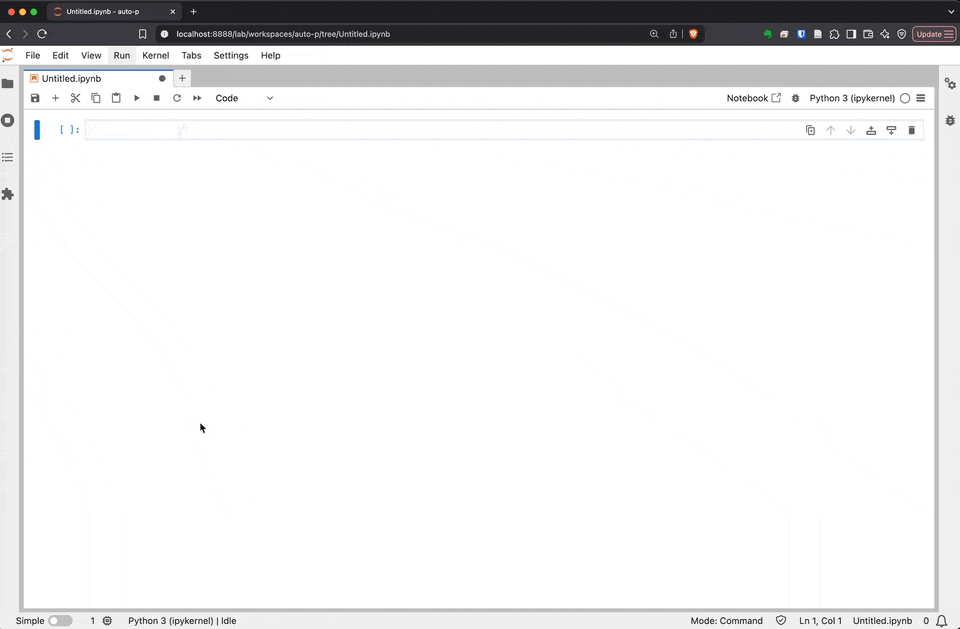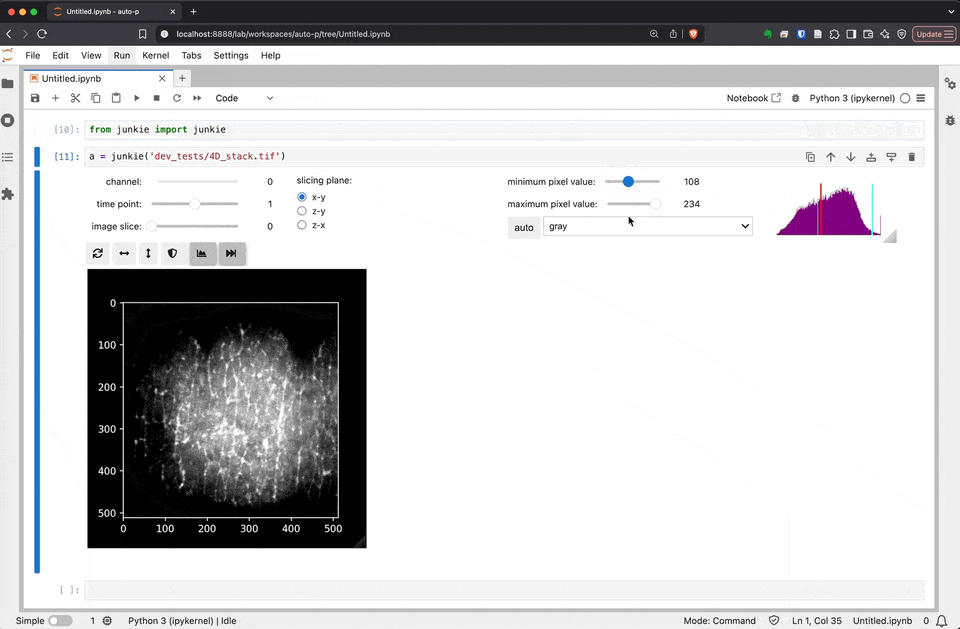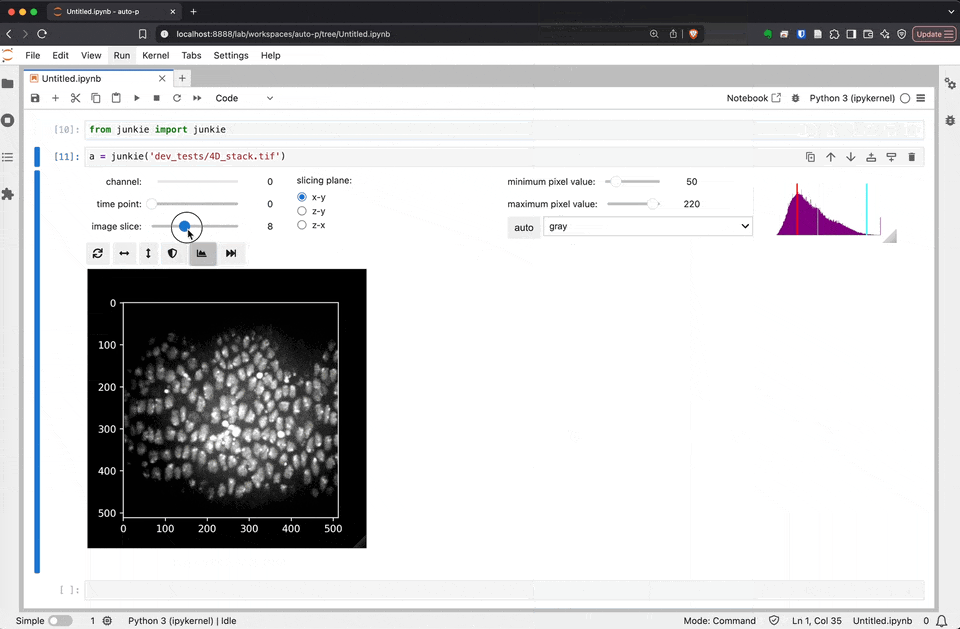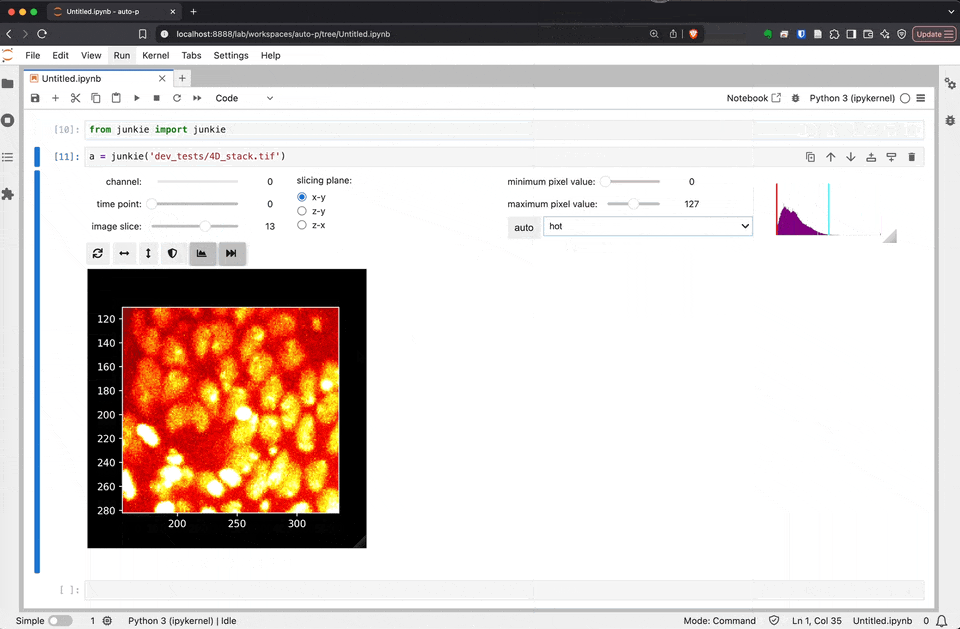
junkie is a JUpyter NotebooK Image Explorer
Project description
JuNkIE (a JUpyter NotebooK Image Explorer)
JuNkIE is a multidimensional image explorer for browsing and simple manipulation of 2D-5D images in Jupyter notebooks. If you are interested in exploring multidimensional images from the command line, checkout CLIMAX, our multidimensional image explorer for the terminal.
Installing JuNkIE
If you want to try out JuNkIE without having to install it, click this button:
JuNkIE is also distributed with a sample notebook that can be run locally, or in a hosted notebook service such as Colab.
We recommend that you install JuNkIE using uv:
$ uv pip install junkie-rfglab
JuNkIE can also be installed using pip:
$ python3 -m pip install --no-cache-dir -U junkie-rfglab
We do most of the development and testing of JuNkIE in Jupyter Lab, so we recommend using Jupyter Lab.
A note on the Python interpreter
JuNkIE requires that you have Python 3.10 or above installed.
Using JuNkIE
JuNkIE is distributed with several sample images that can be found in the docs folder. The images include:
-
cells_movie.tif: single-channel, 150x150 multi-page tiff image with 90 time points; each image in the sequence is the maximum intensity projection of a spinning-disk confocal micrograph depicting epithelial cells in the ectoderm of a fruit fly embryo
-
cells_movie.zarr: three-channel, 512x512 zarr image with 22 time points; each time point represents the maximum intensity projection of a spinning-disk confocal micrograph depicting epithelial cells in the epidermis of a fruit fly embryo; the first channel shows cell nuclei, the second and third channels (repeated) show the cytoskeletal protein filamentous actin
-
slices: two-channel, 512x512 tiff image sequence with 4 time points and 40 Z-slices per time point; each file contains one time point in a specific channel of a spinning-disk confocal micrograph depicting epithelial cells in the head of a fruit fly embryo; the first channel shows the cytoskeletal protein filamentous actin, the second channel shows cell-cell junctions
To start, open a notebook and import JuNkIE:
from junkie import junkie
There are a few ways to open an image with JuNkIE:
-
You can open an image with some other package (e.g. scikit-image, opencv, PIL, etc.) and invoke JuNkIE with a numpy ndarray as the parameter:
import skimage.io as skio astack = skio.imread('cells_movie.tif') myim = junkie(astack) -
You can also specifiy the path to the image that you want to open ...:
myim = junkie('cells_movie.tif') -
... including zarr files:
myim = junkie('cells_movie.zarr') -
Or you can indicate a folder that contains an image sequence:
myim = junkie('./slices') -
If there are image channels split into different files, you can also specify a tuple of strings to distinguish which files in the folder belong to which channel:
myim = junkie('./slices', ('488', '561')) -
You can use a list of numpy arrays and/or paths to concatenate sideways (i.e. display side-by-side, but all the images must have the same dimensions!!):
myim = junkie([astack, 'cells_movie.tif']) -
You can combine the list input with the JuNkIE class method read_image to display image channels side-by-side:
myim = junkie([*junkie.read_image('./slices', ('488', '561'))])
JuNkIE has a couple of additional parameters:
-
cmap (str) indicates the color map used to display the image. It defaults to 'gray'. Check here for a list of color maps.
myim = junkie('cells_movie.tif', cmap='viridis') -
figsize (Tuple[int, int]) specifies the image panel size. The image aspect ratio is maintained. It defaults to (4, 4).
myim = junkie('cells_movie.tif', figsize=(8, 8)
JuNkIE provides access to the standard matplotlib toolbar, and also includes a second toolbar with additional functionality:
| icon | function |
|---|---|
 |
rotate 90° clockwise |
 |
flip horizontally |
 |
flip vertically |
 |
invert color map |
 |
hide/show axes |
 |
continuous/discrete update |
JuNkIE today
As we develop and improve JuNkIE, there may be small changes to the user interface. This is how JuNkIE looks as of today:
Citing JuNkIE
If you use JuNkIE, please cite this repository. We are working on the paper!
Adding functionality onto JuNkIE
If you would like to extend JuNkIE, please make contributions to this repository. You can also check out JuNkIE-picasso, a JuNkIE fork that allows you to define arbitrary image processing pipelines and integrate them into JuNkIE.
Sponsors
We are grateful for the generous support from the following agencies and institutions, which contribute to the development and maintenance of JuNkIE:
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file junkie_rfglab-2026.4.1.tar.gz.
File metadata
- Download URL: junkie_rfglab-2026.4.1.tar.gz
- Upload date:
- Size: 23.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
edd107d0293b46b7f9f28817e15fc8dc6c86214236434c39502d1133f3612db1
|
|
| MD5 |
eec240ea9b19f1771cff462c7f046516
|
|
| BLAKE2b-256 |
2b677271f37a90b56004cbe46c6e98566d3a87b379f3ae03b67d23c99697bc70
|
Provenance
The following attestation bundles were made for junkie_rfglab-2026.4.1.tar.gz:
Publisher:
build_deploy.yaml on rfg-lab/junkie_github
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
junkie_rfglab-2026.4.1.tar.gz -
Subject digest:
edd107d0293b46b7f9f28817e15fc8dc6c86214236434c39502d1133f3612db1 - Sigstore transparency entry: 1266149718
- Sigstore integration time:
-
Permalink:
rfg-lab/junkie_github@d552d29f8e1e7b5ed1f0a93db94a44059e81377d -
Branch / Tag:
refs/tags/v2026.4.1 - Owner: https://github.com/rfg-lab
-
Access:
private
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
build_deploy.yaml@d552d29f8e1e7b5ed1f0a93db94a44059e81377d -
Trigger Event:
push
-
Statement type:
File details
Details for the file junkie_rfglab-2026.4.1-py3-none-any.whl.
File metadata
- Download URL: junkie_rfglab-2026.4.1-py3-none-any.whl
- Upload date:
- Size: 20.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c75069b19194fda49f2293702f7eec52217cc15b35ea623f0c02daeb5e4d5e42
|
|
| MD5 |
1ad8faf8b09edbe36798b44e32008c33
|
|
| BLAKE2b-256 |
1e9ac1356957c06541492fdaab8bfdbe27ee52fdd599744613bae7afa232a13e
|
Provenance
The following attestation bundles were made for junkie_rfglab-2026.4.1-py3-none-any.whl:
Publisher:
build_deploy.yaml on rfg-lab/junkie_github
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
junkie_rfglab-2026.4.1-py3-none-any.whl -
Subject digest:
c75069b19194fda49f2293702f7eec52217cc15b35ea623f0c02daeb5e4d5e42 - Sigstore transparency entry: 1266149798
- Sigstore integration time:
-
Permalink:
rfg-lab/junkie_github@d552d29f8e1e7b5ed1f0a93db94a44059e81377d -
Branch / Tag:
refs/tags/v2026.4.1 - Owner: https://github.com/rfg-lab
-
Access:
private
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
build_deploy.yaml@d552d29f8e1e7b5ed1f0a93db94a44059e81377d -
Trigger Event:
push
-
Statement type: