A program to model disorder in small-molecule X-ray structures
Project description

DSR
The program DSR consists of a text database with fragments of molecules and the DSR program. It acts as a preprocessor for SHELXL .res files. The user inserts a special command in the SHELXL .res file and the DSR program reads this information to put a molecule or fragment with the desired atoms on the position of the target atoms or q-peaks in the unit cell. Bond restraints can be either applied from the database to the molecule or automatically generated.
I apologize for the messy code. This was my first bigger project...
You have either a command line version:
C:\Users\daniel>dsr
----------------------------------------------------- D S R - v238 -------------------------------------
Disordered Structure Refinement (DSR)
Example DSR .res file command line:
REM DSR PUT/REPLACE "Fragment" WITH C1 C2 C3 ON Q1 Q2 Q3 PART 1 OCC -21 =
RESI DFIX
---------------------------------------------------------------------------------------------------------
PUT: Just put the fragment source atoms here.
REPLACE: Replace atoms of PART 0 in 1.3 A distance around target atoms.
---------------------------------------------------------------------------------------------------------
optional arguments:
-h, --help show this help message and exit
-r "res file" ["res file" ...]
res file with DSR command
-re "res file" ["res file" ...]
res file with DSR command (write restraints to external file)
-e "fragment" export fragment as .res/.png file
-c "fragment" export fragment to clipboard
-t inverts the current fragment
-i "tgz file" import a fragment from GRADE (needs .tgz file)
-l list names of all database entries
-s "string" search the database for a name
-g keep group rigid (no restraints)
-u Update DSR to the most current version
-n do not refine after fragment transfer
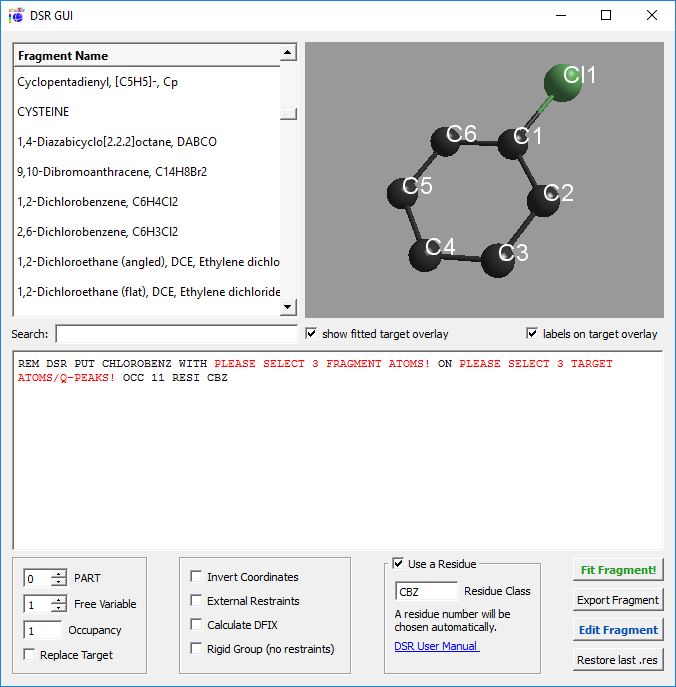
Or a graphical user interface in ShelXle:
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dsr_shelx-242.tar.gz.
File metadata
- Download URL: dsr_shelx-242.tar.gz
- Upload date:
- Size: 1.8 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7d4acea62ebc6e75635f85fec13bbb950b2521050ff74ee1d1a8dfd106e26a44
|
|
| MD5 |
50b16bdb67b9c1e1c3530a130de7976e
|
|
| BLAKE2b-256 |
639be9adf756c05b2c8beb97435cde19946d6fb9b6094ca0c5b76e57b06931bf
|
File details
Details for the file dsr_shelx-242-py3-none-any.whl.
File metadata
- Download URL: dsr_shelx-242-py3-none-any.whl
- Upload date:
- Size: 1.5 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4d6cd0f9baa32cc20acc0e0fdd05971a9936c8ca8aa8a9532ec3b1b00367c036
|
|
| MD5 |
c1eaf1991dc1af174653e23351c1b009
|
|
| BLAKE2b-256 |
e38ee01e5354fda95a3f7af3a55f9700bc5471773db33f40aa4b00efec2cc0e9
|