IBEIS - Image Based Ecological Information System
Project description

NOTE: This IBEIS software is the result of my (Jon Crall’s) PhD work. After I graduated in 2017, the image analysis components of IBEIS and the core HotSpotter program were transferred to WildMe, which as of 2024 has merged with Conservation X Labs.
I continue to independently maintain and support this software, but it can only handle medium scale populations and its GUI interface can be difficult to work with. If you have a larger population or the need for simpler and scalable web interfaces please reach out Conservation X Labs.
If you have the ambition to run and possibly develop software yourself, then keep reading.
IBEIS - Image Analysis
I.B.E.I.S. = Image Based Ecological Information System

Installation Instructions (updated 2020-Nov-01)
The IBEIS software is now available on pypi for Linux systems. This means if you have Python installed. You can simply run:
pip install ibeisto install the software. Then the command to run the GUI is:
ibeisOn Windows / OSX I recommend using a Linux virtual machine. However, if you are computer savvy it is possible to build all of the requirements on from source. The only tricky components are installing the packages with binary dependencies: pyhesaff and vtool_ibeis. If you have these built then the rest of the dependencies can be installed from pypi even on OSX / Windows.
NOTE: When using a VM on windows, you may encounter an error:
qt.qpa.plugin: Could not load the Qt platform plugin "xcb" in
... even though it was found. This application failed to start because no
Qt platform plugin could be initialized. Reinstalling the application may
fix this problem.
Available platform plugins are: xcb, eglfs, ...
Core dumpedThe reason the issue happens appears to be because the opencv-python package includes libraries also packaged with PyQt5 and those are conflicting.
The workaround is to uninstall opencv-python and then install a variant that does not include extra Qt libs:
pip uninstall opencv-python
pip install opencv-python-headlessBasic Usage (updated 2020-Nov-01)
After installation running the ibeis command will open the GUI:
If you have already made a database, it will automatically open the most recently used database.
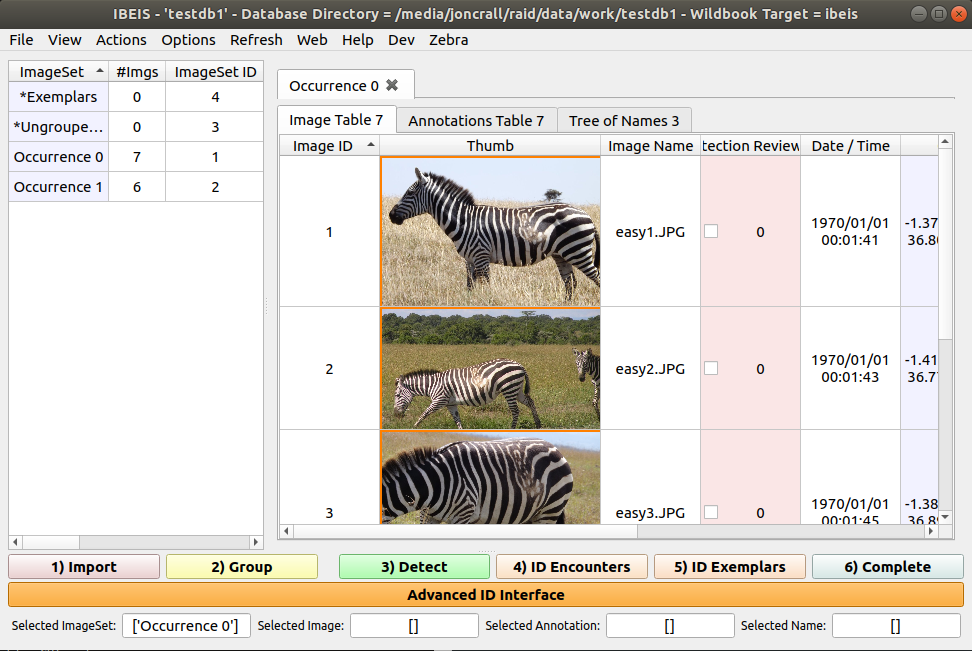
If this is the first time you’ve run the program it will not have a database opened:
Select new database, (which will first ask you to select a work directory where all of your databases will live). Then you will be asked to create a database name. Select one and then create the database in your work directory.
You can drag and drop images into the GUI to add them to the database. Double clicking an image lets you add “annotations”:
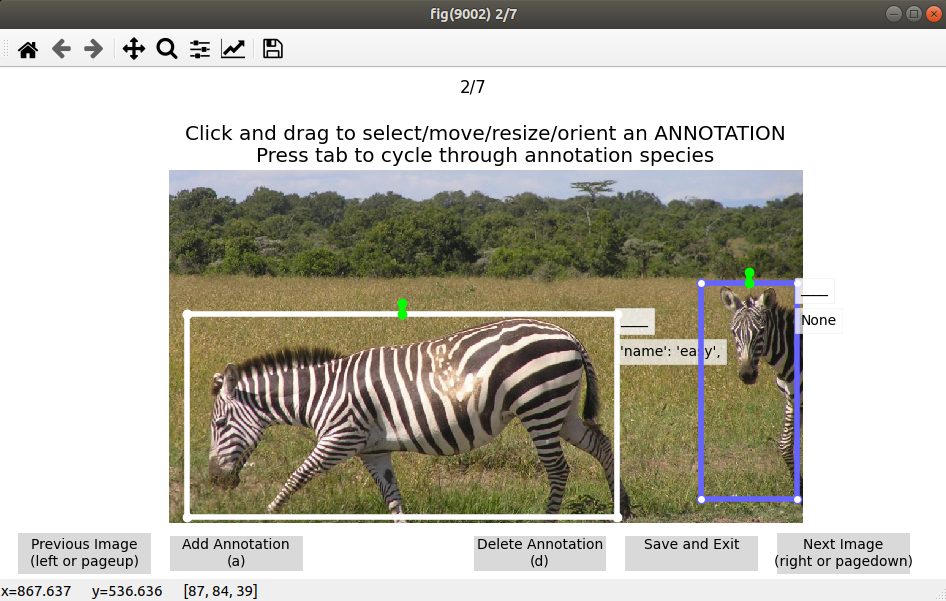
You can also right click one or more images and click “Add annotations from entire images” if your images are already localized to a single individual.
It important than when you add an annotation, you set its species. You can right click multiple annotations and click “set annotation species”. Change this to anything other than “____”.
Once you have annotations with species, you can click one and press “q” to query for matches in the database of other annotations:

Right clicking and marking each match as “True” or “False” (or alternatively selecting a row and pressing “T” or “F”) will mark images as the same or different individuals. Groups marked as the same individual will appear in the “Tree of Names”.
Note there are also batch identification methods in the “ID Encounters” “ID Exemplars” and “Advanced ID Interface” (my personal recommendation). Play around with different right-click menus (although note that some of these are buggy and will crash the program), but the main simple identification procedures are robust and should not crash.
Program Description
IBEIS program for the storage and management of images and derived data for use in computer vision algorithms. It aims to compute who an animal is, what species an animal is, and where an animal is with the ultimate goal being to ask important why biological questions. This This repo Image Analysis image analysis module of IBEIS. It is both a python module and standalone program.
Currently the system is build around and SQLite database, a PyQt5 GUI, and matplotlib visualizations. Algorithms employed are: random forest species detection and localization, hessian-affine keypoint detection, SIFT keypoint description, LNBNN identification using approximate nearest neighbors. Algorithms in development are SMK (selective match kernel) for identification and deep neural networks for detection and localization.
The core of IBEIS is the IBEISController class. It provides an API into IBEIS data management and algorithms. The IBEIS API Documentation can be found here: http://erotemic.github.io/ibeis
The IBEIS GUI (graphical user interface) is built on top of the API. We are also experimenting with a new web frontend that bypasses the older GUI code.
Self Installing Executables
Unfortunately we have not released self-installing-executables for IBEIS yet. We hope to release these soon™, but developer support is needed (contributions welcome).
However there are old [HotSpotter](https://github.com/Erotemic/hotspotter) (the predecessor to IBEIS) binaries available.
Dropbox should still be hosting the download links:
Win32 Installer: https://www.dropbox.com/s/5j1xyx2hq1wzqz2/hotspotter-win32-setup.exe?dl=0
OSX Installer: https://www.dropbox.com/s/q0vzz3xnjbxhsda/hotspotter_installer_mac.dmg?dl=0
IPFS CIDs for the previous installers are QmSnyetkniriHUwTxvzwhkysPKjUj7udBqq5mpK24VJXVM and QmZ3WknrAaxPZhZebdQWZ45EEKwu1Tr6bkFWJzfPRtENs7.
If you are unfamiliar with IPFS use the following gateway links:
Visual Demo
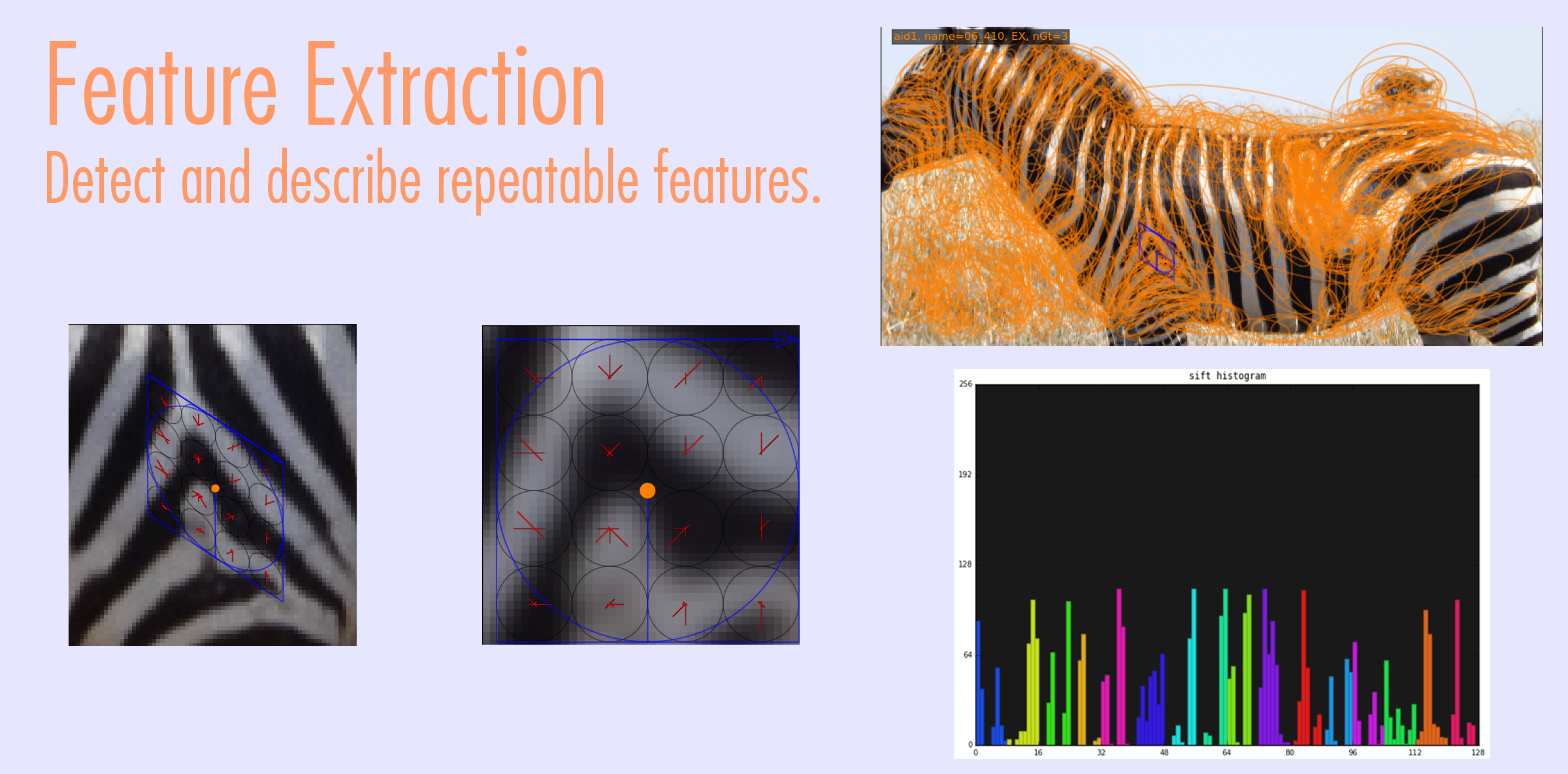
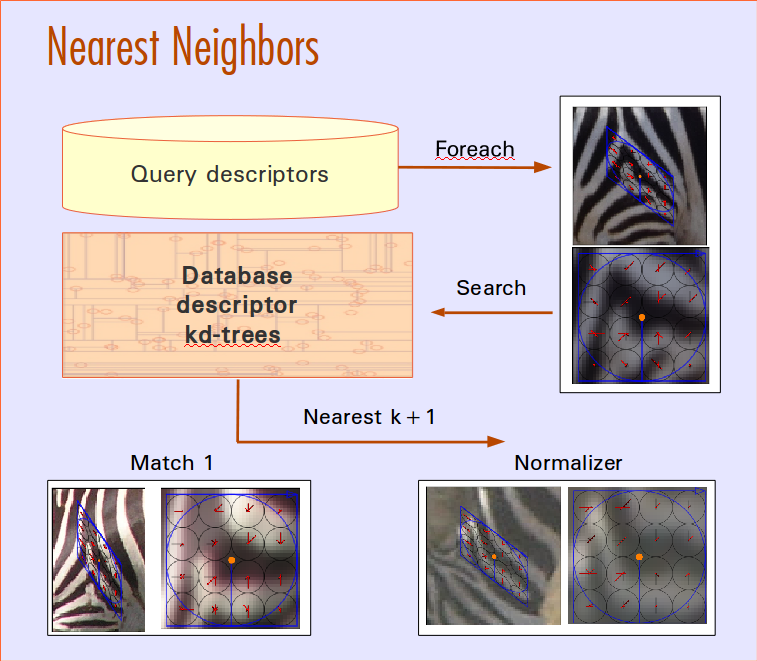
Match Scoring
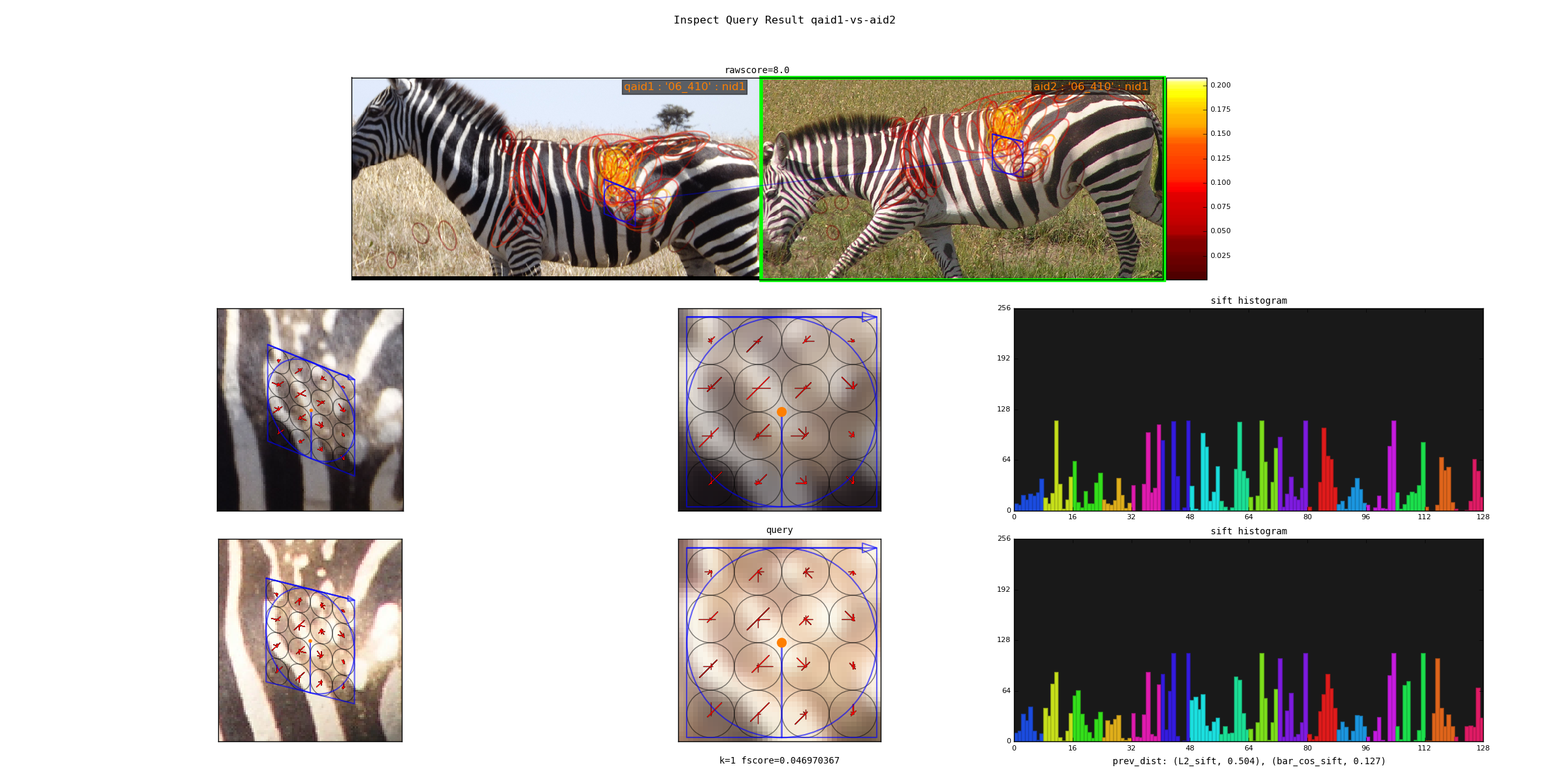
Spatial Verification

python -m vtool.spatial_verification spatially_verify_kpts --showName Scoring
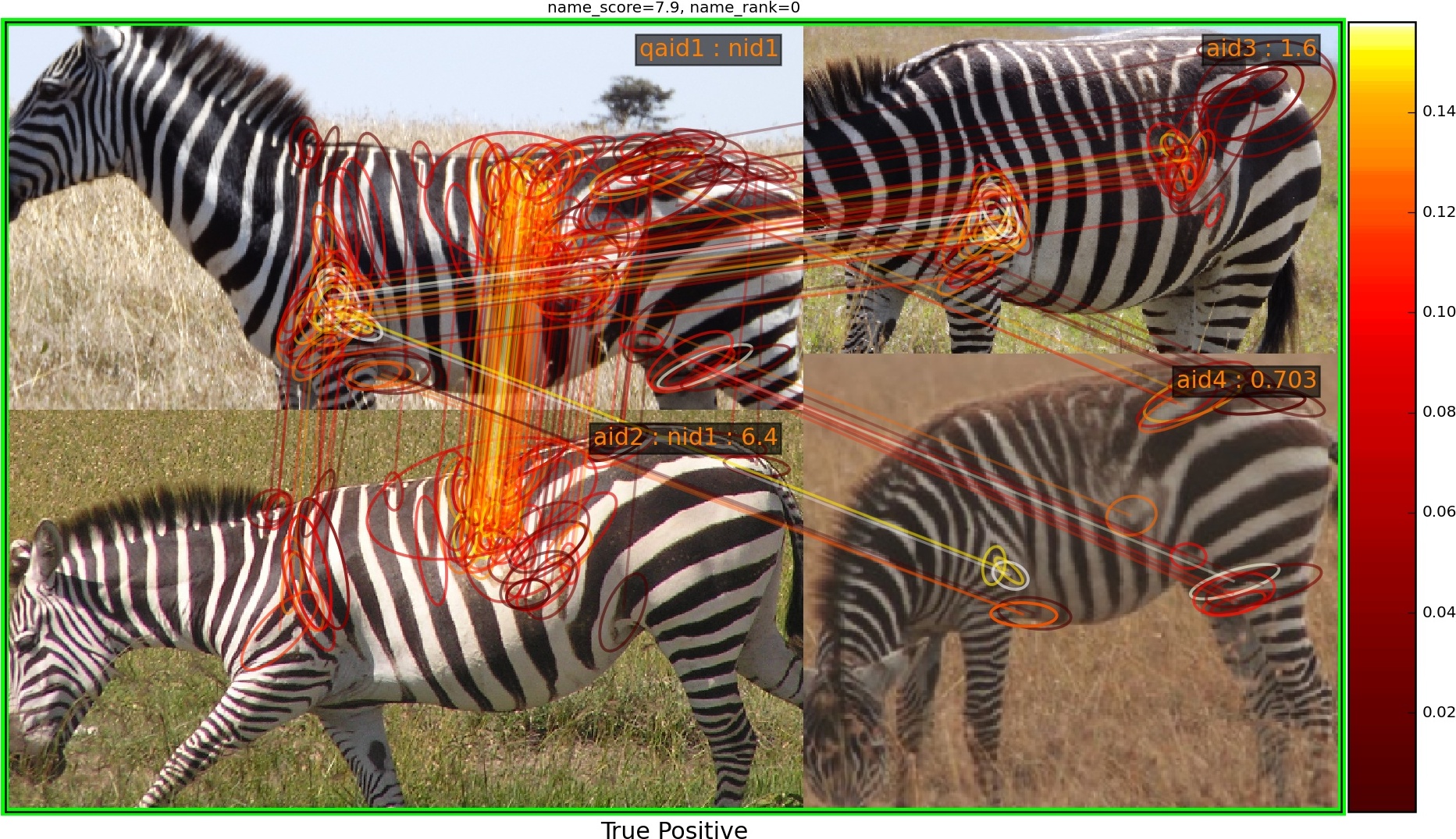

python -m ibeis.algo.hots.chip_match show_single_namematch --qaid 1 --showIdentification Ranking

python -m ibeis.algo.hots.chip_match show_ranked_matches --show --qaid 86Inference
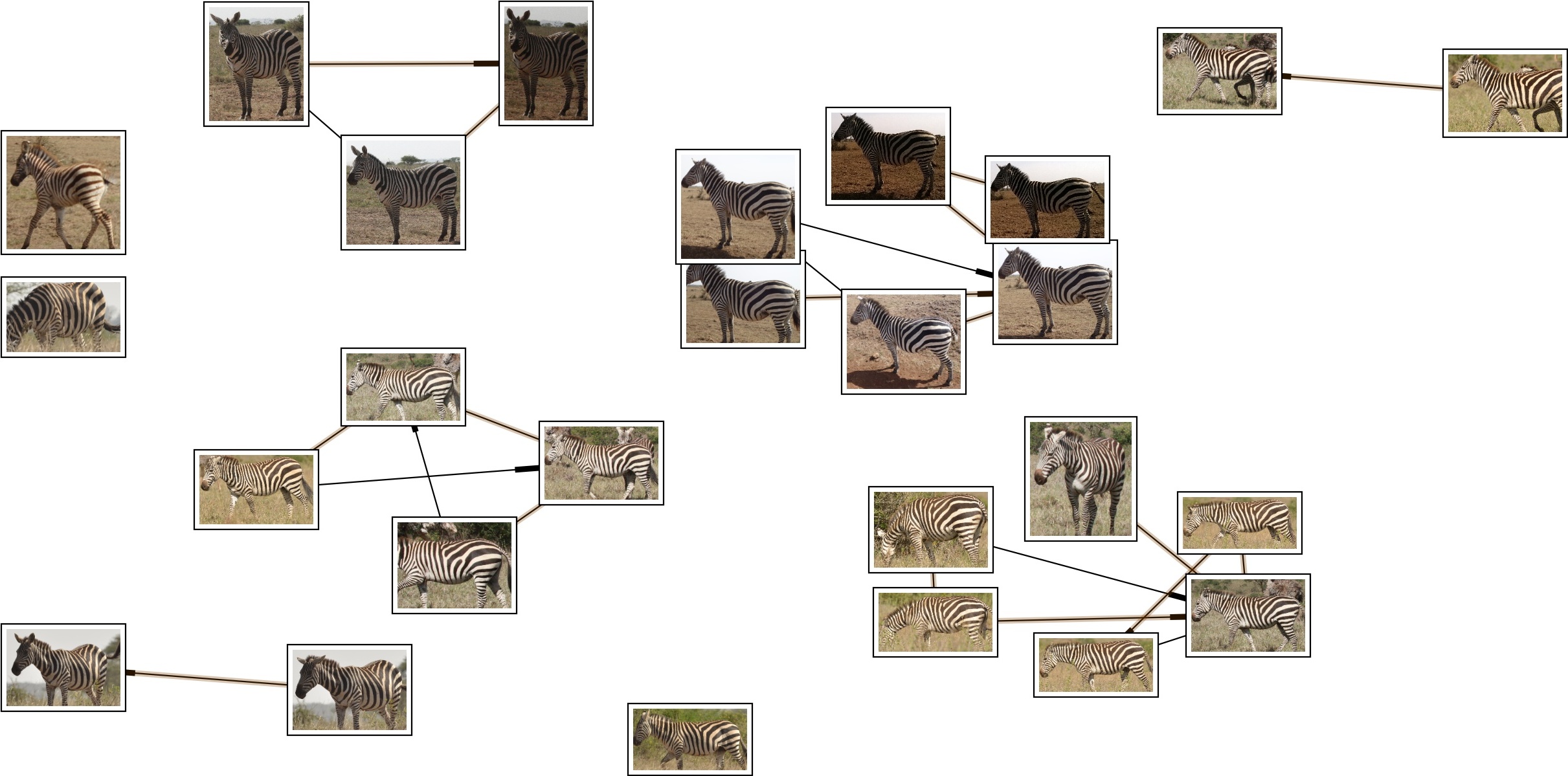
# broken
# python -m ibeis.algo.preproc.preproc_encounter compute_encounter_groups --showInternal Modules
In the interest of modular code we are actively developing several different modules.
— |
bluemellophone’s IBEIS Image Analysis modules
Building from source
To build from source you need to be able to build the following 3 projects with binary dependences. These depened on having a development version of OpenCV and LZ4.
If you are on Linux simply using the wheels for the above projects ( i.e. pip install vtool_ibeis_ext pyflann_ibeis pyhesaff is recommended). On OSX and Win32 these need to be build manually (any contributions to help these build win32 or osx wheels on their respective project CI would be amazing!).
The rest of the dependency repos (guitool_ibeis, plottool_ibeis, dtool_ibeis, vtool_ibeis,) are pure python and can be installed in development mode with the normal clone the repo, and run pip install -e . inside the repo process.
Given a Python environment where each of the dependency modules is installed this repo can be installed with pip install -e . as well.
Demo Databases
A set of small (400MB) demo databases PZ_MTEST1 and NAUT_TEST can be downloaded via.
python dev/reset_dbs.pyNote that it downloads the data from an IPFS gateway, which may be slow and require several attempts before it works.
Running Tests
The tests depend on an even smaller set of test databases, which can be prepared via ibeis –reset-ci-dbs. Once you have the test data you can use the run_doctests.sh or run_tests.py script to execute the system tests.
Known issues
We do not add or remove points from kdtrees. They are always rebuilt
Changing algorithm settings in the preferences window seems bugged and probably requires a fix in utool. The default settings should work and using the Python API with explicit algorithm settings should also be fine.
The code is a mess! PRs are welcome!
The UI has lots of unused or broken components. PRs to disable or make these nicer are welcome!
The only blockers for Windows / OSX pip installs are pyhesaff, pyflann_ibeis, and vtool_ibeis_ext binary wheels. Getting these to build on Github Actions would mean pip installable IBEIS on non-Linux systems.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file ibeis-2.4.0-py3-none-any.whl.
File metadata
- Download URL: ibeis-2.4.0-py3-none-any.whl
- Upload date:
- Size: 1.4 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e6cc450f546c39be13da04e5bddd8577e6ebb4fac6775ff6076d4ab0d95452c5
|
|
| MD5 |
3b37c67e24a0a7d886c197dd000f2f37
|
|
| BLAKE2b-256 |
94d71e4b684d7368560374cfeef2129e8e131b7660757309fed3ebb50340e871
|























