dleamse's encoding and embedding methods, and dleamse's faiss index (IndexIDMap type) write.
Project description
DLEAMSE

A Deep LEArning-based Mass Spectra Embedder for spectral similarity scoring. DLEAMSE (based on Siamese Network) is trained and tested with a larger dataset from PRIDE Cluster. The repository stores the encoder and embedder scripts of DLEAMSE to encode and embed spectra.
The following repo presented the model DLEAMSE and the tool mslookup.
Training data set
A larger spectral set from PRIDE Cluster is used to construct the training and test data, which use high confidence spectra retrieved from high consistency clusters. We chose PRIDE Cluster data to train and test DLEAMSE, for two reasons: 1. The spectra in high consistency clusters are high confidence spectra. 2. The spectral set from PRIDE Cluster covers more species and instrument types. Two filters were used for retrieving high confidence spectra. The first filter controls the quality of collected clusters. We customized clustering-file-converter (https://github.com/spectra-cluster/clustering-file-converter) to retain the high-quality spectral clusters (cluster size >= 30, cluster ratio >= 0.8, and the total ions current (TIC) >= 0.2). The second filter eliminates duplicate clusters assigned with same peptide sequence, only one in the dupli-cates has been chosen, to ensure that the retained clusters are from different peptides. Then 113,362 clusters have been retrained from PRIDE Cluster release 201504. The needed spectra in clusters are acquired from the PRIDE Archive.
Model and Training
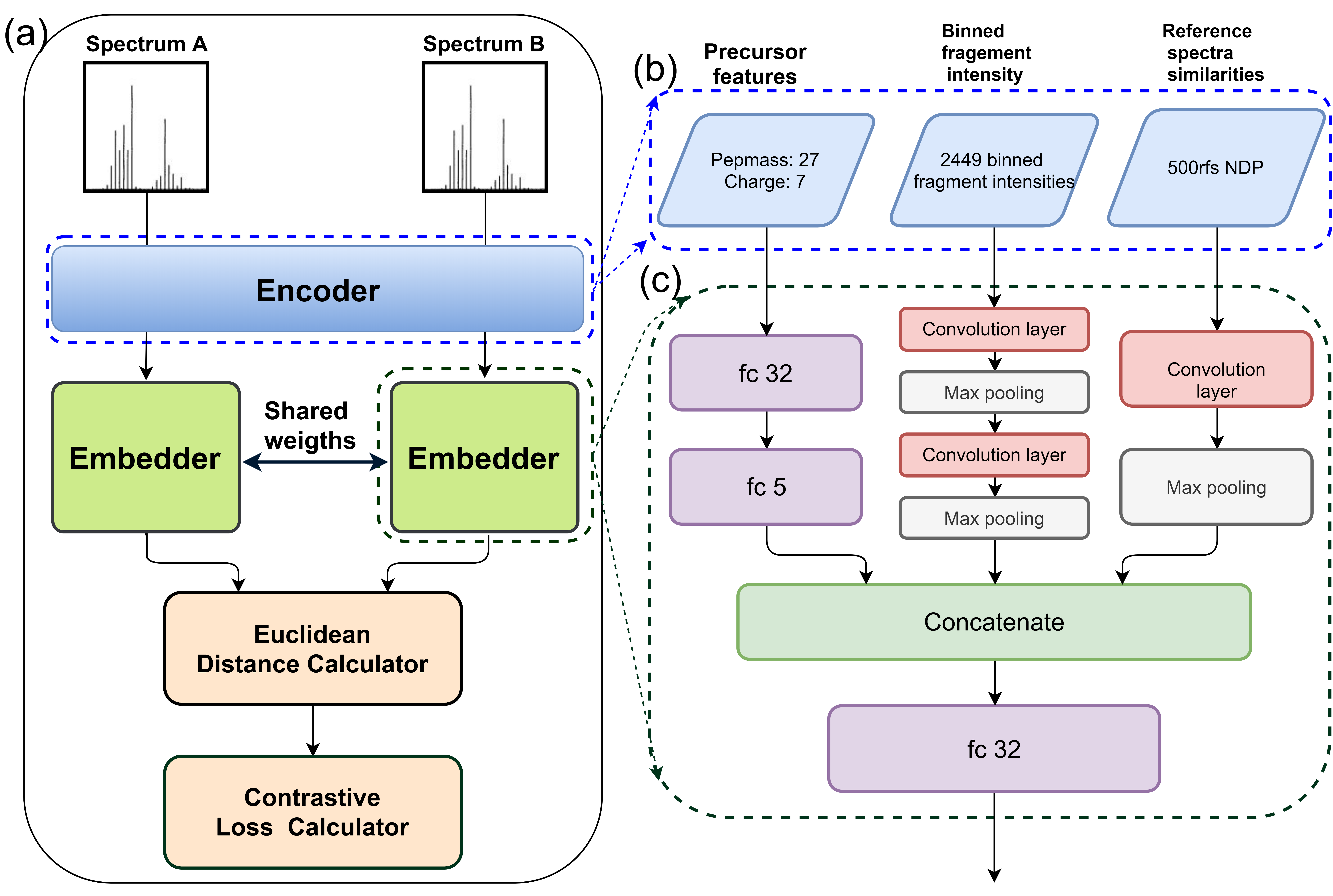
In DLEAMSE, Siamese network (Figure 1a) trains two same embedding models (Figure 1c) with shared weights, and spectra are encoded by the same encoder (Figure 1b) before the embedding. Based on the Euclidean distance between the pair of embedded spectra, the weights of embedding model is learned by contrastive loss function adapted from Hadsell et. al. that penalizes far-apart same-label spectra (label=1) and nearby different-label spectra (label=0). Back propagation from the loss function is used to update the weights in the network. The net-work is trained by stochastic gradient descent with the Adam update rule with a learning rate of 0.005. The codes are implemented in Python3 with the PyTorch framework.
Testing
Requirements
- Python3.7 (or Anaconda3)
- torch==1.0.0 (python -m pip install torch===1.0.0 torchvision===0.2.1 -f https://download.pytorch.org/whl/torch_stable.html)
- pyteomics>=3.5.1
- numpy>=1.13.3
- numba>=0.45.0
- faiss-cpu (conda install faiss-cpu pytorch -c)
- more_itertools==7.1.0
Installation
DLEAMSE’s encoder and embedder have been packaged and uploaded to pypi library, the package’s name is dleamse.
python -m pip install dleamse
Usage
The model file of DLEAMSE: 080802_20_1000_NM500R_model.pkl The 500 reference spectra used in our project: 500_rfs_spectra.mgf
mslookup.py: the commandline script of dleamse
MSLOOKUP
The mslookup is a tool developed using the DLEAMSE model and algorithm and faiss database to encode, index and search previously identified/unidentified spectra in public repositories.
Encode and Embed spectra
python mslookup.py embed-ms-file -i test_cml_index/PXD003552_61576_ArchiveSpectrum.json
Create index files
python mslookup.py make-index -d test_cml_index/database_ids_usi.csv -e test_cml_index/ -o test_cml_index/test_cml_0412.index
Merge index files
python mslookup.py merge-indexes test_cml_index/*.index test_cml_index/test_cml_merge_0412.index
Range Search
In this case, lower_threshold and upper_threshold of range searching are default values, lower_threshold(-lt)=0, upper_threshold(-ut)=0.07.
python mslookup.py range-search -i test_cml_index/test_cml_0412.index -u test_cml_index/test_cml_0412_ids_usi.csv -e test_cml_index/*_embedded.txt -o test_cml_index/test_cml_rangesearch_rlt.json
In this case, lower_threshold(-lt)=0.01, and upper_threshold(-ut) is set to default value 0.07.
python mslookup.py range-search -i test_cml_index/test_cml_0412.index -u test_cml_index/test_cml_0412_ids_usi.csv -e test_cml_index/*_embedded.txt -lt 0.01 -o test_cml_index/test_cml_rangesearch_rlt.json
In this case, lower_threshold(-lt)=0.01, and upper_threshold(-ut) = 0.05.
python mslookup.py range-search -i test_cml_index/test_cml_0412.index -u test_cml_index/test_cml_0412_ids_usi.csv -e test_cml_index/*_embedded.txt -lt 0.01 -ut 0.05 -o test_cml_index/test_cml_rangesearch_rlt.json
About index search
dleamse_faiss_index_search.py
Range Search query 32D spectra vectors against spectra library's index file, Default lower_threshold is 0 and upper_threshold is 0.07.
Databases
We have released a couple of databases for the users of the mslookup tool ftp://ftp.pride.ebi.ac.uk/pride/data/proteogenomics/projects/mslookup/. Databases can be download from the FTP and use locally in your own computer.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dleamse-0.3.6.tar.gz.
File metadata
- Download URL: dleamse-0.3.6.tar.gz
- Upload date:
- Size: 18.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/47.1.0 requests-toolbelt/0.9.1 tqdm/4.49.0 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
896a70876a917fc219b529933754ba564db81a26c10498f5ef74e3948f1ccd57
|
|
| MD5 |
7cc9e8432d980a53e9dcff0141609362
|
|
| BLAKE2b-256 |
b43a5c67073af1f0ac947db6d4859c3af075606eaae5b028c4d4260a6a47c189
|
File details
Details for the file dleamse-0.3.6-py3-none-any.whl.
File metadata
- Download URL: dleamse-0.3.6-py3-none-any.whl
- Upload date:
- Size: 24.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/47.1.0 requests-toolbelt/0.9.1 tqdm/4.49.0 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
98fed217cac9c87bc355cfc73d2ab6b047de9d48111cf856fc3c158304988ff8
|
|
| MD5 |
1cfa7126daefcdd8167b20bb2b02bd93
|
|
| BLAKE2b-256 |
60dd585cae3a6d9d01f897d44996d9d8604ce5a408786cb9b2a58067ea52bb32
|