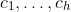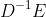TensorFlow 2 implementation of Deep Graph Convolutional Neural Networks.
Project description
DGCNN [TensorFlow]
TensorFlow 2 implementation of An end-to-end deep learning architecture for graph classification based on work by M. Zhang et al., 2018.
Moreover, we offer an attention based modification of the above by utilising graph attention (Veličković et al., 2017) to learn edge weights.
Installation
Simply run pip install dgcnn. The only dependency is tensorflow>=2.0.0.
Usage
The core data structure is the graph signal. If we have N nodes in a graph each having C observed features then the graph signal is the tensor with shape (batch, N, C) corresponding to the data produced by all nodes. Often we have sequences of graph signals in a time series. We will call them temporal graph signals and assume a shape of (batch, time steps, N, C). For each graph signal we also need to have the corresponding adjacency matrices of shape (batch, N, N) or (batch, timesteps, N, N) for temporal and non-temporal data, respectively. While DGCNNs can operate on graphs with different node-counts, C should always be the same and each batch should only contain graphs with the same number of nodes.
The DeepGraphConvolution Layer
This adaptable layer contains the whole DGCNN architecture and operates on both temporal and non-temporal data. It takes the graph signals and their corresponding adjacency matrices and performs the following steps (as described in the paper):
We initialize the layer by providing 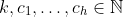
-
It iteratively applies
GraphConvolutionlayers h times with variable hidden feature dimensions.
-
After that, it concatenates all the outputs of the graph convolutions into one tensor which has the shape (..., N,
).
-
Finally it applies
SortPoolingas described in the paper to obtain the output tensor of shape (..., k,).
Import this layer with from gdcnn.components import DeepGraphConvolution.
Initiated it with the following parameters:
Thus, if we have non-temporal graph signals with 10 nodes and 5 features each and we would like to apply a DGCNN containing 3 graph convolutions with hidden feature dimensions of 10, 5 and 2 and SortPooling that keeps the 5 most relevant nodes. Then we would run
from dgcnn.components import DeepGraphConvolution
from tensorflow.keras.layers import Input
from tensorflow.keras import Model
# generating random graph signals as test data
graph_signal = np.random.normal(size=(100, 10, 5)
# corresponding fully connected adjacency matrices
adjacency = np.ones((100, 10, 10))
# inputs to the DGCNN
X = Input(shape=(10, 5), name="graph_signal")
E = Input(shape=(10, 10), name="adjacency")
# DGCNN
# Note that we pass the signals and adjacencies as a tuple.
# The graph signal always goes first!
output = DeepGraphConvolution([10, 5, 2], k=5 )((X, E))
# defining model
model = Model(inputs=[X, E], outputs=output)
Further layers and features
The documentation contains information on how to use the internal SortPooling, GraphConvolution and AttentionMechanism layers and also describes more optional parameters like regularisers, initialisers and constrains that can be used.
Contribute
Bug reports, fixes and additional features are always welcome! Make sure to run the tests with python setup.py test and write your own for new features. Thanks.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file dgcnn-0.3.2.tar.gz.
File metadata
- Download URL: dgcnn-0.3.2.tar.gz
- Upload date:
- Size: 21.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/2.0.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/42.0.2.post20191203 requests-toolbelt/0.9.1 tqdm/4.40.2 CPython/3.7.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7ff8a3bca17c2e4e4f8ae89413db39ac7b811e1576e9e81199813b039405b5c4
|
|
| MD5 |
39d365107d1e11e3124307c60663bbbd
|
|
| BLAKE2b-256 |
eed8b0ae1cb02e323d2619c9e358db98700ee5f3cc8a306318bbcc72d486d6f1
|