Frequently used commands in bioinformatics
Project description





Introduction
The main goal of the fuc package (pronounced “eff-you-see”) is to wrap some of the most frequently used commands in the field of bioinformatics into one place.
The package is written in Python, and supports both command line interface (CLI) and application programming interface (API) whose documentations are available at the Read the Docs.
Currently, fuc can be used to analyze, summarize, visualize, and manipulate the following file formats:
Sequence Alignment/Map (SAM)
Binary Alignment/Map (BAM)
CRAM
Variant Call Format (VCF)
Mutation Annotation Format (MAF)
Browser Extensible Data (BED)
FASTQ
FASTA
General Feature Format (GFF)
Gene Transfer Format (GTF)
delimiter-separated values format (e.g. comma-separated values or CSV format)
Additionally, fuc can be used to parse output data from the following programs:
Your contributions (e.g. feature ideas, pull requests) are most welcome.
Citation
If you use fuc in a published analysis, please report the program version and cite the following article:
Lee et al., 2022. ClinPharmSeq: A targeted sequencing panel for clinical pharmacogenetics implementation. PLOS ONE.
Support fuc
If you find my work useful, please consider becoming a sponsor.
Installation
The following packages are required to run fuc:
biopython lxml matplotlib matplotlib-venn numpy pandas pyranges pysam scipy seaborn statsmodels
There are various ways you can install fuc. The recommended way is via conda (Anaconda):
$ conda install -c bioconda fucAbove will automatically download and install all the dependencies as well. Alternatively, you can use pip (PyPI) to install fuc and all of its dependencies:
$ pip install fucFinally, you can clone the GitHub repository and then install fuc locally:
$ git clone https://github.com/sbslee/fuc
$ cd fuc
$ pip install .The nice thing about this approach is that you will have access to development versions that are not available in Anaconda or PyPI. For example, you can access a development branch with the git checkout command. When you do this, please make sure your environment already has all the dependencies installed.
Getting help
For detailed documentations on the fuc package’s CLI and API, please refer to the Read the Docs.
For getting help on the fuc CLI:
$ fuc -h
usage: fuc [-h] [-v] COMMAND ...
positional arguments:
COMMAND
bam-aldepth Compute allelic depth from a BAM file.
bam-depth Compute per-base read depth from BAM files.
bam-head Print the header of a BAM file.
bam-index Index a BAM file.
bam-rename Rename the sample in a BAM file.
bam-slice Slice a BAM file.
bed-intxn Find the intersection of BED files.
bed-sum Summarize a BED file.
cov-concat Concatenate depth of coverage files.
cov-rename Rename the samples in a depth of coverage file.
fa-filter Filter sequence records in a FASTA file.
fq-count Count sequence reads in FASTQ files.
fq-sum Summarize a FASTQ file.
fuc-bgzip Write a BGZF compressed file.
fuc-compf Compare the contents of two files.
fuc-demux Parse the Reports directory from bcl2fastq.
fuc-exist Check whether certain files exist.
fuc-find Retrieve absolute paths of files whose name matches a
specified pattern, optionally recursively.
fuc-undetm Compute top unknown barcodes using undertermined FASTQ from
bcl2fastq.
maf-maf2vcf Convert a MAF file to a VCF file.
maf-oncoplt Create an oncoplot with a MAF file.
maf-sumplt Create a summary plot with a MAF file.
maf-vcf2maf Convert a VCF file to a MAF file.
ngs-bam2fq Pipeline for converting BAM files to FASTQ files.
ngs-fq2bam Pipeline for converting FASTQ files to analysis-ready BAM
files.
ngs-hc Pipeline for germline short variant discovery.
ngs-m2 Pipeline for somatic short variant discovery.
ngs-pon Pipeline for constructing a panel of normals (PoN).
ngs-quant Pipeline for running RNAseq quantification from FASTQ files
with Kallisto.
ngs-trim Pipeline for trimming adapters from FASTQ files.
tabix-index Index a GFF/BED/SAM/VCF file with Tabix.
tabix-slice Slice a GFF/BED/SAM/VCF file with Tabix.
tbl-merge Merge two table files.
tbl-sum Summarize a table file.
vcf-call Call SNVs and indels from BAM files.
vcf-filter Filter a VCF file.
vcf-index Index a VCF file.
vcf-merge Merge two or more VCF files.
vcf-rename Rename the samples in a VCF file.
vcf-slice Slice a VCF file for specified regions.
vcf-split Split a VCF file by individual.
vcf-vcf2bed Convert a VCF file to a BED file.
vcf-vep Filter a VCF file by annotations from Ensembl VEP.
optional arguments:
-h, --help Show this help message and exit.
-v, --version Show the version number and exit.For getting help on a specific command (e.g. vcf-merge):
$ fuc vcf-merge -hBelow is the list of submodules available in the fuc API:
common : The common submodule is used by other fuc submodules such as pyvcf and pybed. It also provides many day-to-day actions used in the field of bioinformatics.
pybam : The pybam submodule is designed for working with sequence alignment files (SAM/BAM/CRAM). It essentially wraps the pysam package to allow fast computation and easy manipulation. If you are mainly interested in working with depth of coverage data, please check out the pycov submodule which is specifically designed for the task.
pybed : The pybed submodule is designed for working with BED files. It implements pybed.BedFrame which stores BED data as pandas.DataFrame via the pyranges package to allow fast computation and easy manipulation. The submodule strictly adheres to the standard BED specification.
pychip : The pychip submodule is designed for working with annotation or manifest files from the Axiom (Thermo Fisher Scientific) and Infinium (Illumina) array platforms.
pycov : The pycov submodule is designed for working with depth of coverage data from sequence alingment files (SAM/BAM/CRAM). It implements pycov.CovFrame which stores read depth data as pandas.DataFrame via the pysam package to allow fast computation and easy manipulation. The pycov.CovFrame class also contains many useful plotting methods such as CovFrame.plot_region and CovFrame.plot_uniformity.
pyfq : The pyfq submodule is designed for working with FASTQ files. It implements pyfq.FqFrame which stores FASTQ data as pandas.DataFrame to allow fast computation and easy manipulation.
pygff : The pygff submodule is designed for working with GFF/GTF files. It implements pygff.GffFrame which stores GFF/GTF data as pandas.DataFrame to allow fast computation and easy manipulation. The submodule strictly adheres to the standard GFF specification.
pykallisto : The pykallisto submodule is designed for working with RNAseq quantification data from Kallisto. It implements pykallisto.KallistoFrame which stores Kallisto’s output data as pandas.DataFrame to allow fast computation and easy manipulation. The pykallisto.KallistoFrame class also contains many useful plotting methods such as KallistoFrame.plot_differential_abundance.
pymaf : The pymaf submodule is designed for working with MAF files. It implements pymaf.MafFrame which stores MAF data as pandas.DataFrame to allow fast computation and easy manipulation. The pymaf.MafFrame class also contains many useful plotting methods such as MafFrame.plot_oncoplot and MafFrame.plot_summary. The submodule strictly adheres to the standard MAF specification.
pysnpeff : The pysnpeff submodule is designed for parsing VCF annotation data from the SnpEff program. It should be used with pyvcf.VcfFrame.
pyvcf : The pyvcf submodule is designed for working with VCF files. It implements pyvcf.VcfFrame which stores VCF data as pandas.DataFrame to allow fast computation and easy manipulation. The pyvcf.VcfFrame class also contains many useful plotting methods such as VcfFrame.plot_comparison and VcfFrame.plot_tmb. The submodule strictly adheres to the standard VCF specification.
pyvep : The pyvep submodule is designed for parsing VCF annotation data from the Ensembl VEP program. It should be used with pyvcf.VcfFrame.
For getting help on a specific submodule (e.g. pyvcf):
>>> from fuc import pyvcf
>>> help(pyvcf)In Jupyter Notebook and Lab, you can see the documentation for a python function by hitting SHIFT + TAB. Hit it twice to expand the view.
CLI examples
SAM/BAM/CRAM
To print the header of a SAM file:
$ fuc bam-head in.samTo index a CRAM file:
$ fuc bam-index in.cramTo rename the samples in a SAM file:
$ fuc bam-rename in.sam NA12878 > out.samTo slice a BAM file:
$ fuc bam-slice in.bam chr1:100-200 > out.bamBED
To find intersection between BED files:
$ fuc bed-intxn 1.bed 2.bed 3.bed > intersect.bedFASTQ
To count sequence reads in a FASTQ file:
$ fuc fq-count example.fastqFUC
To check whether a file exists in the operating system:
$ fuc fuc-exist example.txtTo find all VCF files within the current directory recursively:
$ fuc fuc-find .vcf.gzTABLE
To merge two tab-delimited files:
$ fuc tbl-merge left.tsv right.tsv > merged.tsvVCF
To merge VCF files:
$ fuc vcf-merge 1.vcf 2.vcf 3.vcf > merged.vcfTo filter a VCF file annotated by Ensembl VEP:
$ fuc vcf-vep in.vcf 'SYMBOL == "TP53"' > out.vcfAPI examples
BAM
To create read depth profile of a region from a CRAM file:
>>> from fuc import pycov
>>> cf = pycov.CovFrame.from_file('HG00525.final.cram', zero=True,
... region='chr12:21161194-21239796', names=['HG00525'])
>>> cf.plot_region('chr12:21161194-21239796')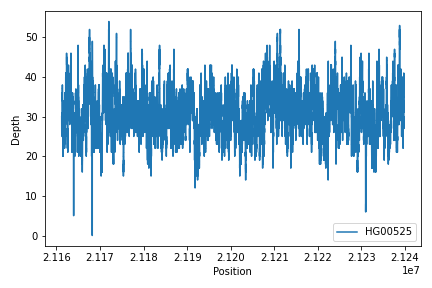
VCF
To filter a VCF file based on a BED file:
>>> from fuc import pyvcf
>>> vf = pyvcf.VcfFrame.from_file('original.vcf')
>>> filtered_vf = vf.filter_bed('targets.bed')
>>> filtered_vf.to_file('filtered.vcf')To remove indels from a VCF file:
>>> from fuc import pyvcf
>>> vf = pyvcf.VcfFrame.from_file('with_indels.vcf')
>>> filtered_vf = vf.filter_indel()
>>> filtered_vf.to_file('no_indels.vcf')To create a Venn diagram showing genotype concordance between groups:
>>> from fuc import pyvcf, common
>>> common.load_dataset('pyvcf')
>>> f = '~/fuc-data/pyvcf/plot_comparison.vcf'
>>> vf = pyvcf.VcfFrame.from_file(f)
>>> a = ['Steven_A', 'John_A', 'Sara_A']
>>> b = ['Steven_B', 'John_B', 'Sara_B']
>>> c = ['Steven_C', 'John_C', 'Sara_C']
>>> vf.plot_comparison(a, b, c)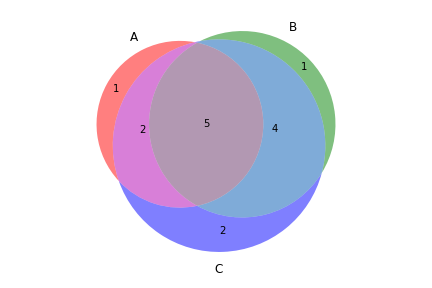
To create various figures for normal-tumor analysis:
>>> import matplotlib.pyplot as plt
>>> from fuc import common, pyvcf
>>> common.load_dataset('pyvcf')
>>> vf = pyvcf.VcfFrame.from_file('~/fuc-data/pyvcf/normal-tumor.vcf')
>>> af = pyvcf.AnnFrame.from_file('~/fuc-data/pyvcf/normal-tumor-annot.tsv', sample_col='Sample')
>>> normal = af.df[af.df.Tissue == 'Normal'].index
>>> tumor = af.df[af.df.Tissue == 'Tumor'].index
>>> fig, [[ax1, ax2], [ax3, ax4]] = plt.subplots(2, 2, figsize=(10, 10))
>>> vf.plot_tmb(ax=ax1)
>>> vf.plot_tmb(ax=ax2, af=af, group_col='Tissue')
>>> vf.plot_hist_format('#DP', ax=ax3, af=af, group_col='Tissue')
>>> vf.plot_regplot(normal, tumor, ax=ax4)
>>> plt.tight_layout()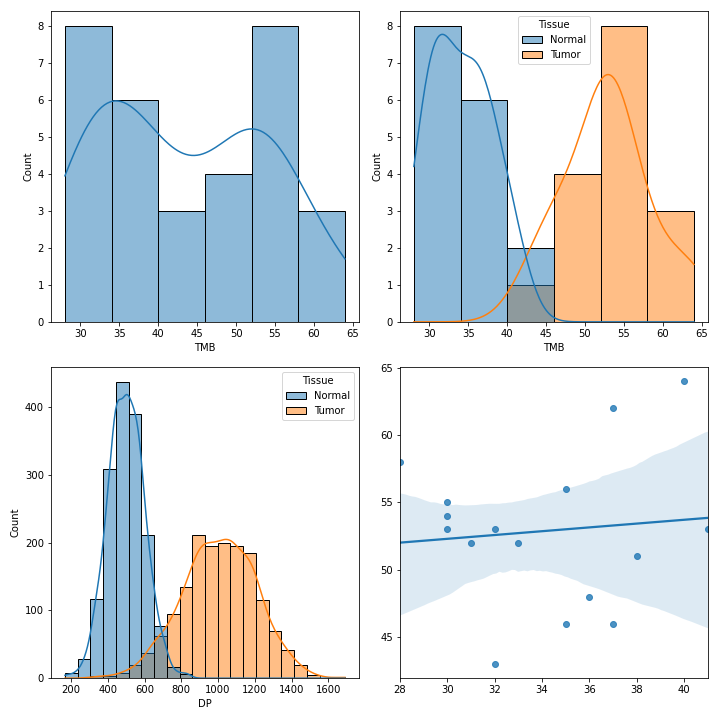
MAF
To create an oncoplot with a MAF file:
>>> from fuc import common, pymaf
>>> common.load_dataset('tcga-laml')
>>> maf_file = '~/fuc-data/tcga-laml/tcga_laml.maf.gz'
>>> mf = pymaf.MafFrame.from_file(maf_file)
>>> mf.plot_oncoplot()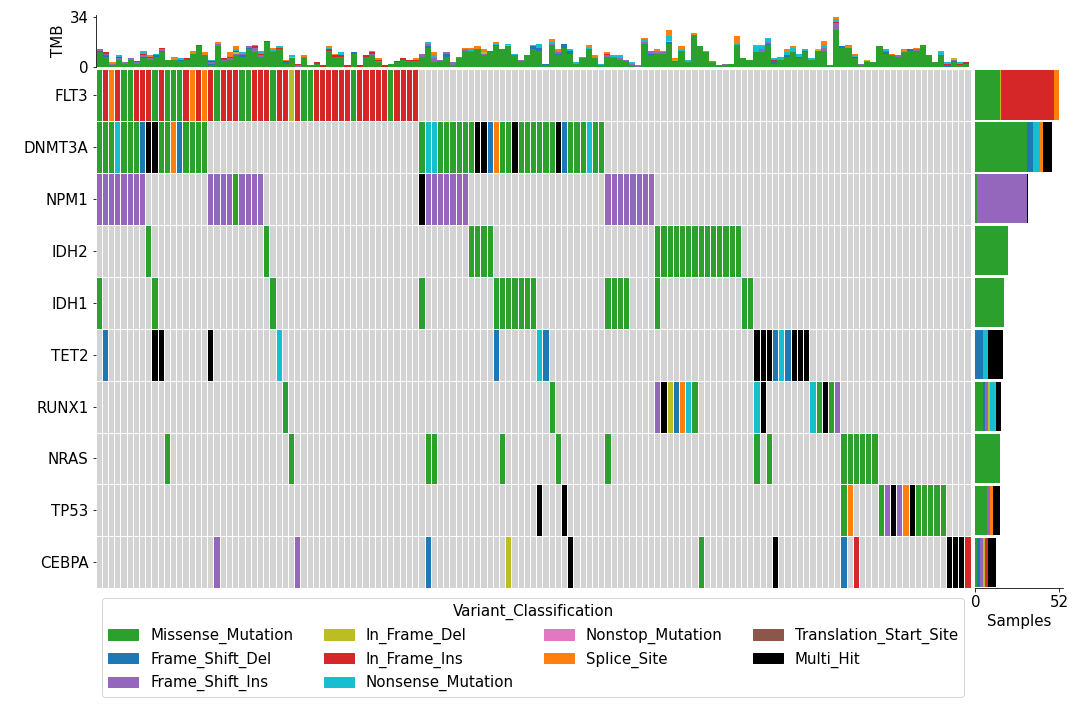
To create a customized oncoplot with a MAF file, see the Create customized oncoplot tutorial:
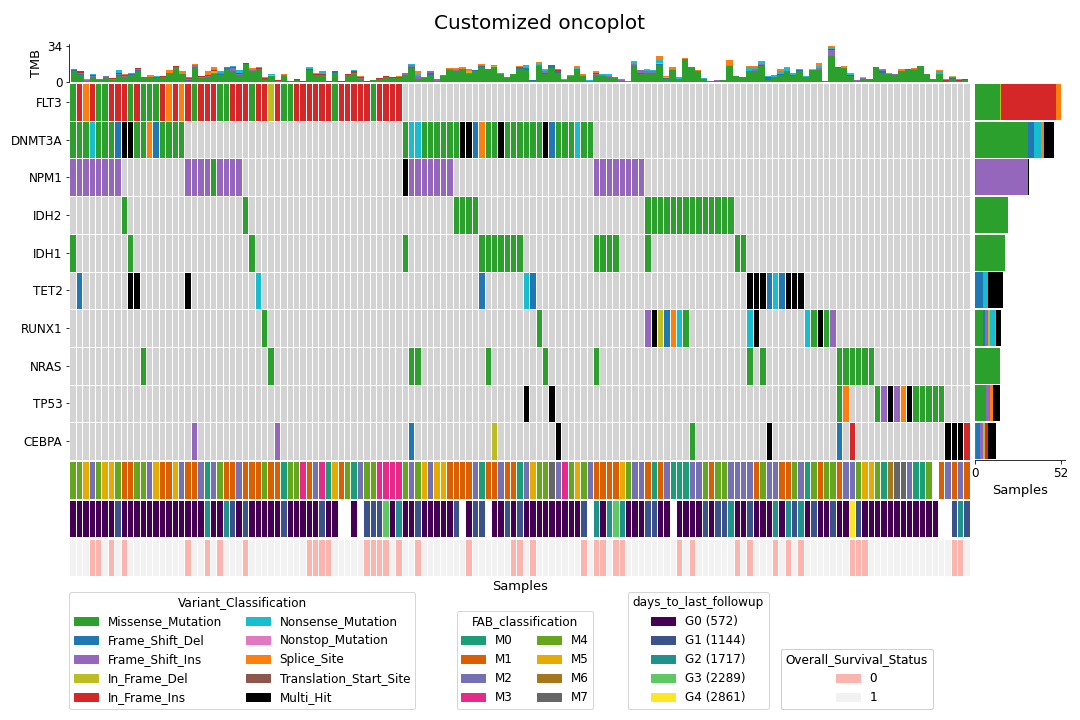
To create a summary figure for a MAF file:
>>> from fuc import common, pymaf
>>> common.load_dataset('tcga-laml')
>>> maf_file = '~/fuc-data/tcga-laml/tcga_laml.maf.gz'
>>> mf = pymaf.MafFrame.from_file(maf_file)
>>> mf.plot_summary()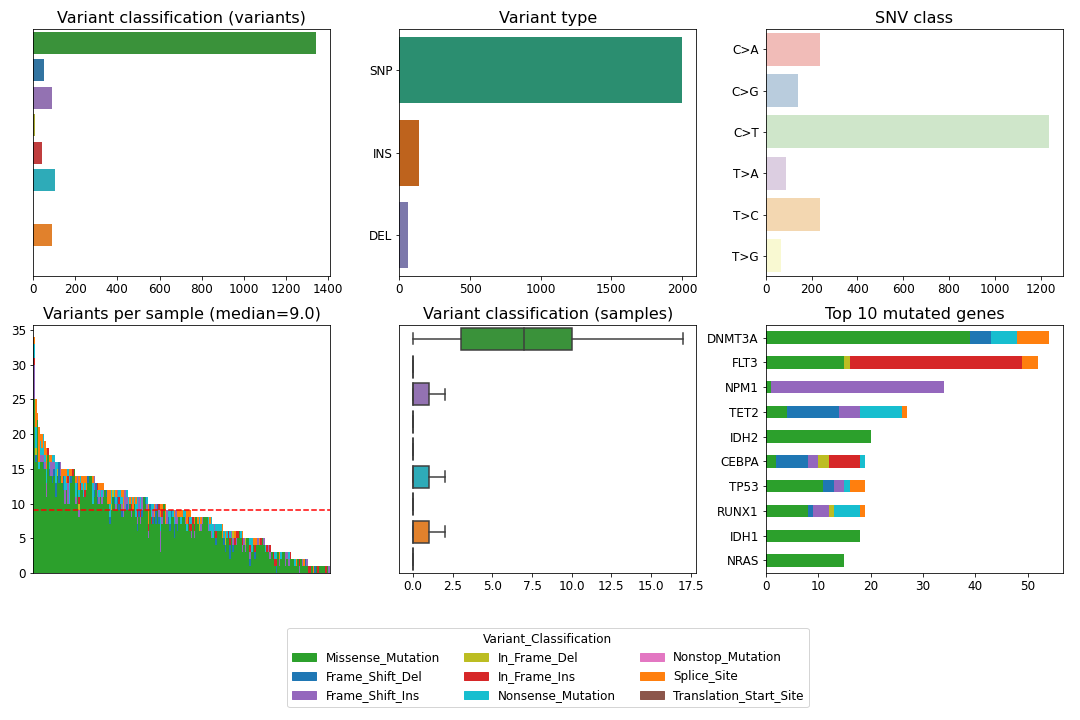
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file fuc-0.38.0.tar.gz.
File metadata
- Download URL: fuc-0.38.0.tar.gz
- Upload date:
- Size: 128.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.0 CPython/3.10.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
379ffdfeaa77ba4d9fd38360e1032986ad3e3da8b380f57652cbcf4b9c1c13dc
|
|
| MD5 |
9e6a7e6a41055e0f64d97340c8ce2363
|
|
| BLAKE2b-256 |
73cdaccbd737f49c9f313d440d17b9562acebece6e5a8214517eab2adfb192bd
|
File details
Details for the file fuc-0.38.0-py3-none-any.whl.
File metadata
- Download URL: fuc-0.38.0-py3-none-any.whl
- Upload date:
- Size: 154.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.0 CPython/3.10.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7b73d5efc220510a735f815eea3ebfa4c775aed45559b325aa390fa3be6490dd
|
|
| MD5 |
e52d83bfb1e25a8b94772aa6f362a0e4
|
|
| BLAKE2b-256 |
deef39b9f73bc9dac83aa7c21ff824807884d35c7af333f739813ec1a958f544
|











