Generalized additive models with a Bayesian twist
Project description
Gammy – Generalized additive models in Python with a Bayesian twist
A Generalized additive model is a predictive mathematical model defined as a sum of terms that are calibrated (fitted) with observation data.
Generalized additive models form a surprisingly general framework for building models for both production software and scientific research. This Python package offers tools for building the model terms as decompositions of various basis functions. It is possible to model the terms e.g. as Gaussian processes (with reduced dimensionality) of various kernels, as piecewise linear functions, and as B-splines, among others. Of course, very simple terms like lines and constants are also supported (these are just very simple basis functions).
The uncertainty in the weight parameter distributions is modeled using Bayesian statistical analysis with the help of the superb package BayesPy. Alternatively, it is possible to fit models using just NumPy.
Table of Contents
Installation
The package is found in PyPi.
pip install gammy
Examples
In this overview, we demonstrate the package's most important features through common usage examples.
Polynomial regression on 'roids
A typical simple (but sometimes non-trivial) modeling task is to estimate an unknown function from noisy data. First we import the bare minimum dependencies to be used in the below examples:
>>> import numpy as np
>>> import gammy
>>> from gammy.models.bayespy import GAM
>>> gammy.__version__
'0.5.6'
Let's simulate a dataset:
>>> np.random.seed(42)
>>> n = 30
>>> input_data = 10 * np.random.rand(n)
>>> y = 5 * input_data + 2.0 * input_data ** 2 + 7 + 10 * np.random.randn(n)
The object x is just a convenience tool for defining input data maps
as if they were just Numpy arrays:
>>> from gammy.arraymapper import x
Define and fit the model:
>>> a = gammy.formulae.Scalar(prior=(0, 1e-6))
>>> b = gammy.formulae.Scalar(prior=(0, 1e-6))
>>> bias = gammy.formulae.Scalar(prior=(0, 1e-6))
>>> formula = a * x + b * x ** 2 + bias
>>> model = GAM(formula).fit(input_data, y)
The model attribute model.theta characterizes the Gaussian posterior
distribution of the model parameters vector.
>>> model.mean_theta
[array([3.20130444]), array([2.0420961]), array([11.93437195])]
Variance of additive zero-mean normally distributed noise is estimated automagically:
>>> round(model.inv_mean_tau, 8)
74.51660744
Predicting with model
>>> model.predict(input_data[:2])
array([ 52.57112684, 226.9460579 ])
Predictions with uncertainty, that is, posterior predictive mean and variance can be calculated as follows:
>>> model.predict_variance(input_data[:2])
(array([ 52.57112684, 226.9460579 ]), array([79.35827362, 95.16358131]))
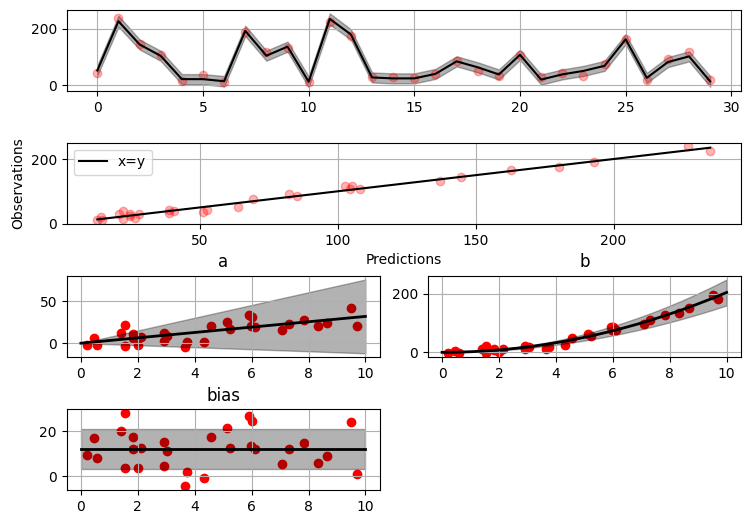
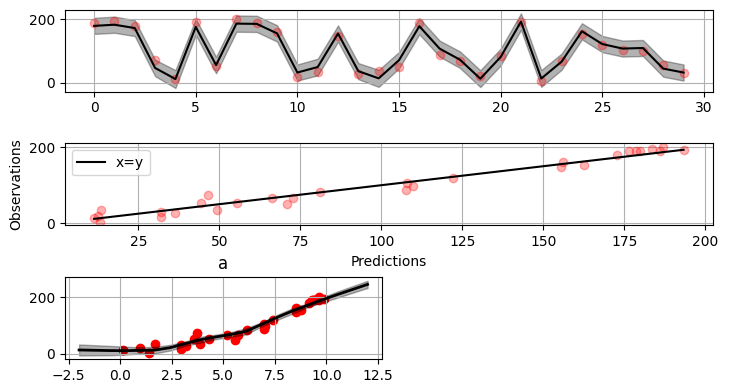
Plotting results
>>> fig = gammy.plot.validation_plot(
... model,
... input_data,
... y,
... grid_limits=[0, 10],
... input_maps=[x, x, x],
... titles=["a", "b", "bias"]
... )
The grey band in the top figure is two times the prediction standard deviation and, in the partial residual plots, two times the respective marginal posterior standard deviation.
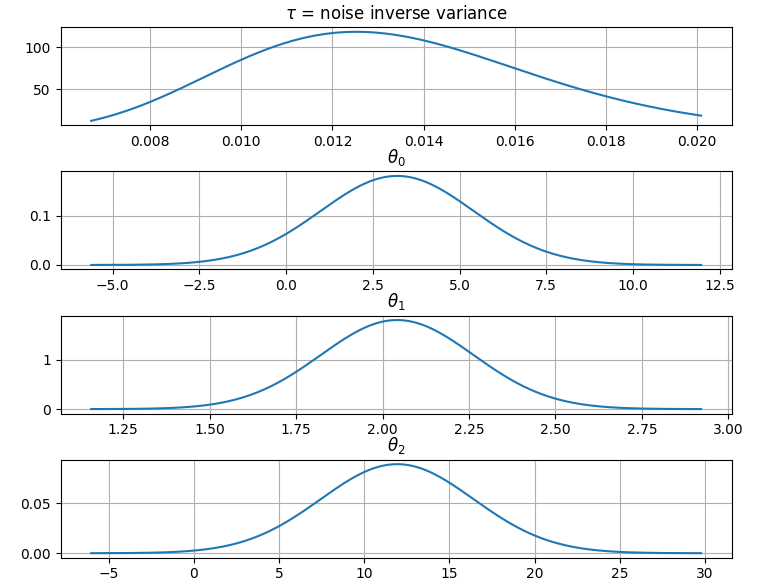
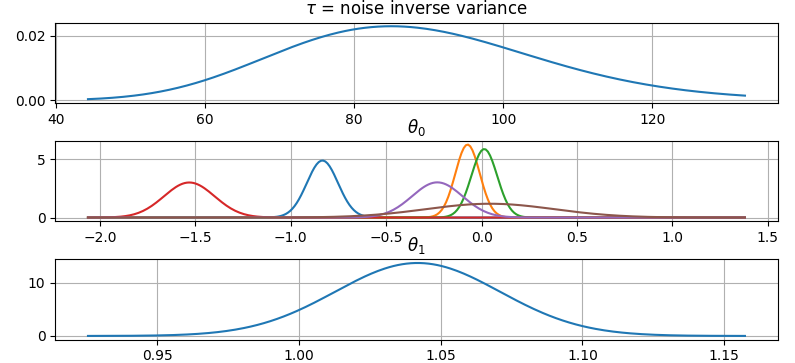
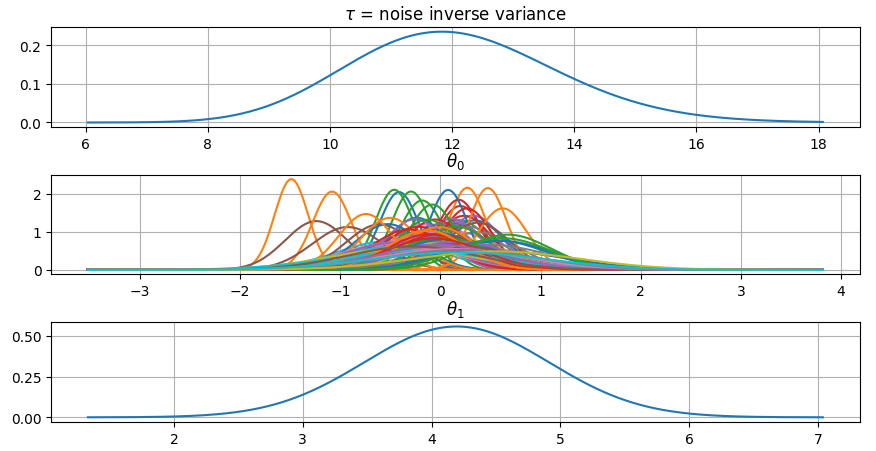
It is also possible to plot the estimated Γ-distribution of the noise precision (inverse variance) as well as the 1-D Normal distributions of each individual model parameter.
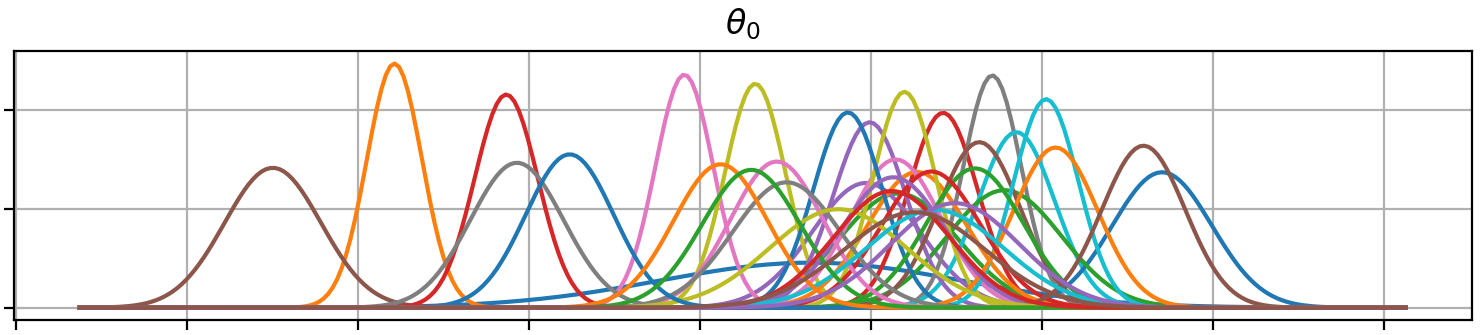
Plot (prior or posterior) probability density functions of all model parameters:
>>> fig = gammy.plot.gaussian1d_density_plot(model)
Saving model on hard disk
Saving:
>> model.save("/home/foobar/test.hdf5")
Loading:
>> model = GAM(formula).load("/home/foobar/test.hdf5")
Gaussian process regression
Create fake dataset:
>>> n = 50
>>> input_data = np.vstack((2 * np.pi * np.random.rand(n), np.random.rand(n))).T
>>> y = (
... np.abs(np.cos(input_data[:, 0])) * input_data[:, 1] +
... 1 + 0.1 * np.random.randn(n)
... )
Define model:
>>> a = gammy.formulae.ExpSineSquared1d(
... np.arange(0, 2 * np.pi, 0.1),
... corrlen=1.0,
... sigma=1.0,
... period=2 * np.pi,
... energy=0.99
... )
>>> bias = gammy.Scalar(prior=(0, 1e-6))
>>> formula = a(x[:, 0]) * x[:, 1] + bias
>>> model = gammy.models.bayespy.GAM(formula).fit(input_data, y)
>>> round(model.mean_theta[0][0], 8)
0.8343458
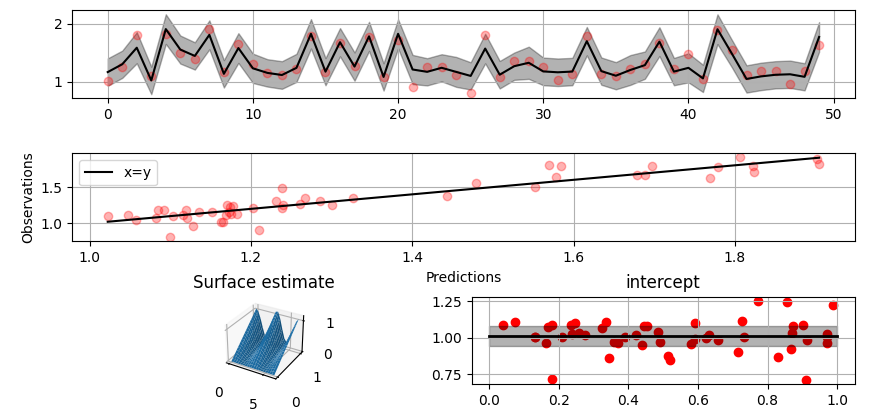
Plot predictions and partial residuals:
>>> fig = gammy.plot.validation_plot(
... model,
... input_data,
... y,
... grid_limits=[[0, 2 * np.pi], [0, 1]],
... input_maps=[x[:, 0:2], x[:, 1]],
... titles=["Surface estimate", "intercept"]
... )
Plot parameter probability density functions
>>> fig = gammy.plot.gaussian1d_density_plot(model)
More covariance kernels
The package contains covariance functions for many well-known options such as the Exponential squared, Periodic exponential squared, Rational quadratic, and the Ornstein-Uhlenbeck kernels. Please see the documentation section More on Gaussian Process kernels for a gallery of kernels.
Defining custom kernels
Please read the documentation section: Customize Gaussian Process kernels
Spline regression
Constructing B-Spline based 1-D basis functions is also supported. Let's define dummy data:
>>> n = 30
>>> input_data = 10 * np.random.rand(n)
>>> y = 2.0 * input_data ** 2 + 7 + 10 * np.random.randn(n)
Define model:
>>> grid = np.arange(0, 11, 2.0)
>>> order = 2
>>> N = len(grid) + order - 2
>>> sigma = 10 ** 2
>>> formula = gammy.BSpline1d(
... grid,
... order=order,
... prior=(np.zeros(N), np.identity(N) / sigma),
... extrapolate=True
... )(x)
>>> model = gammy.models.bayespy.GAM(formula).fit(input_data, y)
>>> round(model.mean_theta[0][0], 8)
-49.00019115
Plot validation figure:
>>> fig = gammy.plot.validation_plot(
... model,
... input_data,
... y,
... grid_limits=[-2, 12],
... input_maps=[x],
... titles=["a"]
... )
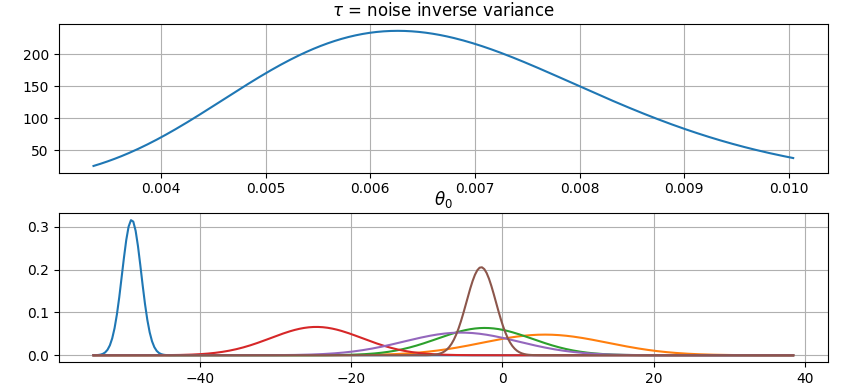
Plot parameter probability densities:
>>> fig = gammy.plot.gaussian1d_density_plot(model)
Non-linear manifold regression
In this example we try estimating the bivariate "MATLAB function" using a Gaussian process model with Kronecker tensor structure (see e.g. PyMC3). The main point in the below example is that it is quite straightforward to build models that can learn arbitrary 2D-surfaces.
Let us first create some artificial data using the MATLAB function!
>>> n = 100
>>> input_data = np.vstack((
... 6 * np.random.rand(n) - 3, 6 * np.random.rand(n) - 3
... )).T
>>> y = (
... gammy.utils.peaks(input_data[:, 0], input_data[:, 1]) +
... 4 + 0.3 * np.random.randn(n)
... )
There is support for forming two-dimensional basis functions given two one-dimensional formulas. The new combined basis is essentially the outer product of the given bases. The underlying weight prior distribution priors and covariances are constructed using the Kronecker product.
>>> a = gammy.ExpSquared1d(
... np.arange(-3, 3, 0.1),
... corrlen=0.5,
... sigma=4.0,
... energy=0.99
... )(x[:, 0]) # NOTE: Input map is defined here!
>>> b = gammy.ExpSquared1d(
... np.arange(-3, 3, 0.1),
... corrlen=0.5,
... sigma=4.0,
... energy=0.99
... )(x[:, 1]) # NOTE: Input map is defined here!
>>> A = gammy.formulae.Kron(a, b)
>>> bias = gammy.formulae.Scalar(prior=(0, 1e-6))
>>> formula = A + bias
>>> model = GAM(formula).fit(input_data, y)
>>> round(model.mean_theta[0][0], 8)
0.37426986
Note that same logic could be used to construct higher dimensional bases, that is, one could define a 3D-formula:
>> formula_3d = gammy.Kron(gammy.Kron(a, b), c)
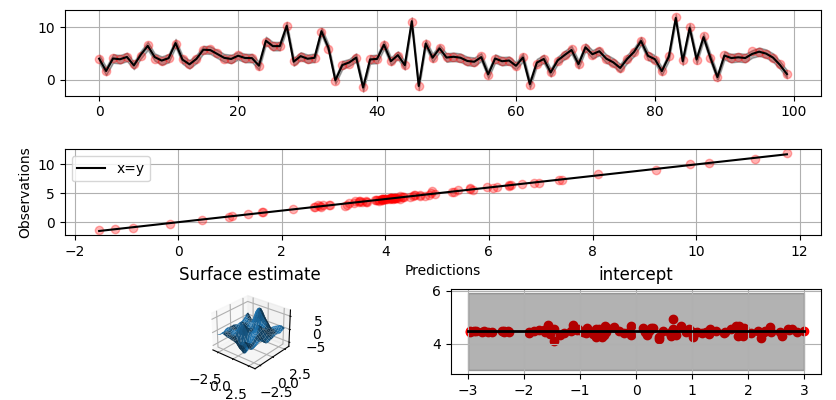
Plot predictions and partial residuals:
>>> fig = gammy.plot.validation_plot(
... model,
... input_data,
... y,
... grid_limits=[[-3, 3], [-3, 3]],
... input_maps=[x, x[:, 0]],
... titles=["Surface estimate", "intercept"]
... )
Plot parameter probability density functions:
>>> fig = gammy.plot.gaussian1d_density_plot(model)
Testing
The package's unit tests can be ran with PyTest (cd to repository root):
pytest -v
Running this documentation as a Doctest:
python -m doctest -v README.md
Package documentation
Documentation of the package with code examples: https://malmgrek.github.io/gammy.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file gammy-0.5.6.tar.gz.
File metadata
- Download URL: gammy-0.5.6.tar.gz
- Upload date:
- Size: 24.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bb80971dc7d7c4fb0deafb32c0db27514030d1e13de665f1fbf34805f3e59537
|
|
| MD5 |
13adf95a0c1e7fef16424e5a8ce9ec85
|
|
| BLAKE2b-256 |
072e1590e9267c3567efd8dc4bc7f74bbc6eb28ceae9d10f6be774101dcd84fc
|
File details
Details for the file gammy-0.5.6-py3-none-any.whl.
File metadata
- Download URL: gammy-0.5.6-py3-none-any.whl
- Upload date:
- Size: 24.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b2f926ad90b09d01c87711a5977b39298ba92aa4dbfb9bce47f97f6bbb62be6e
|
|
| MD5 |
b8e566a94c9e49dfd919ddd28c782ca9
|
|
| BLAKE2b-256 |
c34142f0adaa6b9503b2538a828422063b5e20a411b0c3d8c53c30cc2e6a1c9a
|