Compare one dataset to another at a variety of p-value cutoffs
Project description
A simple script to compare p-values between a test and comparison dataset at a variety of p-value cutoffs. By plotting the enrichment score at a variety of cutoffs, it is possible to pick the optimal cutoff for your data.
Version: 1.0-beta2
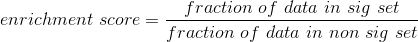
Algorithm
For each p-value in the interval between max_pval (default: 0.05) and min_pval (default: 1e-15), we test at intervals of 1 and 5 for each order of magnitute, e.g. 0.05, 0.01, 0.005, 0.001, 5e-4, 1e-4, 5e-5, 1e-6, … 1e-15.
To test, we simply take all identities with a p-value less than the cutoff and compare them to all identities in the comparison set with p-values below the comp_set_pvalue. We simply ask what percentage or the test set are in the comparison set. We then do exactly the same with the entire set of identities in the comparison set that have a p-value greater than 0.98.
The identities are generally going to be gene or SNP names, but they can be anything (e.g. coordinates) as long as they overlap in the test and comparison data.
Installation
Install via PyPI:
pip install enrich_pvaluesOr install from github:
pip install https://github.com/TheFraserLab/enrich_pvalues/tarball/masterIt should work with python 2 or 3, but python 3 is recommended.
Requirements
In requirements.txt, we use numpy, pandas, matplotlib, seaborn, tabulate, and tqdm.
Usage
First, dump a configuration file to describe your data:
enrich_pvalues dump-config enrich_atac.jsonThis will also print a help table describing each option. You need to describe your comparison data and your test data, and pick your p-value thresholds.
Next, split your comparison dataset into two tables: significant, and not-significant:
enrich_pvalues split -c enrich_atac.json --prefix atac /path/to/comp_data.txt.gzNow, run the enrichment using those two tables and your test data:
enrich_pvalues run -c enrich_atac.json -o atac_scores.xls -p atac /path/to/test_data.txtNote, the second to last argument is the prefix from the second step.
Finally, plot the data. This can also be done by passing e.g. --plot myplot.png to the run step.
enrich_pvalues plot --prefix caQTL atac_scores.xls atac_plot.pdfNote: the scores can be excel format, pickled format, or text format, depending on the suffix. Also, the prefix in this plot step is different, it is used to title the plot only, and so can be whatever you want.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file enrich_pvalues-1.0b2.tar.gz.
File metadata
- Download URL: enrich_pvalues-1.0b2.tar.gz
- Upload date:
- Size: 10.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8c475ea08ccce7299e6fa152784895d3fd2e781d53bd17275102ce9ce2fafeef
|
|
| MD5 |
76895d80ed73ea5988131ead86775182
|
|
| BLAKE2b-256 |
faa36c6c9eaeb6cdee70e4ef5ab60e12343ba042d10647744dddbf9dce7918fb
|










