A SciUnit library for validation testing of spiking networks.
Project description
A SciUnit library for validation testing of spiking networks.

Installation
pip install networkunitConcept
The NetworkUnit module builds upon the formalized validation scheme of the SciUnit package,
which enables the validation of models against experimental data (or other models) via tests.
A test is matched to the model by capabilities and quantitatively evaluated by a score.
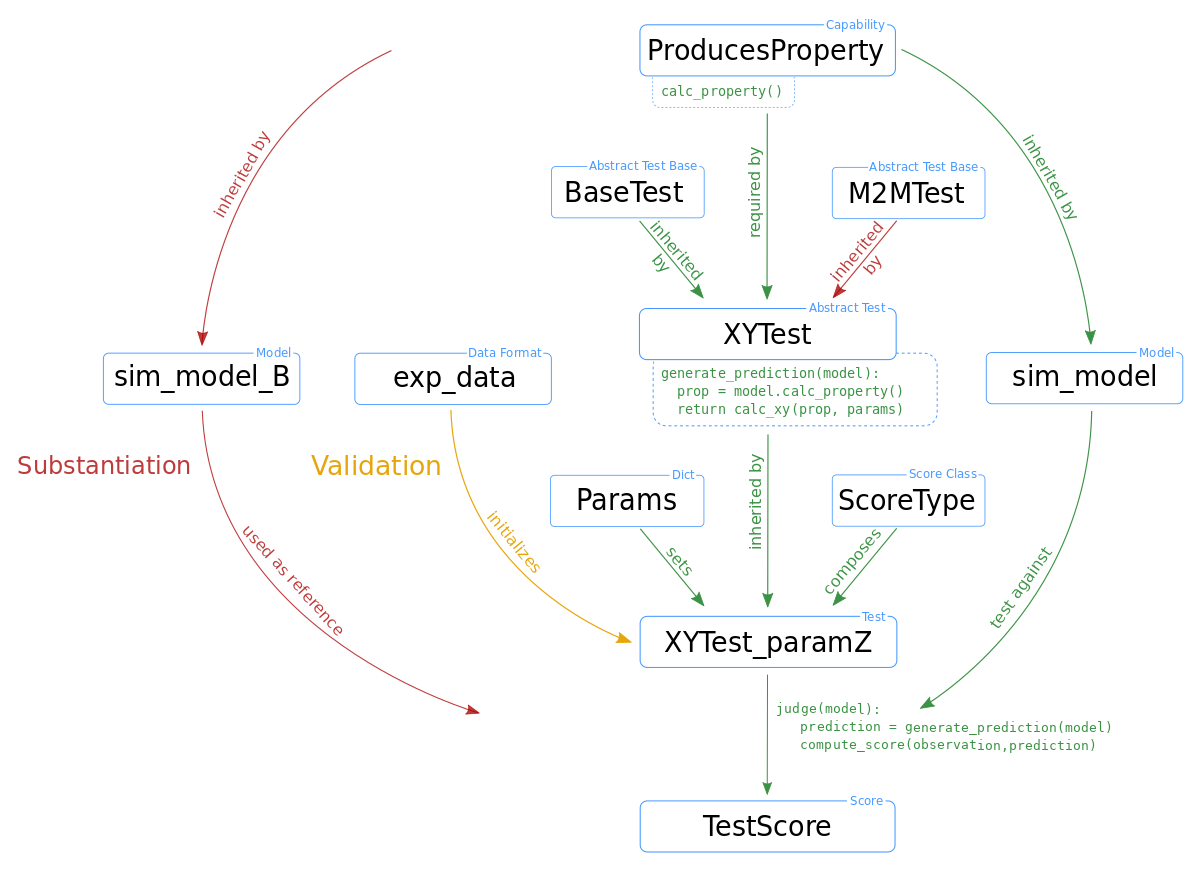
The following figure illustrates a typical test design within NetworkUnit.
The blue boxes indicate the components of the implementation of the validation test, i.e.,
classes, class instances, data sets, and parameters.
The relation between the boxes are indicated by annotated arrows.The basic functionality is
shown by green arrows. The difference in the test design for comparing against experimental
data (validation) and another simulation (substantiation) is indicated by yellow and
red arrows, respectively. The relevant functionality of some components for the
computation of test score is indicated by pseudo-code. The capability
class ProducesProperty contains the function calc_property(). The test XYTest has a function
generate_prediction() which makes use of this capability, inherited by the model class,
to generate a model prediction. The initialized test instance XYTest_paramZ makes use of its
judge() function to evaluate this model prediction and compute the score TestScore.
The XYTest can inherit from multiple abstract test classes (BaseTest),
which is for example used with the M2MTest to add the functionality of evaluating multiple model classes.
To make the test executable it has to be linked to a ScoreType and all free parameters need to be set
(by a Params dict) to ensure a reproducible result.
Showcase examples on how to use NetworkUnit can be found in this repository and interactive reveal.js slides are accessible via the launch-binder button at the top.
Overview of tests
Class name |
Parent class |
Prediction measure |
|---|---|---|
two_sample_test |
- |
- |
firing_rate_test |
two_sample_test |
firing rates |
isi_variation_test |
two_sample_test |
inter-spike-intervals, their CV, CV2, LV, or LVR |
covariance_test |
two_sample_test |
covariances |
correlation_test |
two_sample_test |
- |
correlation_dist_test |
correlation_test |
correlation coefficients |
correlation_matrix_test |
correlation_test |
correlation coefficient matrix |
avg_std_correlation_test |
correlation_test |
avg or std of correlation coefficients |
generalized_correlation_matrix_test |
correlation_matrix_test |
matrix of derived cross-correlation measures |
eigenvalue_test |
correlation_test |
eigenvalues of the correlation coefficient matrix |
power_spectrum_test |
two_sample_test |
power spectral density values |
freqband_power_test |
power_spectrum_test |
power spectral density values in frequency band |
timescale_test |
two_sample_test |
spike train timescale |
graph_centrality_helperclass |
sciunit.Test |
graph centrality measures of given adjacency matrix |
joint_test |
two_sample_test |
joining other test classes for multivariate evaluation |
Inheritance order in case of multiple inheritance for derived test classes:
class new_test(sciunit.TestM2M, graph_centrality_helperclass, <base_test_class>)Overview of scores
Class name |
Test name |
Comparison measure |
|---|---|---|
students_t |
Student’t test |
sample mean |
ks_distance |
Kolmogorov-Smirnov test |
sample distribution |
kl_divergence |
Kullback-Leibler divergence |
sample entropy |
mwu_statistic |
Mann-Whitney U test |
rank sum |
levene_score |
Levene’s test |
sample variance |
effect_size |
Effect size |
standardized mean |
best_effect_size |
Bayesian estimation effect size |
standardized mean |
wasserstein_distance |
Wasserstein Distance |
multivariate score distance |
eigenangle |
Eigenangle Test |
eigenangle similarity |
Overview of model classes
Model name |
Capability |
Parent class |
Purpose |
|---|---|---|---|
loaded_spiketrains |
ProducesSpikeTrains |
sciunit.RunnableModel |
loading simulated spiking data |
stochastic_activity |
ProducesSpikeTrains |
sciunit.RunnableModel |
generating stochastic spiking data |
nest_simulation |
ProducesSpikeTrains |
sciunit.RunnableModel |
template for NEST models |
Other validation test repositories
Acknowledgments
This open source software code was developed in part or in whole in the Human Brain Project, funded from the European Union’s Horizon 2020 Framework Programme for Research and Innovation under Specific Grant Agreements No. 720270, No. 785907 and No. 945539 (Human Brain Project SGA1, SGA2 and SGA3).
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file networkunit-0.2.2.tar.gz.
File metadata
- Download URL: networkunit-0.2.2.tar.gz
- Upload date:
- Size: 45.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.8.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f0ada132cd73078a5ea6ddb60e2c642e6d59355cd90fbdd57cb233116f58f929
|
|
| MD5 |
43119a192b3f87731956e6ebc869e6a5
|
|
| BLAKE2b-256 |
4d717f9b5480d2338c2b639dd7a4a54a9abd7f4f3c7f17cb7b57ebaed287aab2
|
File details
Details for the file networkunit-0.2.2-py3-none-any.whl.
File metadata
- Download URL: networkunit-0.2.2-py3-none-any.whl
- Upload date:
- Size: 67.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.8.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ca24428cb64d4b83102f0bd0b29ec2477246fb833782f4b441837bea486c83e3
|
|
| MD5 |
f0aa9393d6bcc786228a19f970795690
|
|
| BLAKE2b-256 |
0f02d866a71236226b3821709691dc2ef0791cb283c2347130ba0aba01033f48
|












