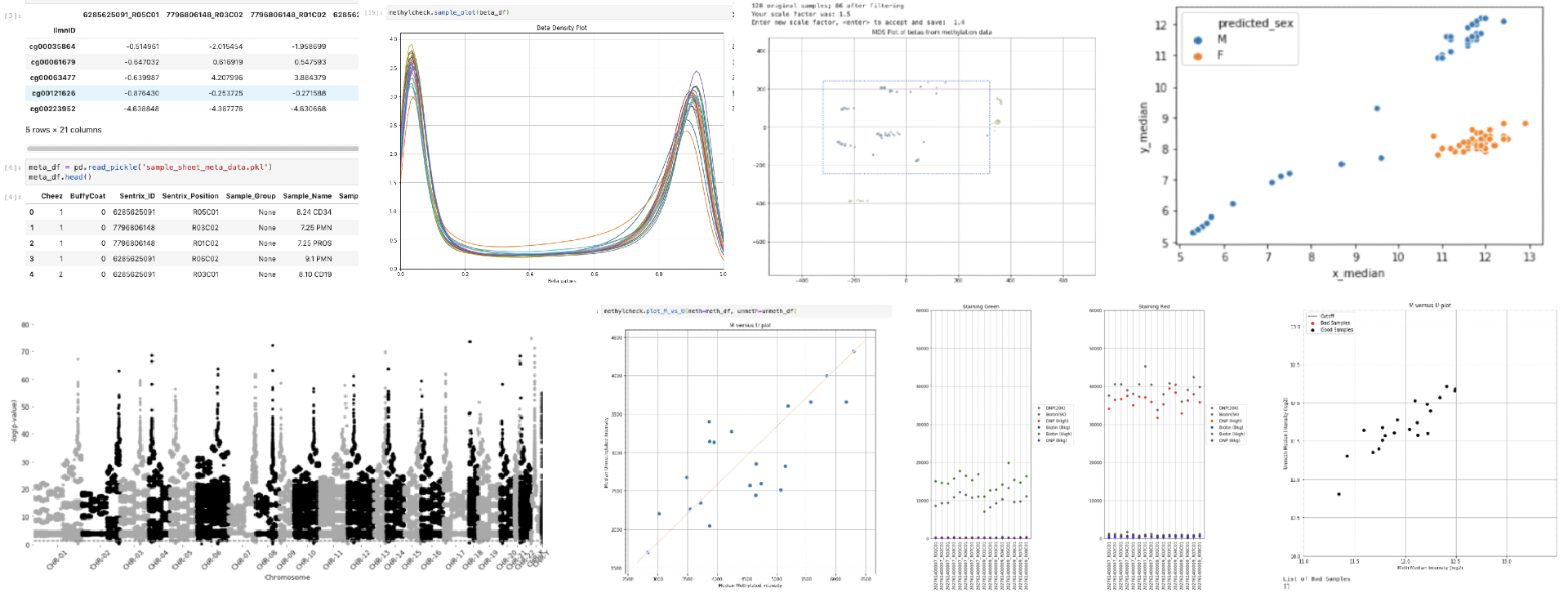
Quality Control (QC), Visualization/plotting, and postprocessing software for Illumina methylation array data. See https://life-epigenetics-methylcheck.readthedocs-hosted.com/en/latest/ for full documentation and examples.
Project description
methylcheck is a Python-based package for filtering and visualizing Illumina methylation array data. The focus is on quality control. View on ReadTheDocs.
methylcheck is part of the methylsuite
methylcheck is part of the methylsuite of python packages that provide functions to process and analyze DNA methylation data from Illumina's Infinium arrays (27k, 450k, and EPIC, as well as mouse arrays). This package is focused on quality control for processed methylation data.
methylcheck functions are designed to work with a minimum of knowledge and specification required. But you can always override the "smart" defaults with custom settings if the default settings don't work for your data. The entire methylsuite is designed in this format: to offer ease of use while still maintaining flexibility for customization as needed.
Methylsuite package components
You should install all three components, as they work together. The parts include:
-
methylprep: for processingidatfiles or downloading GEO datasets from NIH. Processing steps include- infer type-I channel switch
- NOOB (normal-exponential convolution on out-of-band probe data)
- poobah (p-value with out-of-band array hybridization, for filtering low signal-to-noise probes)
- qualityMask (to exclude historically less reliable probes)
- nonlinear dye bias correction (AKA signal quantile normalization between red/green channels across a sample)
- calculate beta-value, m-value, or copy-number matrix
- large batch memory management, by splitting it up into smaller batches during processing
-
methylcheck: (this package) for quality control (QC) and analysis, including- functions for filtering out unreliable probes, based on the published literature
- Note that
methylprep processwill exclude a set of unreliable probes by default. You can disable that using the --no_quality_mask option from CLI.
- Note that
- sample outlier detection
- array level QC plots of staining, bisulfite conversion, hybridization, extension, negative, non-polymorphic, target removal, and specificity
- spreadsheet summary of control probe performance
- data visualization functions based on
seabornandmatplotlibgraphic libraries. - predict sex of human samples from probes
- interactive method for assigning samples to groups, based on array data, in a Jupyter notebook
- functions for filtering out unreliable probes, based on the published literature
-
methylizeprovides more analysis and interpretation functions- differentially methylated probe statistics (between treatment and control samples)
- differentially methylated regions, with gene annotation from the UCSC Human Genome Browser
- volcano plots (to identify probes that are the most different)
- manhattan plots (to identify clusters of probes associated with genomic regions that are different)
Installation
methylcheck maintains configuration files for your Python package manager of choice: pipenv or pip. Conda install is coming soon.
>>> pip install methylcheck
or if you want to install all three packages at once (recommended):
>>> pip install methylsuite
Tutorials and Guides
If you are new to DNA methylation analysis, we recommend reading through this introduction from the methylprep documentation. Otherwise, you are ready to use methylcheck to:
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file methylcheck-0.8.5.tar.gz.
File metadata
- Download URL: methylcheck-0.8.5.tar.gz
- Upload date:
- Size: 37.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/3.10.0 pkginfo/1.7.0 requests/2.27.1 requests-toolbelt/0.9.1 tqdm/4.62.3 CPython/3.8.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c7c2cc6cb49d353c2b4ff7318361ac4642e7c38e7df5ceb61240cf0b2261d3ef
|
|
| MD5 |
f23b9e3a74bb57b8191d4e8d8cd0585d
|
|
| BLAKE2b-256 |
2a51db1d671678e0d825d2e7c129f90275ce64e1b2720785b8942b7dcd11c849
|
File details
Details for the file methylcheck-0.8.5-py3-none-any.whl.
File metadata
- Download URL: methylcheck-0.8.5-py3-none-any.whl
- Upload date:
- Size: 10.8 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/3.10.0 pkginfo/1.7.0 requests/2.27.1 requests-toolbelt/0.9.1 tqdm/4.62.3 CPython/3.8.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d872b6c03c3acde1b51add42e8953b85a9162446620857d86bca6db1f7c1894f
|
|
| MD5 |
a082f536ef56f10b84dee834bef8b902
|
|
| BLAKE2b-256 |
61ab010136c59b9ba6f742926f19fdfc8e928fd01ad87906700e5591c49de3b6
|