Kernel Density Estimation (KDE) and sampling.
Project description
kalepy: Kernel Density Estimation and Sampling
This package performs KDE operations on multidimensional data to: 1) calculate estimated PDFs (probability distribution functions), and 2) resample new data from those PDFs.
Documentation
A number of examples (also used for continuous integration testing) are included in the package notebooks. Some background information and references are included in the JOSS paper.
Full documentation is available on kalepy.readthedocs.io.
README Contents
Installation
from pypi (i.e. via pip)
pip install kalepy
from source (e.g. for development)
git clone https://github.com/lzkelley/kalepy.git
pip install -e kalepy/
In this case the package can easily be updated by changing into the source directory, pulling, and rebuilding:
cd kalepy
git pull
pip install -e .
# Optional: run unit tests (using the `pytest` package)
pytest
Basic Usage
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import kalepy as kale
from kalepy.plot import nbshow
Generate some random data, and its corresponding distribution function
NUM = int(1e4)
np.random.seed(12345)
# Combine data from two different PDFs
_d1 = np.random.normal(4.0, 1.0, NUM)
_d2 = np.random.lognormal(0, 0.5, size=NUM)
data = np.concatenate([_d1, _d2])
# Calculate the "true" distribution
xx = np.linspace(0.0, 7.0, 100)[1:]
yy = 0.5*np.exp(-(xx - 4.0)**2/2) / np.sqrt(2*np.pi)
yy += 0.5 * np.exp(-np.log(xx)**2/(2*0.5**2)) / (0.5*xx*np.sqrt(2*np.pi))
Plotting Smooth Distributions
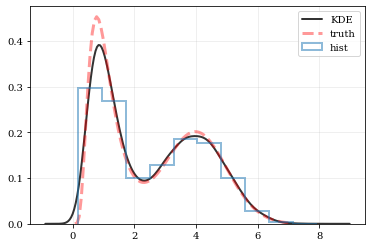
# Reconstruct the probability-density based on the given data points.
points, density = kale.density(data, probability=True)
# Plot the PDF
plt.plot(points, density, 'k-', lw=2.0, alpha=0.8, label='KDE')
# Plot the "true" PDF
plt.plot(xx, yy, 'r--', alpha=0.4, lw=3.0, label='truth')
# Plot the standard, histogram density estimate
plt.hist(data, density=True, histtype='step', lw=2.0, alpha=0.5, label='hist')
plt.legend()
nbshow()
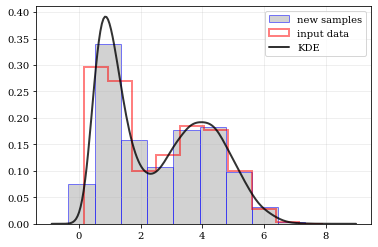
resampling: constructing statistically similar values
Draw a new sample of data-points from the KDE PDF
# Draw new samples from the KDE reconstructed PDF
samples = kale.resample(data)
# Plot new samples
plt.hist(samples, density=True, label='new samples', alpha=0.5, color='0.65', edgecolor='b')
# Plot the old samples
plt.hist(data, density=True, histtype='step', lw=2.0, alpha=0.5, color='r', label='input data')
# Plot the KDE reconstructed PDF
plt.plot(points, density, 'k-', lw=2.0, alpha=0.8, label='KDE')
plt.legend()
nbshow()
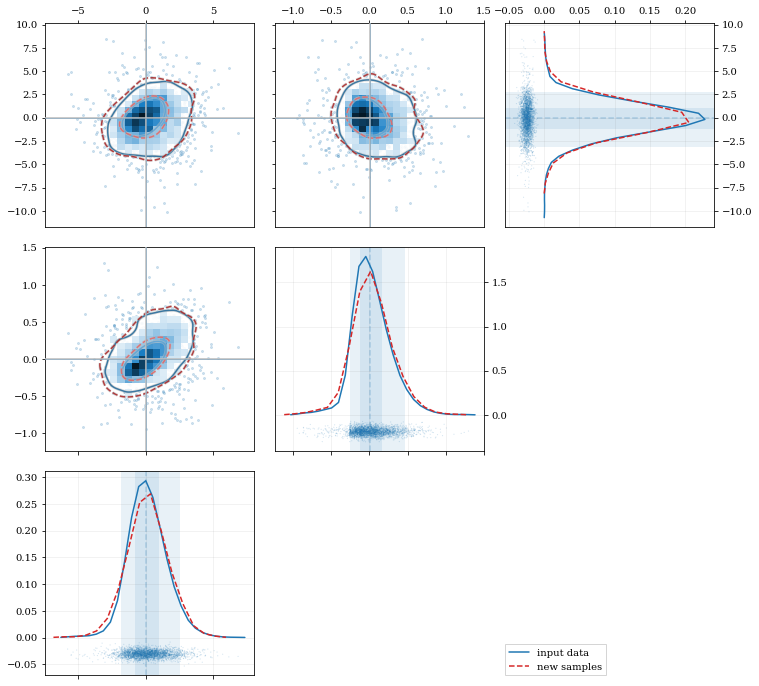
Multivariate Distributions
reload(kale.plot)
# Load some random-ish three-dimensional data
np.random.seed(9485)
data = kale.utils._random_data_3d_02(num=3e3)
# Construct a KDE
kde = kale.KDE(data)
# Construct new data by resampling from the KDE
resamp = kde.resample(size=1e3)
# Plot the data and distributions using the builtin `kalepy.corner` plot
corner, h1 = kale.corner(kde, quantiles=[0.5, 0.9])
h2 = corner.clean(resamp, quantiles=[0.5, 0.9], dist2d=dict(median=False), ls='--')
corner.legend([h1, h2], ['input data', 'new samples'])
nbshow()
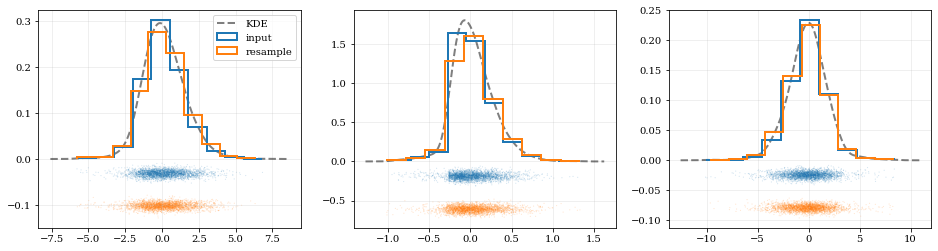
# Resample the data (default output is the same size as the input data)
samples = kde.resample()
# ---- Plot the input data compared to the resampled data ----
fig, axes = plt.subplots(figsize=[16, 4], ncols=kde.ndim)
for ii, ax in enumerate(axes):
# Calculate and plot PDF for `ii`th parameter (i.e. data dimension `ii`)
xx, yy = kde.density(params=ii, probability=True)
ax.plot(xx, yy, 'k--', label='KDE', lw=2.0, alpha=0.5)
# Draw histograms of original and newly resampled datasets
*_, h1 = ax.hist(data[ii], histtype='step', density=True, lw=2.0, label='input')
*_, h2 = ax.hist(samples[ii], histtype='step', density=True, lw=2.0, label='resample')
# Add 'kalepy.carpet' plots showing the data points themselves
kale.carpet(data[ii], ax=ax, color=h1[0].get_facecolor())
kale.carpet(samples[ii], ax=ax, color=h2[0].get_facecolor(), shift=ax.get_ylim()[0])
axes[0].legend()
nbshow()
Fancy Usage
Reflecting Boundaries
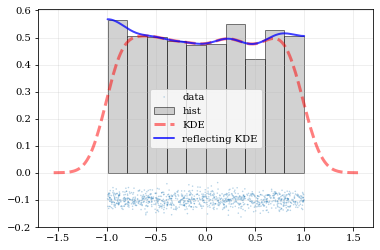
What if the distributions you're trying to capture have edges in them, like in a uniform distribution between two bounds? Here, the KDE chooses 'reflection' locations based on the extrema of the given data.
# Uniform data (edges at -1 and +1)
NDATA = 1e3
np.random.seed(54321)
data = np.random.uniform(-1.0, 1.0, int(NDATA))
# Create a 'carpet' plot of the data
kale.carpet(data, label='data')
# Histogram the data
plt.hist(data, density=True, alpha=0.5, label='hist', color='0.65', edgecolor='k')
# ---- Standard KDE will undershoot just-inside the edges and overshoot outside edges
points, pdf_basic = kale.density(data, probability=True)
plt.plot(points, pdf_basic, 'r--', lw=3.0, alpha=0.5, label='KDE')
# ---- Reflecting KDE keeps probability within the given bounds
# setting `reflect=True` lets the KDE guess the edge locations based on the data extrema
points, pdf_reflect = kale.density(data, reflect=True, probability=True)
plt.plot(points, pdf_reflect, 'b-', lw=2.0, alpha=0.75, label='reflecting KDE')
plt.legend()
nbshow()
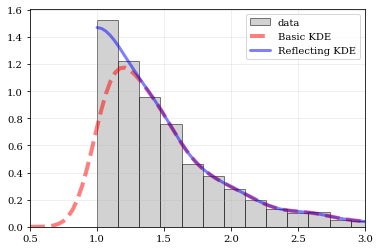
Explicit reflection locations can also be provided (in any number of dimensions).
# Construct random data, add an artificial 'edge'
np.random.seed(5142)
edge = 1.0
data = np.random.lognormal(sigma=0.5, size=int(3e3))
data = data[data >= edge]
# Histogram the data, use fixed bin-positions
edges = np.linspace(edge, 4, 20)
plt.hist(data, bins=edges, density=True, alpha=0.5, label='data', color='0.65', edgecolor='k')
# Standard KDE with over & under estimates
points, pdf_basic = kale.density(data, probability=True)
plt.plot(points, pdf_basic, 'r--', lw=4.0, alpha=0.5, label='Basic KDE')
# Reflecting KDE setting the lower-boundary to the known value
# There is no upper-boundary when `None` is given.
points, pdf_basic = kale.density(data, reflect=[edge, None], probability=True)
plt.plot(points, pdf_basic, 'b-', lw=3.0, alpha=0.5, label='Reflecting KDE')
plt.gca().set_xlim(edge - 0.5, 3)
plt.legend()
nbshow()
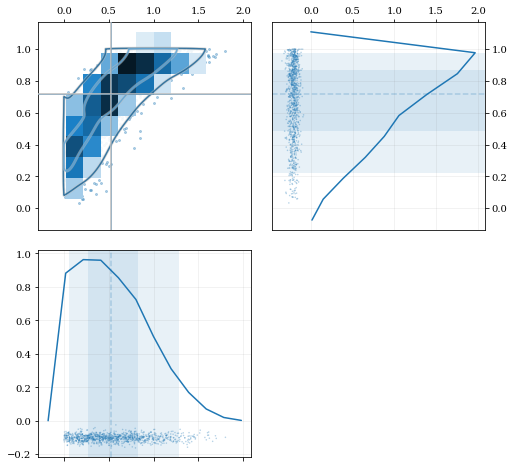
Multivariate Reflection
# Load a predefined dataset that has boundaries at:
# x: 0.0 on the low-end
# y: 1.0 on the high-end
data = kale.utils._random_data_2d_03()
# Construct a KDE with the given reflection boundaries given explicitly
kde = kale.KDE(data, reflect=[[0, None], [None, 1]])
# Plot using default settings
kale.corner(kde)
nbshow()
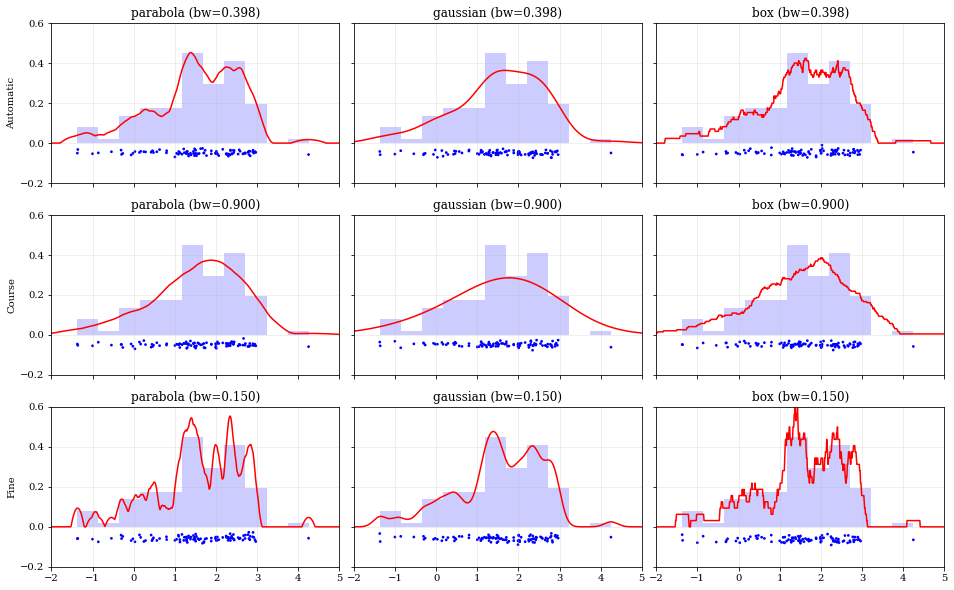
Specifying Bandwidths and Kernel Functions
# Load predefined 'random' data
data = kale.utils._random_data_1d_02(num=100)
# Choose a uniform x-spacing for drawing PDFs
xx = np.linspace(-2, 8, 1000)
# ------ Choose the kernel-functions and bandwidths to test ------- #
kernels = ['parabola', 'gaussian', 'box'] #
bandwidths = [None, 0.9, 0.15] # `None` means let kalepy choose #
# ----------------------------------------------------------------- #
ylabels = ['Automatic', 'Course', 'Fine']
fig, axes = plt.subplots(figsize=[16, 10], ncols=len(kernels), nrows=len(bandwidths), sharex=True, sharey=True)
plt.subplots_adjust(hspace=0.2, wspace=0.05)
for (ii, jj), ax in np.ndenumerate(axes):
# ---- Construct KDE using particular kernel-function and bandwidth ---- #
kern = kernels[jj] #
bw = bandwidths[ii] #
kde = kale.KDE(data, kernel=kern, bandwidth=bw) #
# ---------------------------------------------------------------------- #
# If bandwidth was set to `None`, then the KDE will choose the 'optimal' value
if bw is None:
bw = kde.bandwidth[0, 0]
ax.set_title('{} (bw={:.3f})'.format(kern, bw))
if jj == 0:
ax.set_ylabel(ylabels[ii])
# plot the KDE
ax.plot(*kde.pdf(points=xx), color='r')
# plot histogram of the data (same for all panels)
ax.hist(data, bins='auto', color='b', alpha=0.2, density=True)
# plot carpet of the data (same for all panels)
kale.carpet(data, ax=ax, color='b')
ax.set(xlim=[-2, 5], ylim=[-0.2, 0.6])
nbshow()
Resampling
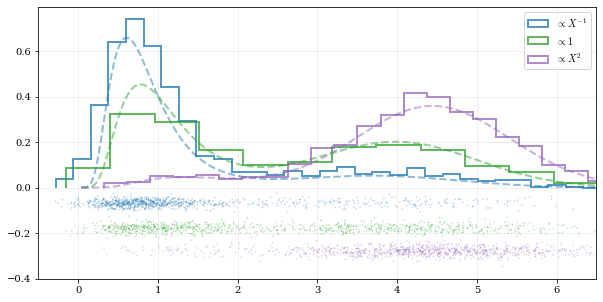
Using different data weights
# Load some random data (and the 'true' PDF, for comparison)
data, truth = kale.utils._random_data_1d_01()
# ---- Resample the same data, using different weightings ---- #
resamp_uni = kale.resample(data, size=1000) #
resamp_sqr = kale.resample(data, weights=data**2, size=1000) #
resamp_inv = kale.resample(data, weights=data**-1, size=1000) #
# ------------------------------------------------------------ #
# ---- Plot different distributions ----
# Setup plotting parameters
kw = dict(density=True, histtype='step', lw=2.0, alpha=0.75, bins='auto')
xx, yy = truth
samples = [resamp_inv, resamp_uni, resamp_sqr]
yvals = [yy/xx, yy, yy*xx**2/10]
labels = [r'$\propto X^{-1}$', r'$\propto 1$', r'$\propto X^2$']
plt.figure(figsize=[10, 5])
for ii, (res, yy, lab) in enumerate(zip(samples, yvals, labels)):
hh, = plt.plot(xx, yy, ls='--', alpha=0.5, lw=2.0)
col = hh.get_color()
kale.carpet(res, color=col, shift=-0.1*ii)
plt.hist(res, color=col, label=lab, **kw)
plt.gca().set(xlim=[-0.5, 6.5])
# Add legend
plt.legend()
# display the figure if this is a notebook
nbshow()
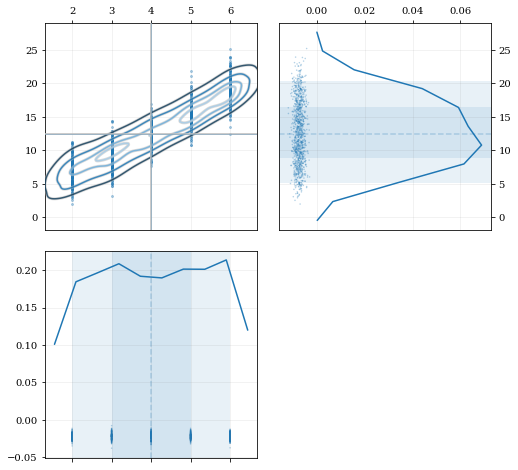
Resampling while 'keeping' certain parameters/dimensions
# Construct covariant 2D dataset where the 0th parameter takes on discrete values
xx = np.random.randint(2, 7, 1000)
yy = np.random.normal(4, 2, xx.size) + xx**(3/2)
data = [xx, yy]
# 2D plotting settings: disable the 2D histogram & disable masking of dense scatter-points
dist2d = dict(hist=False, mask_dense=False)
# Draw a corner plot
kale.corner(data, dist2d=dist2d)
nbshow()
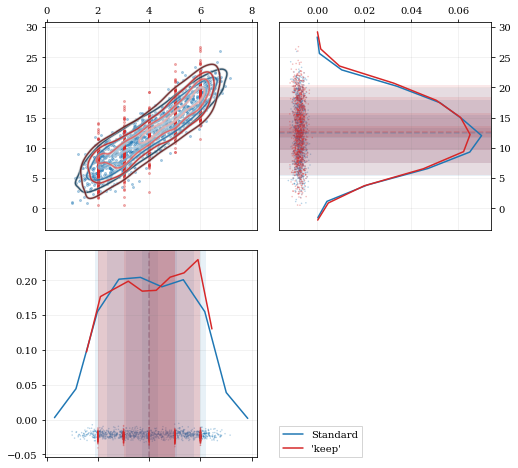
A standard KDE resampling will smooth out the discrete variables, creating a smooth(er) distribution. Using the keep parameter, we can choose to resample from the actual data values of that parameter instead of resampling with 'smoothing' based on the KDE.
kde = kale.KDE(data)
# ---- Resample the data both normally, and 'keep'ing the 0th parameter values ---- #
resamp_stnd = kde.resample() #
resamp_keep = kde.resample(keep=0) #
# --------------------------------------------------------------------------------- #
corner = kale.Corner(2)
dist2d['median'] = False # disable median 'cross-hairs'
h1 = corner.plot(resamp_stnd, dist2d=dist2d)
h2 = corner.plot(resamp_keep, dist2d=dist2d)
corner.legend([h1, h2], ['Standard', "'keep'"])
nbshow()
Development & Contributions
Please visit the github page <https://github.com/lzkelley/kalepy>_ for issues or bug reports. Contributions and feedback are very welcome.
Contributors:
- Luke Zoltan Kelley (@lzkelley)
- Zachary Hafen (@zhafen)
JOSS Paper:
- Kexin Rong (@kexinrong)
- Arfon Smith (@arfon)
- Will Handley (@williamjameshandley)
Attribution
A JOSS paper has been submitted. If you have found this package useful in your research, please add a reference to the code paper:
.. code-block:: tex
@article{kalepy,
author = {Luke Zoltan Kelley},
title = {kalepy: a python package for kernel density estimation and sampling},
journal = {The Journal of Open Source Software},
publisher = {The Open Journal},
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file kalepy-1.4.3.tar.gz.
File metadata
- Download URL: kalepy-1.4.3.tar.gz
- Upload date:
- Size: 54.4 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.9.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dcac1fa78db55f88222eac655d6b11cecb63f61b5a37483aff48cb29374a2da8
|
|
| MD5 |
8a7a7c12bf140dfaa14b8ae015f91d49
|
|
| BLAKE2b-256 |
a27a801732537622f20230d4b062f1af1b1c709c7757bf9b50abdb4ed9e4923e
|
File details
Details for the file kalepy-1.4.3-py3-none-any.whl.
File metadata
- Download URL: kalepy-1.4.3-py3-none-any.whl
- Upload date:
- Size: 555.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.9.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
43647be31a811c5c59deb21bb5017e1a85a00c06019d38b2d4e0cc192b37deb8
|
|
| MD5 |
59bd43fda5db7cd13d619b4280dfcb66
|
|
| BLAKE2b-256 |
93f1e8b3905c7ba534aaccca69ee8ac1d38c80ceab7c3e08f70aea46da24192d
|