Deep-learning Tool for White Matter (WM) lesions and Claustrum structure segmentation in brain magnetic resonance imaging (MRI).
Project description
deepNeuroSeg
There are two different segmentation tasks you can perform with deepNeuroSeg: White Matter Hyperintensities (WMH) and Claustrum Segmentation. To learn details about the used deep-learning models please refer to the publications [1] and [2]. As soon as the underlying models are required, they will be downloaded to your computer and placed in ~/.deepNeuroSeg folder.
White Matter Hyperintensities (WMH) Segmentation
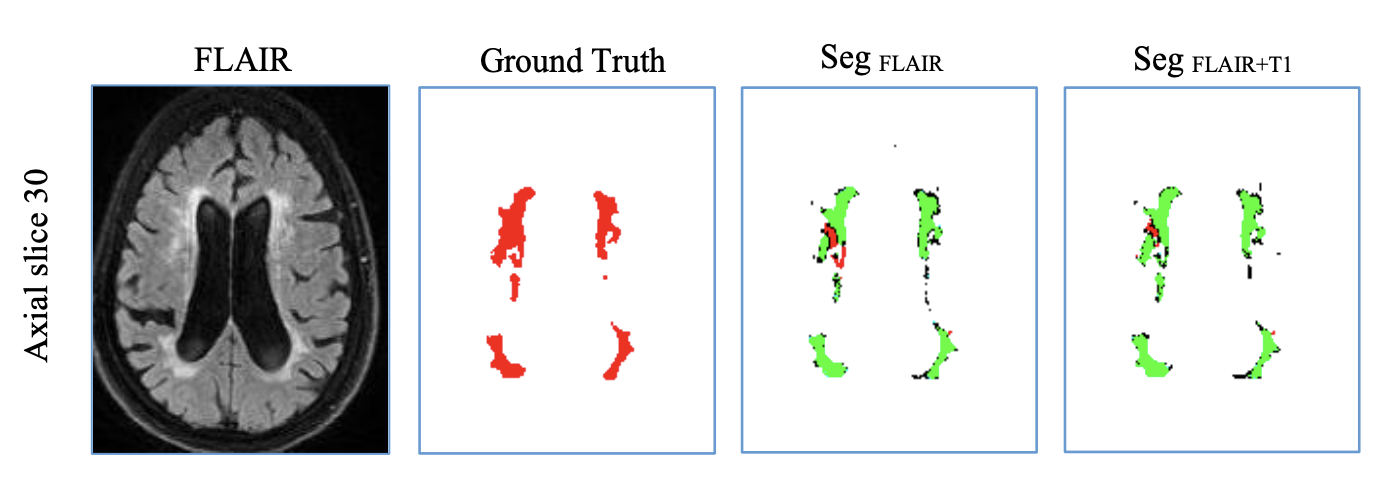
WMH segmentation can be performed either using FLAIR only or both FLAIR and T1 scans. The results would be similar to our experience [1], but the default is using both of them.
Claustrum Segmentation
For Claustrum Segmentation the T1 scan must be provided.
How to:
Run deepNeuroSeg either in command line or Python.
Command line
deepNeuroSeg performs two different segmentation tasks. The desired task must be specified with --type that is either equals to wmh (White Matter Hyperintensities) or c (Claustrum). For example:
deepNeuroSeg --type wmh --flair YOUR_PATH.nii.gz --t1 YOUR_PATH.nii.gz --o YOUR_PATH
Or:
deepNeuroSeg --type c --t1 YOUR_PATH.nii.gz --o YOUR_PATH
For more details see:
deepNeuroSeg --help
Options:
--type [wmh|c] Either 'wmh' (White Matter Hyperintensities) or 'c'
(Claustrum)
--flair PATH Path to nii.gz file with a FLAIR scan.
--t1 PATH Path to nii.gz file with a T1 scan.
--o TEXT Path where to save the resulting segmentation. Directory path or specific nii.gz file path.
[required]
--help Show this message and exit.
The resulting mask will be saved with user-specified .nii.gz file name or in the user-specified directory under the name out_mask.nii.gz.
Python
In Python user will have to follow the next steps:
Step 1: Import deepNeuroSeg
from deepNeuroSeg import SegmentationFactory, SegmentationType
Step 2: Create a SegmentationFactory object with segmentation type either SegmentationType.Claustrum or SegmentationType.WMH. An example for WMH Segmentation with both FLAIR and T1 modalities:
segmenter = SegmentationFactory.create_segmenter(SegmentationType.WMH,
FLAIR_path='YOUR_PATH',
T1_path='YOUR_PATH')
Or claustrum segmentation:
segmenter = SegmentationFactory.create_segmenter(SegmentationType.Claustrum,
T1_path='YOUR_PATH')
Step 3: Next the segmentation can be performed.
Option a: The user can specify the output path directly in perform_segmentation method.
prediction = segmenter.perform_segmentation(outputDir='YOUR_PATH')
Option b: The output numpy array can be inspected first, and then saved with save_segmentation function.
prediction = segmenter.perform_segmentation()
segmenter.save_segmentation(mask=prediction, outputDir='YOUR_PATH')
In both cases, the output mask will be saved with user-specified .nii.gz file name or in user-specified directory under the name out_mask.nii.gz.
See also the jupyter notebook for a quick start.
Special feature of Claustrum Segmentation:
The user can check the orientation of the coronal and axial images by selecting the special feature in perform_segmentation method:
prediction = segmenter.perform_segmentation(check_orientation=True)
check_orientation=True will save the images of coronal and axial slices under ~/.deepNeuroSeg/images/.
References:
[1]: Li, Hongwei, et al. "Fully convolutional network ensembles for white matter hyperintensities segmentation in MR images." NeuroImage 183 (2018): 650-665.
[2]: Li, Hongwei, et al. "Complex Grey Matter Structure Segmentation in Brains via Deep Learning: Example of the Claustrum." arXiv preprint arXiv:2008.03465 (2020).
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file deepNeuroSeg-0.4.tar.gz.
File metadata
- Download URL: deepNeuroSeg-0.4.tar.gz
- Upload date:
- Size: 13.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/4.0.1 pkginfo/1.7.0 requests/2.25.1 requests-toolbelt/0.9.1 tqdm/4.60.0 CPython/3.9.1
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1ce5ae64d0d7b21457848271144e9d2748b1f1647ceb13f838e9a6813986387d
|
|
| MD5 |
cd5dbfb9f81f53de1587aaf0a4e2130b
|
|
| BLAKE2b-256 |
75485481db4865a2102503bdd2a778a34cc20efa3082a6dc3050d896b9e596ad
|











