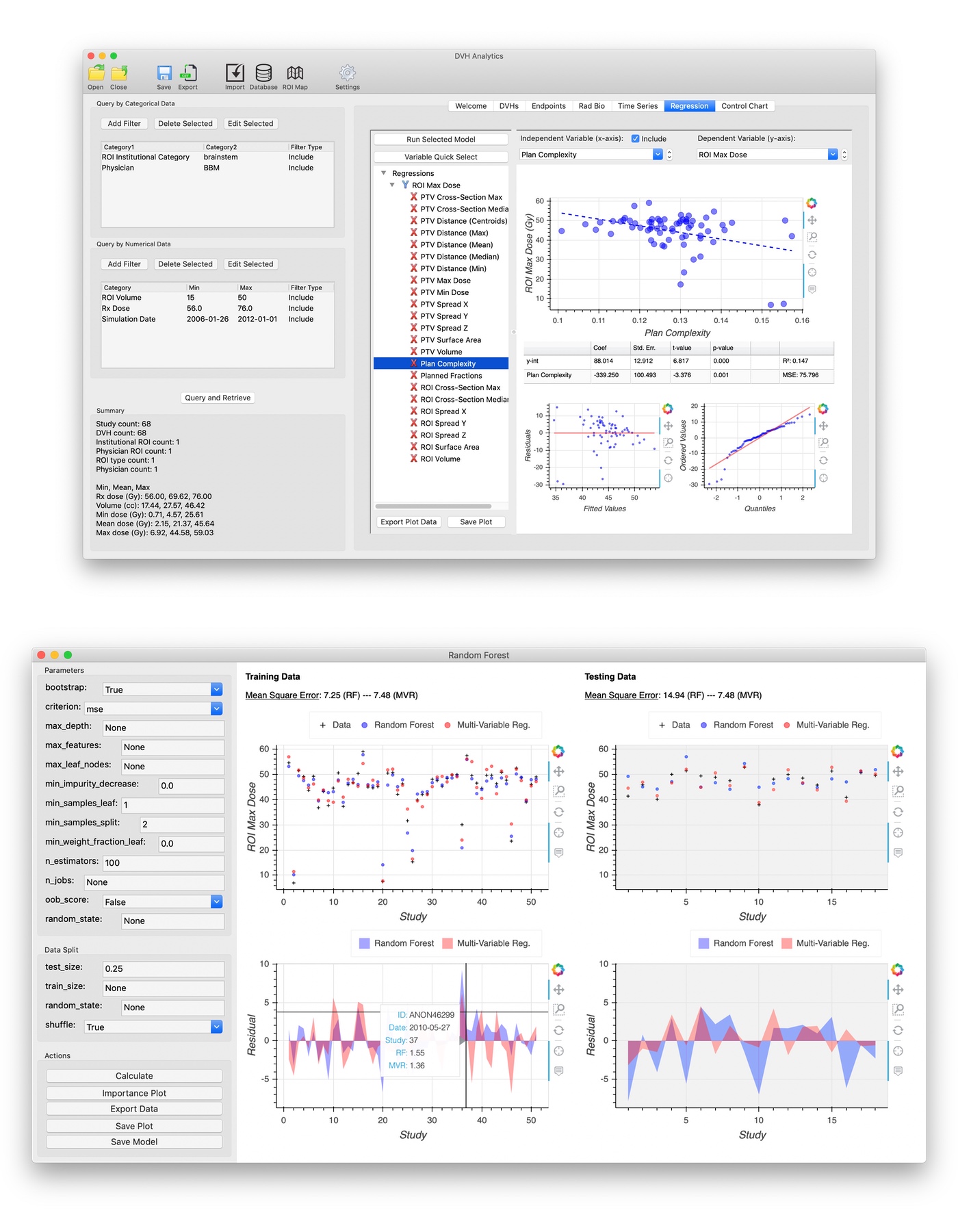
Create a database of DVHs, GUI with wxPython, plots with Bokeh
Project description
DVH Analytics (DVHA) is a software application for building a local database of radiation oncology treatment planning data. It imports data from DICOM-RT files (i.e., plan, dose, and structure), creates a SQL database, provides customizable plots, and provides tools for generating linear, multi-variable, and machine learning regressions.
Documentation
Be sure to check out the latest release for the user manual PDF, which is geared towards the user interface. For power-users, dvha.readthedocs.io contains detailed documentation for backend tools (e.g., if you want to perform queries with python commands).
Executables
Executable versions of DVHA can be found here. Please keep in mind this software is still in beta. If you have issues, compiling from source may be more informative.
About
In addition to viewing DVH data, this software provides methods to:
download queried data
create time-series plots of various planning and dosimetric variables
calculate correlations
generate multi-variable linear and machine learning regressions
share regression models with other DVHA users
additional screenshots available here
The code is built with these core libraries:
wxPython Phoenix - Build a native GUI on Windows, Mac, or Unix systems
Pydicom - Read, modify and write DICOM files with python code
dicompyler-core - A library of core radiation therapy modules for DICOM RT
Bokeh - Interactive Web Plotting for Python
scikit-learn - Machine Learning in Python
Installation
To install via pip:
$ pip install dvhaIf you’ve installed via pip or setup.py, launch from your terminal with:
$ dvhaIf you’ve cloned the project, but did not run the setup.py installer, launch DVHA with:
$ python dvha_app.pySee our installation notes for potential Shapely install issues on MS Windows and help setting up a PostgreSQL database if it is preferred over SQLite3.
Dependencies
Python >3.5
wxPython Phoenix >= 4.0.4, < 4.1.0
Pydicom >=1.4.0
dicompyler-core >= 0.5.4
Bokeh >= 1.2.0, < 2.0.0
PostgreSQL (optional) and psycopg2
Shapely < 1.7.0
Statsmodels >=0.8.0
Scikit-learn >= 0.21.0
Support
If you like DVHA and would like to support our mission, all we ask is that you cite us if we helped your publication, or help the DVHA community by submitting bugs, issues, feature requests, or solutions on the issues page.
Cite
- DOI: https://doi.org/10.1002/acm2.12401
Cutright D, Gopalakrishnan M, Roy A, Panchal A, and Mittal BB. “DVH Analytics: A DVH database for clinicians and researchers.” Journal of Applied Clinical Medical Physics 19.5 (2018): 413-427.
The previous web-based version described in the above publication can be found here but is no longer being developed.
Selected Studies Using DVHA
5,000 Patients National Cancer Institute (5R01CA219013-03): Active 8/1/17 → 7/31/22 Retrospective NCI Phantom-Monte Carlo Dosimetry for Late Effects in Wilms Tumor Brannigan R (Co-Investigator), Kalapurakal J (PD/PI), Kazer R (Co-Investigator)
265 Patients DOI: https://doi.org/10.1016/j.ijrobp.2019.06.2509 Gross J, et al. “Determining the organ at risk for lymphedema after regional nodal irradiation in breast cancer.” International Journal of Radiation Oncology* Biology* Physics 105.3 (2019): 649-658.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dvha-0.9.7.tar.gz.
File metadata
- Download URL: dvha-0.9.7.tar.gz
- Upload date:
- Size: 1.2 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.6.1 requests/2.25.0 setuptools/51.0.0 requests-toolbelt/0.9.1 tqdm/4.54.1 CPython/3.6.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3c7234db1634b35d29ac47903a3aae51e08bca95d386d120ab1571fdce82d3cb
|
|
| MD5 |
c567f404ded10c72a2b0132ba8813a8f
|
|
| BLAKE2b-256 |
aeda879cce0f9e3419f98ca3f1da7cb01f4d074c79a5db0083d34984cfa76071
|
File details
Details for the file dvha-0.9.7-py3-none-any.whl.
File metadata
- Download URL: dvha-0.9.7-py3-none-any.whl
- Upload date:
- Size: 1.2 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.6.1 requests/2.25.0 setuptools/51.0.0 requests-toolbelt/0.9.1 tqdm/4.54.1 CPython/3.6.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a0ec3db51e587538071a6c4cea182745501eaf9f78cac86432010df717233aed
|
|
| MD5 |
82788f4bb31770026a9800c0236033be
|
|
| BLAKE2b-256 |
7c22b0afeff9e6807f4077979d992baef99e8295e94913254f32ae0957d9bd9e
|