.pdf
│ ├── ProjectName_t1_LouvainUMAPClustering.pdf
│ ├── ProjectName_t1_LouvainUMAPClusteringCoordinates.txt
│ ├── ProjectName_t2_ForceDirectedLayout.pdf
│ ├── ProjectName_t2_LouvainUMAPClustering.pdf
│ ├── ProjectName_t2_LouvainUMAPClusteringCoordinates.txt
│ └── ...
└── SupplementaryResults
├── ProjectName_BetweenTimePoints_CellStatesConnectionProbabilities.txt
├── ProjectName_t1_FilteredCellInfo.txt
├── ProjectName_t1_FilteredGeneInfo.txt
├── ProjectName_t1_CellStatesConnectionProbabilities.txt
├── ProjectName_t2_FilteredCellInfo.txt
├── ProjectName_t2_FilteredGeneInfo.txt
├── ProjectName_t2_CellStatesConnectionProbabilities.txt
└── ...
Parameter Details
The parameter details of CStreet are as follows:
usage: CStreet [-h] <-i ExpMatrix1 ExpMatrix2 ExpMatrix3 ...> [-s CellStates1 CellStates2 CellStates3 ...] [-n ProjectName] [-o OutputDir] [options]
CStreet is a cell states trajectory inference method for time-series single-cell RNA-seq data.
optional arguments:
-h, --help show this help message and exit
-v, --version show version number of CStreet and exit
-i INPUT_EXPMATRIX [INPUT_EXPMATRIX ...], --Input_ExpMatrix INPUT_EXPMATRIX [INPUT_EXPMATRIX ...]
This indicates expression matrixes, which contain the
time-series expression level as read counts or
normalized values in tab-delimited format. For
example: '-i ExpressionMatrix_t1.txt
ExpressionMatrix_t2.txt ExpressionMatrix_t3.txt'
indicates the input of 3 timepoint expression
matrixes.
-T, --Input_CellinCol
This determines whether the expression level of one
cell is displayed as a column in the expression
matrixes. For example, '-T' indicates that the gene
expression levels of one cell are displayed in a
column and the expression levels of one gene across
all cells are displayed in a row.
-o OUTPUT_DIR, --Output_Dir OUTPUT_DIR
The project directory, which is used to save all
output files. DEFAULT: "./".
-n OUTPUT_NAME, --Output_Name OUTPUT_NAME
The project name, which is used to generate output
file names as a prefix. DEFAULT: "CStreet"
-s [INPUT_CELLSTATES [INPUT_CELLSTATES ...]], --Input_CellStates [INPUT_CELLSTATES [INPUT_CELLSTATES ...]]
An optional parameter that uses CStreet's built-in
dimensionality reduction and clustering methods to
perform clustering without knowing the cell states
(DEFAULT) or accepts the user’s input. The input files
should contain the cell state information sharing the
same cell ID in the expression matrixes in tab-
delimited format.
--CellClusterParam_PCAn CELLCLUSTERPARAM_PCAN
The number of principal components to use, which is
enabled only if the cell state information is not
provided. It can be set from 1 to the minimum
dimension size of the expression matrixes. DEFAULT:
10.
--CellClusterParam_k CELLCLUSTERPARAM_K
The number of nearest neighbors to be searched, which
is enabled only if the cell state information is not
provided. It should be in the range of 2 to 100 in
general. DEFAULT: 15.
--CellClusterParam_Resolution CELLCLUSTERPARAM_RESOLUTION
The resolution of the Louvain algorithm, which is
enabled only if the cell state information is not
provided. A higher resolution means that more and
smaller clusters are found. DEFAULT: 0.1.
--Switch_DeadCellFilter {ON,OFF}
The switch for the dead cell filter, which filters
cell outliers based on the count percent of
mitochondrial genes. DEFAULT: "ON".
--Threshold_MitoPercent THRESHOLD_MITOPERCENT
The maximum count percent of mitochondrial genes
needed for a cell to pass filtering, which is enabled
only if '--Switch_DeadCellFilter' is "ON". DEFAULT:
0.2.
--Switch_LowCellNumGeneFilter {ON,OFF}
The switch for the low cell number gene filter, which
retains genes that are expressed in at least a certain
number of cells. DEFAULT: "ON".
--Threshold_LowCellNum THRESHOLD_LOWCELLNUM
The minimum number of cells expressed that is required
for a gene to pass filtering, which is enabled only if
'--Switch_LowCellNumGeneFilter' is "ON". DEFAULT: 3.
--Switch_LowGeneCellsFilter {ON,OFF}
The switch for the low gene number cell filter, which
retains cells with at least a certain number of genes
expressed. DEFAULT: "ON".
--Threshold_LowGeneNum THRESHOLD_LOWGENENUM
The minimum number of genes expressed that is required
for a cell to pass filtering, which is enabled only if
'--Switch_LowGeneCellsFilter' is "ON". DEFAULT: 200.
--Switch_Normalize {ON,OFF}
The switch to enable total read count normalization
for each cell. DEFAULT: "ON".
--Threshold_NormalizeBase THRESHOLD_NORMALIZEBASE
The base of normalization, which is enabled only if '
--Switch_Normalize' is "ON". If the DEFAULT is chosen,
it is CPM normalization. DEFAULT: 1e6.
--Switch_LogTransform {ON,OFF}
The switch to logarithmize the expression matrix.
DEFAULT: "ON".
--KNNParam_metric KNNPARAM_METRIC
The distance metric to use for kNN. It can be set to
"euclidean" or "correlation". DEFAULT: "euclidean".
--WithinTimePointParam_PCAn WITHINTIMEPOINTPARAM_PCAN
The number of principal components to use within a
timepoint. It can be set from 1 to the minimum
dimension size of the expression matrixes. DEFAULT:
10.
--WithinTimePointParam_k WITHINTIMEPOINTPARAM_K
The number of nearest neighbors to be searched within
a timepoint. It should be in the range of 2 to 100 in
general. DEFAULT: 15.
--BetweenTimePointParam_PCAn BETWEENTIMEPOINTPARAM_PCAN
The number of principal components to use between
timepoints. It can be set from 1 to the minimum
dimension size of the expression matrixes. DEFAULT:
10.
--BetweenTimePointParam_k BETWEENTIMEPOINTPARAM_K
The number of nearest neighbors to be searched between
timepoints. It should be in the range of 2 to 100 in
general. DEFAULT: 15.
--ProbParam_SamplingSize PROBPARAM_SAMPLINGSIZE
The number of repeated sampling trials used to
estimate the connection probability. DEFAULT: 5.
--ProbParam_RandomSeed PROBPARAM_RANDOMSEED
The seed of repeated sampling trials used to estimate
the connection probability. DEFAULT: 0.
--FigureParam_FigureSize FIGUREPARAM_FIGURESIZE FIGUREPARAM_FIGURESIZE
Figure size of the result figure. For example: '--
FigureParam_FigureSize 6 7' indicates width is 6 and
height is 7. DEFAULT: 6 7.
--FigureParam_LabelBoxWidth FIGUREPARAM_LABELBOXWIDTH
The width of the label box in the result figure. For
example, '--FigureParam_LabelBoxWidth 10' means that
10 characters will be shown in the label box of the
resulting figure. DEFAULT: 10.
--Threshold_MinProbability THRESHOLD_MINPROBABILITY
The minimum probability of edge for each cell state
that is displayed, which will only be used for
visualization. It can be a number between 0 and 1 or
"OTSU". When OTSU is selected, it will be
automatically estimated using OTSU's Method (OTSU,
1979). DEFAULT: "OTSU".
--Threshold_MaxOutDegree THRESHOLD_MAXOUTDEGREE
The maximum number of outdegrees for each cell state
that is displayed, which will only be used for
visualization. DEFAULT: 10.
--Threshold_MinCellNumofStates THRESHOLD_MINCELLNUMOFSTATES
The minimum cell number of each cell state that is
displayed, which will only be used for visualization.
DEFAULT: 0.
Run CStreet in Python interface
CStreet can also be used step by step in the Python interface and easily integrated into custom scripts. Here is a tutorial written using Jupyter Notebook.
Supplementary Data&Code
All supplementary code and data, including cell type annotations, used to run the comparisons are available on here. The Anndata h5ad file format is used to stored single-cell expression data and additional annotations in our supplementary file.
Citation
Zhao C, Xiu W, Hua Y, Zhang N, Zhang Y. CStreet: a computed Cell State trajectory inference method for time-series single-cell RNA sequencing data. Bioinformatics (Oxford, England). 2021 Jul. DOI: 10.1093/bioinformatics/btab488.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cstreet-1.1.2.tar.gz.
File metadata
- Download URL: cstreet-1.1.2.tar.gz
- Upload date:
- Size: 26.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/50.3.0 requests-toolbelt/0.9.1 tqdm/4.50.1 CPython/3.6.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ac5ab00ca14a1544bc31fdc88c5d8fbad880f9858d9af231771fe2667d6654e6
|
|
| MD5 |
bb443cc9e0c9cdc956d863498ceeb2b1
|
|
| BLAKE2b-256 |
f9cd4c74db35a79bf79c48aedcba8a3b2a38561c93be8e8c41a199d1349f604c
|
File details
Details for the file cstreet-1.1.2-py3-none-any.whl.
File metadata
- Download URL: cstreet-1.1.2-py3-none-any.whl
- Upload date:
- Size: 22.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/50.3.0 requests-toolbelt/0.9.1 tqdm/4.50.1 CPython/3.6.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bcb4f3052beba1ba5804ca7f35becdcc98c4bc4d8af9b5ecb3279b83c59cdeb4
|
|
| MD5 |
2d749dc2d3bcc09bacba2e8eebeb4b6a
|
|
| BLAKE2b-256 |
98ccc34efd8dfd10b554db00dd6337c54f1e0e68c77a88d312efcf852f5972ba
|