Interactive visualization and processing of 3D medical images in python
Project description
imvis
Interactive visualization of 3D medical images in python
Installation
The package can be installed from PyPI using pip:
pip install imvis
Development Setup
This project uses UV for dependency management.
Install UV
# Windows (PowerShell)
irm https://astral.sh/uv/install.ps1 | iex
# Linux/macOS
curl -LsSf https://astral.sh/uv/install.sh | sh
Setup Development Environment
# Clone the repository
git clone https://github.com/MengXiangxi/imvis.git
cd imvis
# Install dependencies
uv sync
# Run tests
uv run pytest tests/test_imvis.py -v
Features
3D image visualization
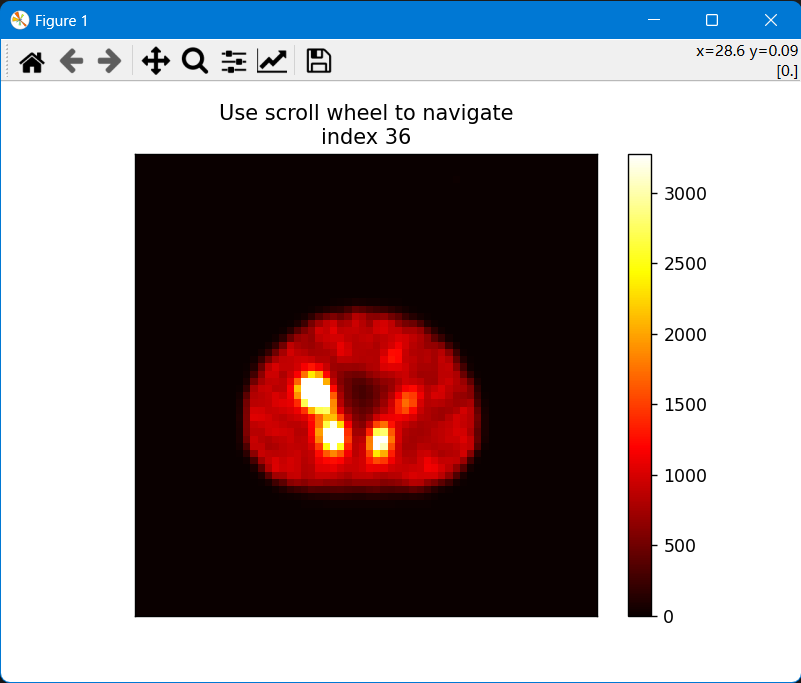
imagesc3s is a function that allows to visualize 3D images in a 2D slice-by-slice fashion. It is based on the matplotlib library and allows to interactively scroll through the slices of a 3D image. It also allows to change the color map and the windowing of the image.
import imvis as iv
import pydicom
ds = pydicom.dcmread('path/to/dicom/file')
img = ds.pixel_array
iv.imagesc3s(img)
In cases where scrolling is not possible (e.g. in a Jupyter notebook), the alternative version imagesc3slider can be used. It allows to scroll through the slices of a 3D image using a slider.
iv.imagesc3slider(img)
When using Jupyter notebook, the matplotlib backend can be changed to tk or qt to enable scrolling. This can be done using the following magic command:
%matplotlib tk
iv.imagesc3slider(img)
MIP with rotation angles
mipz allows the user to obtain a maximum intensity projection (MIP) of a 3D image along the z-axis. The user can also specify the rotation angles of the MIP.
import SimpleITK as sitk
import numpy as np
img = sitk.ReadImage("/path/to/nifti")
imarray = sitk.GetArrayFromImage(img)
mip_array = np.zeros((36, imarray.shape[0], imarray.shape[1]))
for i in range(0, 360, 10):
mip_array[int(i/10),:,:] = iv.mipz(imarray, i)
iv.imagesc3s(mip_array, [0, 10])
NIFTI image resampling in reference to another image
resample_nifti_to allows to resample a NIFTI image in reference to another image. This is useful when you want to resample a NIFTI image to the same resolution as a DICOM image.
def resample_nifti_to(nifti_in, nifti_ref, fname_out, img_type='intensity'):
"""Resample a nifti image to the same space as another nifti image.
Parameters
----------
nifti_in : string
Path to the nifti image to be resampled.
nifti_ref : string
Path to the nifti image to be used as reference.
fname_out : string
Path to the resampled nifti image.
img_type : string, optional
Type of the image. Default is 'intensity'.
'intensity': general type, no conversion.
'BQML': PET or quantitative SPECT image, total counts are preserved.
'mask': interger mask, interpolation will not change the value.
"""
Convert PET DICOM to NIFTI with SUV
dicom2niftiSUV allows to convert a PET DICOM image to a NIFTI image with SUV values. The SUV values are computed using the corresponding DICOM tags.
- bodyweight: "TBW" (total body weight) or "LBW" (lean body weight)
def dicom2niftiSUV(dicomdir, niftiname,bodyweight="TBW"):
"""Convert a folder of dicom files to nifti files and apply SUV conversion.
Parameters
----------
dicomdir : string
Path to the folder containing dicom files.
niftiname : string
Path and filename to the output nifti file.
"""
Sort files in the DICOMDIR file into hierarchical folders
dicomdir_split allows to sort the files in a DICOMDIR file into hierarchical folders in the Patient/Study/Series fashion. This might be useful when extracting the desired DICOM series from a DICOMDIR file.
def dicomdir_split(dicomdir_path, output_folder):
''' Split DICOM files in the DICOMDIR into different folders based according to patient, studies, and series.
Parameters
----------
dicomdir_path : string
Path to the DICOMDIR file.
output_folder : string
Path to the output folder.
'''
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file imvis-0.1.1.tar.gz.
File metadata
- Download URL: imvis-0.1.1.tar.gz
- Upload date:
- Size: 710.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.10.10 {"installer":{"name":"uv","version":"0.10.10","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":null,"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":null}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b2ebc4007f01bbce230185fe4514835d5d8f03b2054d87a2407332b2db38fdb5
|
|
| MD5 |
32c903ecbd7a0c760b93beabf53b32c1
|
|
| BLAKE2b-256 |
351b52468ead88caac100e435ab7dc37ff588e8bbe6764f512296caa331536e4
|
File details
Details for the file imvis-0.1.1-py3-none-any.whl.
File metadata
- Download URL: imvis-0.1.1-py3-none-any.whl
- Upload date:
- Size: 10.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.10.10 {"installer":{"name":"uv","version":"0.10.10","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":null,"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":null}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2f47a432b97b950da893a6cca19907e7e080de7b9e14f9fc9daf0ae9c5f543e2
|
|
| MD5 |
b45f84240046cd19d5b19ade471cee18
|
|
| BLAKE2b-256 |
0c07919b1c7719d4b786654beb8b25755c97e010bba0a45f055834ec2c2e15fb
|