Python Framework for Materials Discovery and Design
Project description
PyChemia: Python Framework for Materials Discovery and Design
PyChemia is an open-source Python Library for materials structural search. The purpose of the initiative is to create a method agnostic framework for materials discovery and design using a variety of methods from Minima Hoping to Soft-computing based methods. PyChemia is also a library for data-mining, using several methods to discover interesting candidates among the materials already processed.
The core of the library is the Structure python class, it is able to describe periodic and non-periodic structures. As the focus of this library is structural search the class defines extensive capabilities to modify atomic structures.
The library includes capability to read and write in several ab-initio codes. At the level of DFT, PyChemia support VASP, ABINIT and Octopus. At Tight-binding level development is in process to support DFTB+ and Fireball. This allows the library to compute electronic-structure properties using state-of-the-art ab-initio software packages and extract properties from those calculations.
Installation
You can install pychemia in several ways. We are showing 3 ways of installing PyChemia inside a Virtual environment. A virtual environment is a good way of isolating software packages from the pacakges installed with the Operating System. The decision on which method to use depends if you want to use the most recent code or the package uploaded from time to time to PyPi. The last method is particularly suited for developers who want to change the code and get those changes operative without an explicit instalation.
Installing with pip from pypi.org on a virtual environment
This method installs PyChemia from the packages uploaded to PyPi every month. It will provides a version of PyChemia that is stable.
First, create and activate the virtual environment. We are using the name pychemia_ve, but that is arbitrary.
python3 -m venv pychemia_ve source pychemia_ve/bin/activate
When the virtual environment is activated, your prompt changes to (pychemia_ve)...$. Now, install pychemia with pip
python3 -m pip install pychemia
Installing with pip from a cloned repo on a virtual environment
This method installs PyChemia from the Github repo. The method will install PyChemia from the most recent sources.
First, create and activate the virtual environment. We are using the name pychemia_ve, but that is arbitrary.
python3 -m venv pychemia_ve cd pychemia_ve source bin/activate
Second, clone the repository from GitHub
git clone https://github.com/MaterialsDiscovery/PyChemia.git
Finally, install from the repo folder
python3 -m pip install PyChemia
Using PyChemia from repo folder on a virtual environment
This method is mostly used for development. In this way PyChemia is not actually installed and changes to the code will take inmediate effect.
First, create and activate the virtual environment. We are using the name pychemia_ve, but that is arbitrary.
python3 -m venv pychemia_ve cd pychemia_ve source bin/activate
Clone the repository
git clone https://github.com/MaterialsDiscovery/PyChemia.git
Go to repo folder, install Cython with pip and execute setup.py to build the Cython modules.
cd PyChemia python3 -m pip install Cython python3 setup.py build_ext --inplace python3 setup.py build
Finally, install the packages required for PyChemia to work
python3 -m pip install -r requirements.txt
Set the variable $PYTHONPATH to point to PyChemia folder, in the case of bash it will be:
export PYTHONPATH=`path`
On C shell (csh or tcsh)
setenv PYTHONPATH `path`
To run the internal testsuite install pytest
python3 -m pip install pytest
Execute the testsuite:
pytest
The output of the testsuite looks like this:
=================== test session starts =========================== platform linux -- Python 3.11.3, pytest-7.3.0, pluggy-1.0.0 rootdir: /gpfs20/scratch/gufranco/PyChemia plugins: hypothesis-6.71.0, anyio-3.6.2 collecting 2 items collecting 29 items collected 67 items tests/test_0.py . [ 1%] tests/test_1_doctest_core.py ... [ 5%] tests/test_1_doctest_crystal.py .. [ 8%] tests/test_1_doctest_utils.py .... [ 14%] tests/test_3_scripts.py .... [ 20%] tests/test_analysis.py ... [ 25%] tests/test_code_abinit.py ..... [ 32%] tests/test_code_fireball.py . [ 34%] tests/test_code_vasp.py ...... [ 43%] tests/test_core.py ...... [ 52%] tests/test_crystal_kpoints.py . [ 53%] tests/test_crystal_symmetry.py . [ 55%] tests/test_db_queue.py . [ 56%] tests/test_io.py .. [ 59%] tests/test_population.py ..... [ 67%] tests/test_population_orbitals.py . [ 68%] tests/test_searcher_clusters.py .... [ 74%] tests/test_searcher_functions.py .... [ 80%] tests/test_searcher_noncollinear.py ... [ 85%] tests/test_utils_metaheuristics.py . [ 86%] tests/test_utils_periodic.py ...... [ 95%] tests/test_utils_serializer.py . [ 97%] tests/test_zexample1.py . [ 98%] tests/test_zexample2.py . [100%] =================== 67 passed in 54.60s ===========================
PyChemia requirements
PyChemia relies on a number of other python packages to operate. Some of them are mandatory and they must be installed. Other packages are optional and their absence will only constrain certain capabilities.
Python >= 3.6 < 3.12
The library is tested using versions of Python from 3.6 up to 3.11
https://travis-ci.org/MaterialsDiscovery/PyChemia
Mandatory
Numpy >= 1.19 Fundamental library for numerical intensive computation in Python. Numpy arrays are essential for efficient array manipulation.
SciPy >= 1.5 Used mostly for Linear Algebra, FFT and spatial routines.
Spglib >= 2.0 Used to determine symmetry groups for periodic structures
Matplotlib >= 3.3 Used to plot band structures, densities of states and other 2D plots
PyMongo >= 4.1 Used for structural search PyChemia relies strongly in MongoDB and its python driver. For the MongoDB server, any version beyond 3.11 should be fine. We have tested pychemia on MongoDB 4.0
psutil >= 5.9 Cross-platform lib for process and system monitoring in Python
netCDF4 > 1.5 Python/numpy interface to the netCDF C library
Optional
pytest >= 7.0 A python library for testing, simply go to the source directory and execute:
pytest -v .
Pandas Library for Data Analysis used by the datamining modules
PyMC PyMC is a python module that implements Bayesian statistical models and fitting algorithms Important for the datamining capabilities of PyChemia
Mayavi >= 4.1 Some basic visualization tools are incorporated using this library
ScientificPython >2.6 This library is used for reading and writing NetCDF files
pymatgen >= 2.9 pymatgen is an excellent library for materials analysis
ASE Atomic Simulation Environment is another good library for ab-initio calculations. Quite impressive for the number of ab-initio packages supported
qmpy The Python library behind the [Open Quantum Materials Database](http://oqmd.org). The OQMD is a database of DFT calculated structures. For the time being the database contains more than 300000 structures, with more than 90% of them with the electronic ground-state computed.
- coverage >= 4.0.1
Provides code coverage analysis
python-coveralls To submit coverage information to coveralls.io
Documentation
Instructions for installation, using and programming scripts with PyChemia can be found on two repositories for documentation:
Read The Docs:
Python Hosted:
Documentation is hosted on Read the Docs also available with Short URLs readthedocs and rtfd
Documentation is also hosted on Python Hosted
Sources
The main repository is on GitHub
Sources and wheel binaries are also distrubuted on PyPI Python.org or PyPI pypi.org
Structure of the Library
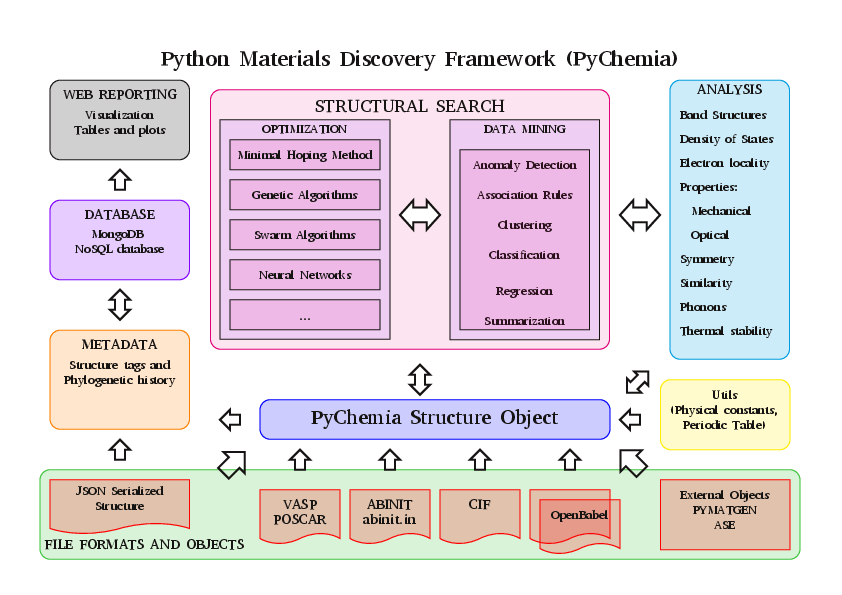
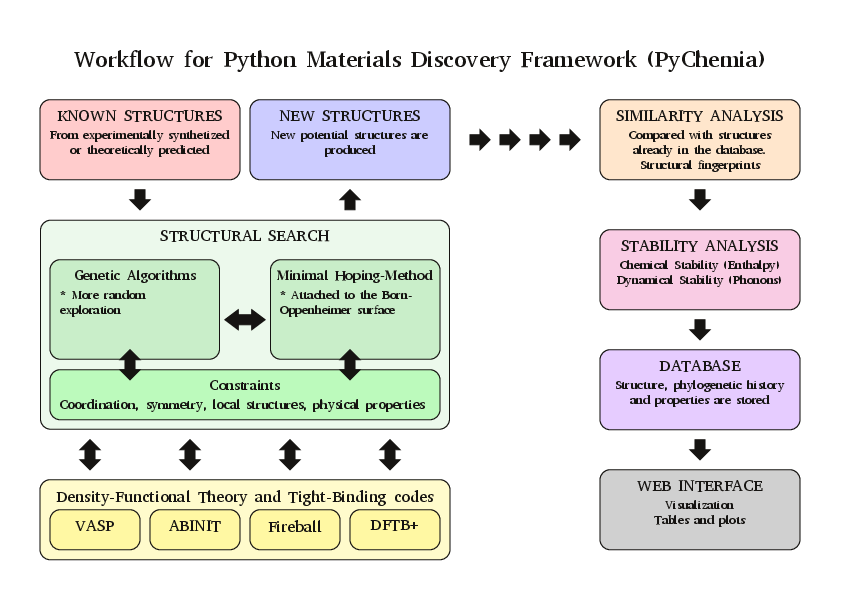
Contributors
Prof. Aldo H. Romero [West Virginia University] (Project Director)
Guillermo Avendaño-Franco [West Virginia University] (Basic Infrastructure)
Adam Payne [West Virginia University] (Bug fixes (Populations, Relaxators, and KPoints) )
Irais Valencia Jaime [West Virginia University] (Simulation and testing)
Sobhit Singh [West Virginia University] (Data-mining)
Francisco Muñoz [Universidad de Chile] (PyPROCAR)
Wilfredo Ibarra Hernandez [West Virginia University] (Interface with MAISE)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file pychemia-0.24.2.3.tar.gz.
File metadata
- Download URL: pychemia-0.24.2.3.tar.gz
- Upload date:
- Size: 35.7 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
29237b04c2433b4847519ab44a83511537cf24e44b21c5e8ad4ceb981bb6aa70
|
|
| MD5 |
7f50fff878f04be8a1ecd5f620a63642
|
|
| BLAKE2b-256 |
632cd1618368b5900378d1ef142dbc35d72575f42e9b83dc2ab7fbcd09cce47d
|











