Neural network modeled after the olfactory system of the hawkmoth.
Project description
pymoth
This package contains a Python version of MothNet

Neural network modeled after the olfactory system of the hawkmoth, Manduca sexta (shown above).
This repository contains a Python version of the code used in:
- "Putting a bug in ML: The moth olfactory network learns to read MNIST", Neural Networks 2019
Docs (via Sphinx)
Installation
Built for use with Mac/Linux systems - not tested in Windows.
- Requires Python 3.6+
Via pip
pip install mothnet
From source
First, clone this repo and cd into it. Then run:
# Install dependencies:
pip install -r pymoth/docs/requirements.txt
# Run sample experiment:
python pymoth/examples.py
Dependencies (also see requirements.txt)
- scipy
- matplotlib
- scikit-learn(for kNN and SVM models)
- pillow
- keras (for loading MNIST)
- tensorflow (also for loading MNIST)
Example experiment (also see examples.py)
import os
import pymoth
def experiment():
# instantiate the MothNet object
mothra = pymoth.MothNet({
'screen_size': (1920, 1080), # screen size (width, height)
'num_runs': 1, # how many runs you wish to do with this moth
'goal': 15, # defines the moth's learning rates
'tr_per_class': 1, # (try 3) the number of training samples per class
'num_sniffs': 1, # (try 2) number of exposures each training sample
'num_neighbors': 1, # optimization param for nearest neighbors
'box_constraint': 1e1, # optimization parameter for svm
'n_thumbnails': 1, # show N experiment inputs from each class
'show_acc_plots': True, # True to plot, False to ignore
'show_time_plots': True, # True to plot, False to ignore
'show_roc_plots': True, # True to plot, False to ignore
'results_folder': 'results', # string
'results_filename': 'results', # will get the run number appended to it
'data_folder': 'MNIST_all', # string
'data_filename': 'MNIST_all', # string
})
# loop through the number of simulations specified:
for run in range(mothra.NUM_RUNS):
# generate dataset
feature_array = mothra.load_mnist()
train_X, test_X, train_y, test_y = mothra.train_test_split(feature_array)
# load parameters
mothra.load_moth() # define moth model parameters
mothra.load_exp() # define parameters of a time-evolution experiment
# run simulation (SDE time-step evolution)
sim_results = mothra.simulate(feature_array)
# future: mothra.fit(X_train, y_train)
# collect response statistics:
# process the sim results to group EN responses by class and time
EN_resp_trained = mothra.collect_stats(sim_results, mothra.experiment_params,
mothra._class_labels, mothra.SHOW_TIME_PLOTS, mothra.SHOW_ACC_PLOTS,
images_filename=mothra.RESULTS_FILENAME, images_folder=mothra.RESULTS_FOLDER,
screen_size=mothra.SCREEN_SIZE)
# reveal scores
# score MothNet
mothra.score_moth_on_MNIST(EN_resp_trained)
# score KNN
mothra.score_knn(train_X, train_y, test_X, test_y)
# score SVM
mothra.score_svm(train_X, train_y, test_X, test_y)
# plot each model in a subplot of a single figure
if mothra.SHOW_ROC_PLOTS:
mothra.show_multi_roc(['MothNet', 'SVM', 'KNN'], mothra._class_labels,
images_filename=mothra.RESULTS_FOLDER+os.sep+mothra.RESULTS_FILENAME+'_ROC_multi')
Sample results
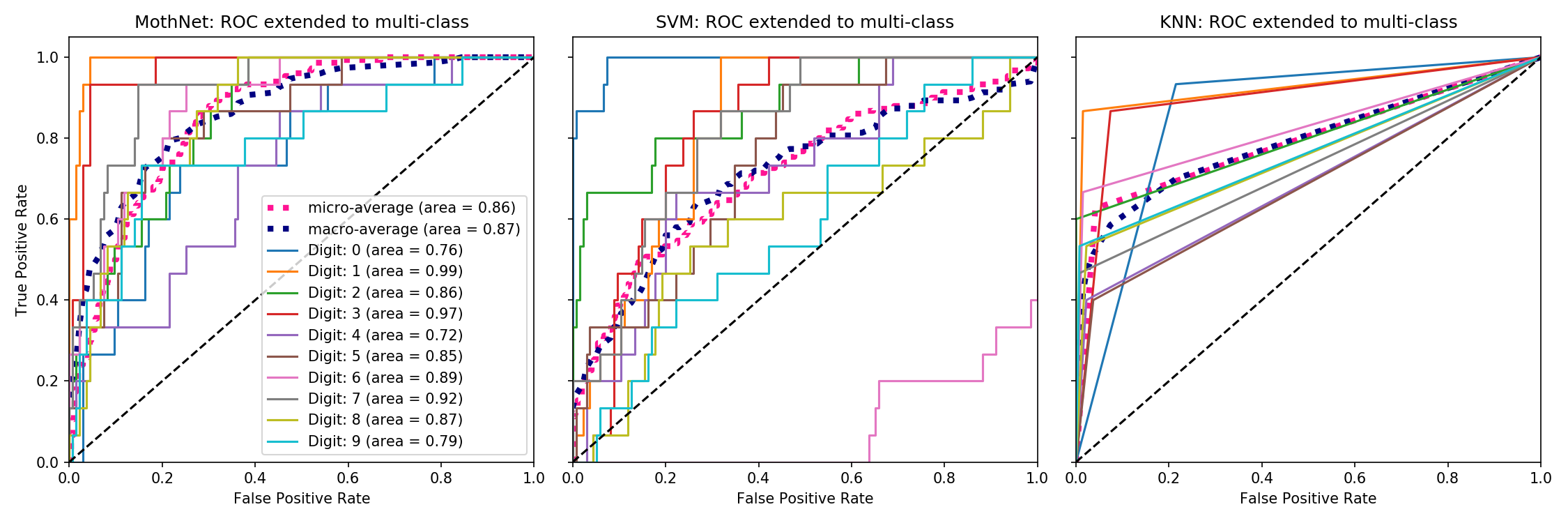
Dataset
Modules
- classify.py Classify output from MothNet model.
- generate.py Download (if absent) and prepare down-sampled MNIST dataset.
- params.py Experiment and model parameters.
- sde.py Run stochastic differential equation simulation.
- show_figs.py Figure generation module.
- MNIST_make_all.py Downloads and saves MNIST data to .npy file.
Questions, comments, criticisms? Feel free to drop us an e-mail!
Bug reports, suggestions, or requests are also welcome! Feel free to create an issue.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file mothnet-0.0.5.tar.gz.
File metadata
- Download URL: mothnet-0.0.5.tar.gz
- Upload date:
- Size: 44.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.13.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.1.0 requests-toolbelt/0.9.1 tqdm/4.33.0 CPython/3.7.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
afefec33fdff9d13a74538c7fc70bb1b89df4a7c36083e23ef950ca215d8f6e4
|
|
| MD5 |
a32f5732082257a4c19cb104a6ed2c24
|
|
| BLAKE2b-256 |
579c0f25210d2189391c4a246e6fb0fc2ffee7b8723fd091d9290da9e2a80900
|
File details
Details for the file mothnet-0.0.5-py3-none-any.whl.
File metadata
- Download URL: mothnet-0.0.5-py3-none-any.whl
- Upload date:
- Size: 49.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.13.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.1.0 requests-toolbelt/0.9.1 tqdm/4.33.0 CPython/3.7.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
29430ceb75d6a3852ee8efa5f65432d8975980946d3794a8dd07cd19f3935b66
|
|
| MD5 |
e49c288f8d57723d4874bbca817c9405
|
|
| BLAKE2b-256 |
d5e339593fadd0ea2ccff1c36b3eb300db8e65d26b71b108d4ad0b5f230ce607
|














