PySAMOSA is a software framework for processing open ocean and coastal waveforms from SAR satellite altimetry to measure sea surface heights, wave heights, and wind speed for the oceans and inland water bodies. Satellite altimetry is a space-borne remote sensing technique used for Earth observation.
Project description
PySAMOSA

PySAMOSA is a Python-based software for processing open ocean and coastal waveforms from SAR satellite altimetry to measure sea surface heights, wave heights, and wind speed for the oceans and inland waters. Satellite altimetry is a space-borne remote sensing technique used for Earth observation. More details on satellite altimetry can be found here.
The process of extracting of the three geophysical parameters from the reflected echoes/waveforms is called retracking. The measured (noisy) waveforms are fitted against the open ocean power return echo waveform model SAMOSA2 [1,2].
In the coastal zone, the return echoes are affected by spurious signals from strongly reflective targets such as sand and mud banks, tidal flats, shipping platforms, sheltered bays, or calm waters close to the shoreline.
The following European Space Agency (ESA) satellite altimetry missions are supported:
- Sentinel-3 (S3)
- Sentinel-6 Michael Freilich (S6-MF)
The software retracks the waveforms, i.e. the Level-1b (L1b) data, to extract the retracked variables SWH, range, and Pu.
The open ocean retracker implementation specification documents from the official EUMETSAT baseline are used (S3 [1], S6-MF [2]).
For retracking coastal waveforms the following retrackers are implemented:
- SAMOSA+ [3]
- CORAL [4,5]
In addition, FF-SAR-processed S6-MF data can be retracked using the zero-Doppler beam of the SAMOSA2 model and a specially adapted $\alpha_p$ LUT table, created by the ESA L2 GPP project [7]. The application of the FF-SAR-processed data has been validated in [5].
Not validated (experimental) features:
- CryoSat-2 (CS2) support
- SAMOSA++ coastal retracker [2]
- Monte-carlo SAMOSA2 simulator
Getting-started
Usage
Install pysamosa into your environment
$ pip install pysamosa
This is the sample to retrack a single L1b file from the S6-MF mission
from pathlib import Path
import numpy as np
from pysamosa.common_types import L1bSourceType
from pysamosa.data_access import data_vars_s3, data_vars_s6
from pysamosa.retracker_processor import RetrackerProcessor
from pysamosa.settings_manager import get_default_base_settings, SettingsPreset
l1b_files = []
# l1b_files.append(Path('S6A_P4_1B_HR______20211120T051224_20211120T060836_20220430T212619_3372_038_018_009_EUM__REP_NT_F06.nc'))
l1b_files.append(Path.cwd().parent / '.data' / 's6' / 'l1b' / 'S6A_P4_1B_HR______20211120T051224_20211120T060836_20220430T212619_3372_038_018_009_EUM__REP_NT_F06.nc')
l1b_src_type = L1bSourceType.EUM_S6_F06
data_vars = data_vars_s6
# configure coastal CORAL retracker
pres = SettingsPreset.CORALv2
rp_sets, retrack_sets, fitting_sets, wf_sets, sensor_sets = get_default_base_settings(settings_preset=pres, l1b_src_type=l1b_src_type)
rp_sets.nc_dest_dir = l1b_files[0].parent / 'processed'
rp_sets.n_offset = 0
rp_sets.n_inds = 0 #0 means all
rp_sets.n_procs = 6 #use 6 cores in parallel
rp_sets.skip_if_exists = False
additional_nc_attrs = {
'L1B source type': l1b_src_type.value.upper(),
'Retracker preset': pres.value.upper(),
}
rp = RetrackerProcessor(l1b_source=l1b_files, l1b_data_vars=data_vars['l1b'],
rp_sets=rp_sets,
retrack_sets=retrack_sets,
fitting_sets=fitting_sets,
wf_sets=wf_sets,
sensor_sets=sensor_sets,
nc_attrs_kw=additional_nc_attrs,
bbox=[np.array([-29.05, -29.00, 0, 360])],
)
rp.process() #start processing
print(rp.output_l2) #retracked L2 output can be found in here
A running minimal working example for retracking is shown in notebooks/retracking_example.ipynb.
Development
It is highly recommended to use a proper Python IDE, such as PyCharm Community or Visual Studio Code. Using the IDE will allow you to familiarise yourself better with the code, debug and extend it.
Clone the repo
$ git clone {repo_url}
Enter cloned directory
$ cd pysamosa
Install dependencies into your conda env/virtualenv
$ pip install -r requirements.txt
Compile the .pyx files (e.g. model_helpers.pyx) by running cython to build the extensions For Windows users: An installed C/C++ compiler may be required for installation, e.g. MSCV, which comes with the free Visual Studio Community
$ python setup.py build_ext --inplace
Optional: Compile on an HPC cluster (not normally required)
$ LDSHARED="icc -shared" CC=icc python setup.py build_ext --inplace
Tips
The following list provides a brief description of the recommended use of the software.
-
Getting-started with Jupyter Notebook The
notebooks/retracking_example.ipynbcontains a sample script how to retrack a sample EUMETSAT baseline L1b file. The retracked SWH and SWH data are compared with the EUMETSAT baseline L2 data. Thenotebooks/demo_script.pyprovides the code example from above to quickly launch a small retracking example. -
More entry points The files
main_s3.py,main_s6.py,main_cs.py, (main_*.py) etc. serve as entry points for batch processing of multiple nc files. A list of L1b files (or a single file) is read for retracking, which are fully retracked or based on the given bounding box (bbox) paramater. A retracked L2 file is written out per processed L1b file. -
Settings The
RetrackerProcessorinputs require theRetrackerProcessorSettings,RetrackerSettings,FittingSettings,WaveformSettings, andSensorSettingsobjects to be inserted during initialisation. The default settings of these settings objects can be retrieved with theget_default_base_settingsfunction based on the three settingsL1bSourceTypeandSettingsPreset. For instance, the following code snippet is taken from themain_s3.pyfile and retracks Sentinel-3 data with the default SAMOSA-based open ocean retracker with no SettingsPreset (100 waveforms from measurement index 25800, and using 6 cores).
l1b_src_type = L1bSourceType.EUM_S3
pres = SettingsPreset.NONE #use this for the standard SAMOSA-based retracker [2]
# pres = SettingsPreset.CORALv2 #use this for CORALv2 [5]
# pres = SettingsPreset.NONE #use this for SAMOSA+ [3]
rp_sets, retrack_sets, fitting_sets, wf_sets, sensor_sets = get_default_base_settings(settings_preset=pres, l1b_src_type=l1b_src_type)
rp_sets.nc_dest_dir = nc_dest_path / run_name
rp_sets.n_offset = 25800
rp_sets.n_inds = 100
rp_sets.n_procs = 6
rp_sets.skip_if_exists = False
- Evaluation environment
There are several unit tests located in
./pysamosa/tests/that aim to analyse the retracked output in more detail. The most important test scripts aretest_retrack_multi.py, which includes retracking of small along-track segments of the S3A, S6, CS2 missions (and a generic input nc file).test_retrack_singleallows you to check the retracking result of a single waveform and compare it to reference retracking result.
Please uncomment the line mpl.use('TkAgg') in file conftest.py to
plot the test output, which is particularly useful for the retracking tests in files tests/test_retrack_multi. py and tests/test_retrack_multi.py.
- Difference between CORALv1 and CORALv2
- v2 has two additional extensions that were required for S6-MF
- retrack_sets.interference_masking_mask_before_le = True Interference signals before the leading edge are also masked out by the adaptive inteference mitigation scheme (AIM, CORAL feature)
- fitting_sets.Fit_Var_2_MinMax_Hs = (0.0, 20) lower SWH boundary for fitting procedure is set to 0.0, as defined in [2]
- Quality flag
During the retracking process, the quality flag variables
swh_qual' andrange_qual' (where the latter is just a copy of the former) are part of the retracked output and indicate the quality of the retracking of each individual waveform (0=good, 1=bad). This makes a difference particularly in coastal scenarios where the waveforms are affected by spurious signals which tend to cause incorrectly retracked waveforms. The CORAL coastal retracker maximises the number of valid records in the coastal zone. We therefore emphasise the importance of consideringswh_qual/range_qualquality flags in the retracked product.
Validation
Run tests
To run all the unit tests (using the pytest framework), run
$ pytest
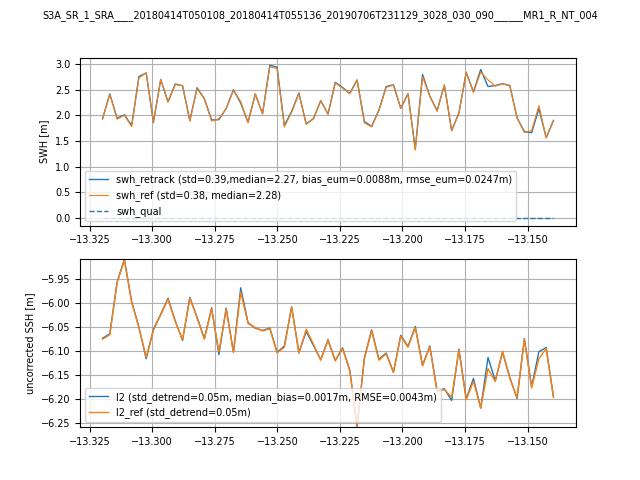
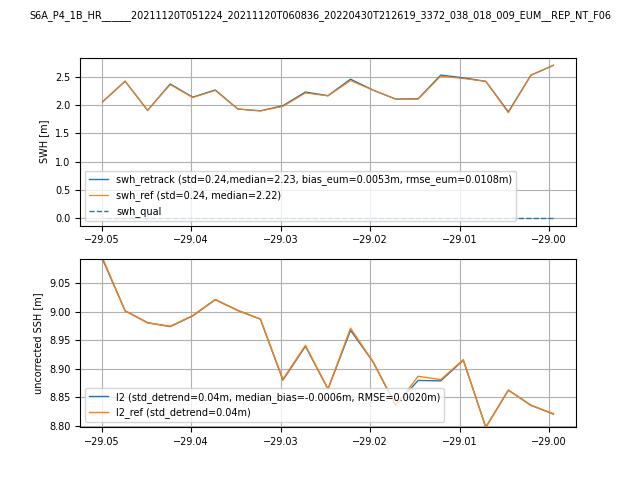
Comparison with EUMETSAT L2 baseline data
Comparison of a retracked open ocean segment from S3 and S6-MF missions with the EUMETSAT L2 baseline (S3: 004,
S6-MF: F06)
(generated by notebooks/retracking_example.ipynb Jupyter notebook)
| S3 | S6-MF |
|---|---|
 |
 |
Contributions
This software is intended to be a community-based project. Contributions to this project are very welcome. In this case:
- Fork this repository
- Submit a pull request to be merged back into this repository.
Before submitting changes, please check that your changes pass flake8, black, isort and the tests, including testing other Python versions with tox:
$ flake8 pysamosa tests scripts
$ black . --check --diff
$ isort . --check-only --diff
$ pytest
$ tox
If your pull request is accepted, you will be included in the next official release and will be listed as a co-author for the DOI link created by Zenodo.
Future work
Possible developments of this project are:
Retracking-related
- Align CS-2 retracking with the CS-2 baseline processing chain, validate against SAMpy developed as part of the ESA Cryo-TEMPO project
- Implement evolutions of the EUMETSAT's baseline processing chain [6], e.g. the numerical retracking planned for Q3/2023
Software-related
- Create notebook for a coastal retracking demo
- Create richer documentation (readthedocs)
Citation
If you use this software or the code, please cite this DOI:
Florian Schlembach; Marcello Passaro. PySAMOSA: An Open-source Software Framework for Retracking SAMOSA-based, Open Ocean and Coastal Waveforms of SAR Satellite Altimetry. Zenodo. https://zenodo.org/badge/latestdoi/646028227.
Acknowledgement
The authors are grateful to
Salvatore Dinardo for his support in implementing the SAMOSA-based and SAMOSA+ [3] retracking algorithms.
This package was created with Cookiecutter and the audreyr/cookiecutter-pypackage project template.
References
[1] SAMOSA Detailed Processing Model: Christine Gommenginger, Cristina Martin-Puig, Meric Srokosz, Marco Caparrini, Salvatore Dinardo, Bruno Lucas, Marco Restano, Américo, Ambrózio and Jérôme Benveniste, Detailed Processing Model of the Sentinel-3 SRAL SAR altimeter ocean waveform retracker, Version 2.5.2, 31 October 2017, Under ESA-ESRIN Contract No. 20698/07/I-LG (SAMOSA), Restricted access as defined in the Contract, Jérôme Benveniste (Jerome.Benvensite@esa.int) pers. comm.
[2] EUMETSAT. Sentinel-6/Jason-CS ALT Level 2 Product Generation Specification (L2 ALT PGS), Version V4D; 2022. https://www.eumetsat.int/media/48266.
[3] Dinardo, Salvatore. ‘Techniques and Applications for Satellite SAR Altimetry over Water, Land and Ice’. Dissertation, Technische Universität, 2020. https://tuprints.ulb.tu-darmstadt.de/11343/.
[4] Schlembach, F.; Passaro, M.; Dettmering, D.; Bidlot, J.; Seitz, F. Interference-Sensitive Coastal SAR Altimetry Retracking Strategy for Measuring Significant Wave Height. Remote Sensing of Environment 2022, 274, 112968. https://doi.org/10.1016/j.rse.2022.112968.
[5] Schlembach, F.; Ehlers, F.; Kleinherenbrink, M.; Passaro, M.; Dettmering, D.; Seitz, F.; Slobbe, C. Benefits of Fully Focused SAR Altimetry to Coastal Wave Height Estimates: A Case Study in the North Sea. Remote Sensing of Environment 2023, 289, 113517. https://doi.org/10.1016/j.rse.2023.113517.
[6] Scharroo, R.; Martin-Puig, C.; Meloni, M.; Nogueira Loddo, C.; Grant, M.; Lucas, B. Sentinel-6 Products Status. Ocean Surface Topography Science Team (OSTST) meeting in Venice 2022. https://doi.org/10.24400/527896/a03-2022.3671.
[7] ESA L2 GPP Project: FF-SAR SAMOSA LUT generation was funded under ESA contract 4000118128/16/NL/AI.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pysamosa-1.0.0.tar.gz.
File metadata
- Download URL: pysamosa-1.0.0.tar.gz
- Upload date:
- Size: 8.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5ebdb48ea9ea47d8f329122ad2cae6bea6868b220260257f17ad254eca08c617
|
|
| MD5 |
11082554e7284ec55c482c34284eec64
|
|
| BLAKE2b-256 |
0e8ec1caca44fee3272cec7b88cef924b1c9f5865d23b14ab27f636e32ae6788
|
File details
Details for the file pysamosa-1.0.0-pp310-pypy310_pp73-win_amd64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp310-pypy310_pp73-win_amd64.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f9997432ce3406d377121e0128b58103e50efb55539a130ae28fe97a02c8f99a
|
|
| MD5 |
4080002bee3421b2efebc00486b84fc7
|
|
| BLAKE2b-256 |
f5bccc5e5481e753d0ecffd33d7a810999728949e2e82cc92d15c5ca49fa5e42
|
File details
Details for the file pysamosa-1.0.0-pp310-pypy310_pp73-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp310-pypy310_pp73-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
95ae5d2917060d22484d2e6069467f0583f92fe4a86a6ae59ecca7eed2224170
|
|
| MD5 |
6ae67d91663456ed3cdd479979405124
|
|
| BLAKE2b-256 |
9b7e57f377982ad4fdd8414d31e1d112551f9e90751199d3a35a65acea2dd551
|
File details
Details for the file pysamosa-1.0.0-pp310-pypy310_pp73-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp310-pypy310_pp73-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, manylinux: glibc 2.17+ i686, manylinux: glibc 2.5+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cb02ed84420614ff71189499d19ccb5bdac7bd191c7ea3d7588d05ac18ebc684
|
|
| MD5 |
5fa68572ecdda6e0ec9045c2381b98be
|
|
| BLAKE2b-256 |
4656ee964b093652bf5c9f17f8c0290637e080b37806875243a09a665715a546
|
File details
Details for the file pysamosa-1.0.0-pp310-pypy310_pp73-macosx_10_9_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp310-pypy310_pp73-macosx_10_9_x86_64.whl
- Upload date:
- Size: 3.9 MB
- Tags: PyPy, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9a77370cf27cb62ead2d530fb07b1fcb1464b26745b05a9adc482c3fd32faffa
|
|
| MD5 |
7a48f66760dd9673f3ac9ec3aa5b5081
|
|
| BLAKE2b-256 |
3a5e0ad4a5b9b7ab70a79892082b52b21f22dbd3497a4b246e859c78fa093923
|
File details
Details for the file pysamosa-1.0.0-pp39-pypy39_pp73-win_amd64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp39-pypy39_pp73-win_amd64.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c6ac1ddd9b61636a806977ac80bb8dbdd1e0962e634448e51417e90f21f17cdf
|
|
| MD5 |
a013634f3ae338efd05e8205e0afc588
|
|
| BLAKE2b-256 |
c5401ab3efd906dfb090ce517f11eff50e6d168efe9f26735ff9383b52241dc3
|
File details
Details for the file pysamosa-1.0.0-pp39-pypy39_pp73-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp39-pypy39_pp73-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cef436779b650840e905abd2b82f1709b05004ffa19a662bc2762a28e34d8198
|
|
| MD5 |
c423ce191c87b0453f7cfa72fc555da0
|
|
| BLAKE2b-256 |
54cbca2a66b6378e6b989f2d7fd0be2ba18e9a9d129883217492408b71091df4
|
File details
Details for the file pysamosa-1.0.0-pp39-pypy39_pp73-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp39-pypy39_pp73-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, manylinux: glibc 2.17+ i686, manylinux: glibc 2.5+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5213eb10fdc2e0523a01b7b080731a6543e47e380a4fdb7b86fe5ded3565cfa1
|
|
| MD5 |
eb944288e16bb35130682d893983267f
|
|
| BLAKE2b-256 |
1b9c863d9430c60e4ca657dd621f42f269a5ca20fad4f4a89bd7834464f33477
|
File details
Details for the file pysamosa-1.0.0-pp39-pypy39_pp73-macosx_10_9_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp39-pypy39_pp73-macosx_10_9_x86_64.whl
- Upload date:
- Size: 3.9 MB
- Tags: PyPy, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cd80455c0ff9a557fc9239bec91c9b368c1d32ec2e96019e621001238e8e65b3
|
|
| MD5 |
12fa463c4db7119add1b6b1406830ad2
|
|
| BLAKE2b-256 |
72d43493304d033a0767e416851153b292acdc15646a712a34ca71b7c2f7fa8e
|
File details
Details for the file pysamosa-1.0.0-pp38-pypy38_pp73-win_amd64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp38-pypy38_pp73-win_amd64.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e4ece37e4f545f14f2921a69f80b16ab091884143308d14f898efb048b0ccaf3
|
|
| MD5 |
d9934c33dbc9c5c13faca6ecbb772019
|
|
| BLAKE2b-256 |
d7e59285d41907f7ec5b7d6fa3f04dd85ea0818f1ebe820065be0e8e2859b88c
|
File details
Details for the file pysamosa-1.0.0-pp38-pypy38_pp73-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp38-pypy38_pp73-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
78b3d2af73cb3d0f372873f7a5543123e5c2df7578227c472c7b5a86d90de901
|
|
| MD5 |
751cc31f673116f02141464f77b6b215
|
|
| BLAKE2b-256 |
12b3edb4043894b523ddb7da0239fd5edd18a8aad182d8df61deab838610aaa9
|
File details
Details for the file pysamosa-1.0.0-pp38-pypy38_pp73-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp38-pypy38_pp73-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl
- Upload date:
- Size: 4.0 MB
- Tags: PyPy, manylinux: glibc 2.17+ i686, manylinux: glibc 2.5+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
82bbbbecf8dddc05c768a4b8cbb645f2d2f7644a83ee51f830c6a18f4a309a31
|
|
| MD5 |
5f7a97b8e7f88a96b3a29cb56a7eb6b6
|
|
| BLAKE2b-256 |
67765e394e9c4561586adf2e76574193f9cc59ccd5d05184b2f8dd904f949342
|
File details
Details for the file pysamosa-1.0.0-pp38-pypy38_pp73-macosx_10_9_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-pp38-pypy38_pp73-macosx_10_9_x86_64.whl
- Upload date:
- Size: 3.9 MB
- Tags: PyPy, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
51b1b87b183482dc4d3d41774e20a182d03eb72fcfdb31ff20932781b564a7e7
|
|
| MD5 |
b281fbe6e63e7791be836448a406811f
|
|
| BLAKE2b-256 |
6c640074168a7f2515fb0bf07d4990284fb73f1b9200c9d988bceafb9c4c5d74
|
File details
Details for the file pysamosa-1.0.0-cp311-cp311-win_amd64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp311-cp311-win_amd64.whl
- Upload date:
- Size: 4.0 MB
- Tags: CPython 3.11, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e0b33ba362c228b3c5bd7447e3b2feedd09fad468f6965e99e8480f09f682bb3
|
|
| MD5 |
f500d9055642948a37b4ba3b554757f2
|
|
| BLAKE2b-256 |
2ed41a860b6352a3c750914645dc391cf1b12492dfc2ad59ead526435341de74
|
File details
Details for the file pysamosa-1.0.0-cp311-cp311-win32.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp311-cp311-win32.whl
- Upload date:
- Size: 3.9 MB
- Tags: CPython 3.11, Windows x86
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0ac777e1aa92ad0de0479b1a773dabc876bf479f0835331fbb2636058752ed0e
|
|
| MD5 |
f88b8cb3e2ee311805ee7e5e3a9eab88
|
|
| BLAKE2b-256 |
ca597045bb0fadbeae391d4f4f6756e9bc753ba1a74deadae685c560a99a917b
|
File details
Details for the file pysamosa-1.0.0-cp311-cp311-musllinux_1_1_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp311-cp311-musllinux_1_1_x86_64.whl
- Upload date:
- Size: 4.4 MB
- Tags: CPython 3.11, musllinux: musl 1.1+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
28baebf7a4a0ea213b2db7738af7391fd3a6b2f8b8eed9fb156bc84baa8f4ee3
|
|
| MD5 |
6074feace6cb2bd2992cba9a221d3af2
|
|
| BLAKE2b-256 |
6613dcadb3e698bffb4f6007c65bccaf35306208be8d598728b728db240ecc08
|
File details
Details for the file pysamosa-1.0.0-cp311-cp311-musllinux_1_1_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp311-cp311-musllinux_1_1_i686.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.11, musllinux: musl 1.1+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
65e76da0d46d64fb1e6811758dbaa20707f75dddadfae3eff2d70e7c28d60117
|
|
| MD5 |
7120fef785e67d40b5aeda7698566e85
|
|
| BLAKE2b-256 |
535ef1005bfd8f0a0139f870298171a18a9dd044138ec794a074c44a178f08c9
|
File details
Details for the file pysamosa-1.0.0-cp311-cp311-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp311-cp311-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 4.4 MB
- Tags: CPython 3.11, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
913821ef6eff31dfc32915076a51ed15e7d38533344258ffe30b69771eba74b8
|
|
| MD5 |
4a0bbe213c81b2196bba61315ea559a3
|
|
| BLAKE2b-256 |
460a1ebcaacb775863b9135d3ad550612ba521d25b33e9d4ddf9b9b0cf688152
|
File details
Details for the file pysamosa-1.0.0-cp311-cp311-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp311-cp311-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.11, manylinux: glibc 2.17+ i686, manylinux: glibc 2.5+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3900ed95e94c36ae1f397d956e863be28bb2e271df6d7470287cf0d6929473a9
|
|
| MD5 |
e47d384ca98639829e4f5f19ea22537a
|
|
| BLAKE2b-256 |
376e2eff6ea49442ebdc87f7fd93db1aefdd043326d88cf5887095de9557f90c
|
File details
Details for the file pysamosa-1.0.0-cp311-cp311-macosx_10_9_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp311-cp311-macosx_10_9_x86_64.whl
- Upload date:
- Size: 4.0 MB
- Tags: CPython 3.11, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
14987a3ff08e07fad189d3c09e8e9a2a73fe080d1a8382121af02127ca1ced33
|
|
| MD5 |
d66a61575174eb9234d18896adeed60a
|
|
| BLAKE2b-256 |
397b58bd557670c6c2f72baab18287681d5ffb63f096d2c3374562b6d4dcb723
|
File details
Details for the file pysamosa-1.0.0-cp310-cp310-win_amd64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp310-cp310-win_amd64.whl
- Upload date:
- Size: 4.0 MB
- Tags: CPython 3.10, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
afa34063ae42fcda7c500c3441cdebfb2321a7adf9dc4d2c9c9753ffa96340ce
|
|
| MD5 |
cdd92c461d5d6365a2c464969a1ef5b7
|
|
| BLAKE2b-256 |
00df54bb2e1c21d6d775164392affad86b38638ff4c2c71fb482fd288a43c3ba
|
File details
Details for the file pysamosa-1.0.0-cp310-cp310-win32.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp310-cp310-win32.whl
- Upload date:
- Size: 3.9 MB
- Tags: CPython 3.10, Windows x86
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fdb2533f7950c1f9efaf813e06606b987e98ed0ec9f094d817c4fbd76acc9ebc
|
|
| MD5 |
a4f78ffc3e755228782ca9a358a908ae
|
|
| BLAKE2b-256 |
bf61c7bc7ce59bf3360389c19c7d1a87714526091c90dfb935fa261f947d7915
|
File details
Details for the file pysamosa-1.0.0-cp310-cp310-musllinux_1_1_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp310-cp310-musllinux_1_1_x86_64.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.10, musllinux: musl 1.1+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1a54a8881003a9b05d03b1d30495f8c8ec5c1c69e3a42b80645f1484bc97c772
|
|
| MD5 |
7b2fefdc8e86afab7ce25951c962b543
|
|
| BLAKE2b-256 |
1b3e2e45fc25df8dbccdb235c7de865021375ea35874a90e8edc380402d5b480
|
File details
Details for the file pysamosa-1.0.0-cp310-cp310-musllinux_1_1_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp310-cp310-musllinux_1_1_i686.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.10, musllinux: musl 1.1+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7c67d27cb42d9ce27dfd9d73e9c853ab3037ee10e19767582a80ffe9b7bfb657
|
|
| MD5 |
353bd0862f2189b78389c9f571ba8fbf
|
|
| BLAKE2b-256 |
c0e6980eb7c3171a4e2b331894bc82bbc8cce7abdc9ea5b9aa1212c5e17552f6
|
File details
Details for the file pysamosa-1.0.0-cp310-cp310-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp310-cp310-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.10, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b58525da785ef7dea3b23c342370e55081754e0c86645f7a0502f1e6d4da0ab9
|
|
| MD5 |
fef98306f7060bea91e829804a587396
|
|
| BLAKE2b-256 |
7626e4e285ea7523a5af88e12bf85a4ad73c71c1cdb363709d2c5804cc32a811
|
File details
Details for the file pysamosa-1.0.0-cp310-cp310-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp310-cp310-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.10, manylinux: glibc 2.17+ i686, manylinux: glibc 2.5+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c853453b5aa053b5fe95dd876b8d8fe1a1174b06630fc3131c4861623c6d553c
|
|
| MD5 |
292190e5ee715cc2c1acd00b2d280db2
|
|
| BLAKE2b-256 |
b57233fa4fe3a0ab8dce02363fb17b4333f2d213e6a922422e2ff3fcef6f9492
|
File details
Details for the file pysamosa-1.0.0-cp310-cp310-macosx_10_9_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp310-cp310-macosx_10_9_x86_64.whl
- Upload date:
- Size: 4.0 MB
- Tags: CPython 3.10, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b117f598b1fe52999c00284e7f19485f44eb895f9064f8fcd7f1c29a13756baf
|
|
| MD5 |
3232c61ba7bcda8a2be59e86d31d54e8
|
|
| BLAKE2b-256 |
3ab052244b4ccbdf088b322e90c22eb4ab1b9cfae83875c0c4bbcc3019b34167
|
File details
Details for the file pysamosa-1.0.0-cp39-cp39-win_amd64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp39-cp39-win_amd64.whl
- Upload date:
- Size: 4.0 MB
- Tags: CPython 3.9, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d5201c5606ab964a5599c78ab80977ddda8ebc14060485b45b0cffe0a5fc73f9
|
|
| MD5 |
c4cb405976660b0a2f42595923e6bad1
|
|
| BLAKE2b-256 |
12742aa6abce695908211d75e8d40b8a6521125fb2cb9acfc77b62234b9cfd59
|
File details
Details for the file pysamosa-1.0.0-cp39-cp39-win32.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp39-cp39-win32.whl
- Upload date:
- Size: 3.9 MB
- Tags: CPython 3.9, Windows x86
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cad0b25d02cd8dd6ce71e4299cbc44ba7464a4d6b4cf65ded77bcb2964935590
|
|
| MD5 |
fcdc39cc4d55356f105bcbc41d039a00
|
|
| BLAKE2b-256 |
71090e6667916319f1e35e0fcdc07b7494c0ce85a38bb6e4aa6670c5a8850f98
|
File details
Details for the file pysamosa-1.0.0-cp39-cp39-musllinux_1_1_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp39-cp39-musllinux_1_1_x86_64.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.9, musllinux: musl 1.1+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8624b5146731e1c2cecbe72ed1ce085bd744fe635b1abdc13bcaece4524322f6
|
|
| MD5 |
34b78338c9adc642a1f6c10c253e7a92
|
|
| BLAKE2b-256 |
a6d1ee6efb352912462fbd121efd833c30b2bb9a9eafb1f510acbf70cf0fdaf1
|
File details
Details for the file pysamosa-1.0.0-cp39-cp39-musllinux_1_1_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp39-cp39-musllinux_1_1_i686.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.9, musllinux: musl 1.1+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
99f48c5f9e432c1e2b3b32dc1c61f8ba8730c5c8577d97f393227dc3ca895e95
|
|
| MD5 |
a4263541fb9014632fe8f93515bc8211
|
|
| BLAKE2b-256 |
4342ab1b997cc20ed02948034210f63860186df8db8e35498445ab393ae6f327
|
File details
Details for the file pysamosa-1.0.0-cp39-cp39-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp39-cp39-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.9, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5464e16090a266fbcfc047982babe40089436197343ac902606db03697b4045c
|
|
| MD5 |
b0a7cd527f03f8ad520697fb326c6e71
|
|
| BLAKE2b-256 |
b2fa4f53817637b8c8d15ed3088e7633d78fc8ff24fda25063be372ce37e3641
|
File details
Details for the file pysamosa-1.0.0-cp39-cp39-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp39-cp39-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.9, manylinux: glibc 2.17+ i686, manylinux: glibc 2.5+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
caea9210329426d212d87e8f3bcb482c6877eb5cf73a26e19602e871696292cf
|
|
| MD5 |
92a02ed2b4fa28d0856409b4aa8cac9a
|
|
| BLAKE2b-256 |
09896f0327dfa5e3b84a1a88efc75a6ed9761a61ad1d48afc14a8ee06c650b05
|
File details
Details for the file pysamosa-1.0.0-cp39-cp39-macosx_10_9_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp39-cp39-macosx_10_9_x86_64.whl
- Upload date:
- Size: 4.0 MB
- Tags: CPython 3.9, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
207664d94c53789dedaa1e6cb3028a89aa8b8247f1581cabb1575acb7157eea3
|
|
| MD5 |
90f073f908c4075b8e7c2e84ef53ee78
|
|
| BLAKE2b-256 |
746b32156471f4566789c6340f1a394608082b0874102eb9cdbf57726df9cd96
|
File details
Details for the file pysamosa-1.0.0-cp38-cp38-win_amd64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp38-cp38-win_amd64.whl
- Upload date:
- Size: 4.0 MB
- Tags: CPython 3.8, Windows x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5d7452191f24e4b121d4082b61c803ce004cf944402ac40528b114471ca52202
|
|
| MD5 |
d6f034a1a1e0117cf3b55d50dff55a5d
|
|
| BLAKE2b-256 |
1160b7d4a9819d84906746009b42ecbee2b5c23a8eabe47ddf9aa89f4afefbb4
|
File details
Details for the file pysamosa-1.0.0-cp38-cp38-win32.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp38-cp38-win32.whl
- Upload date:
- Size: 3.9 MB
- Tags: CPython 3.8, Windows x86
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
88adfe1fdf674b587b770a2c6e2128a97eef9cc4be3aa4fe63a99671a2aba3c2
|
|
| MD5 |
0e77936e785a2ccfb994d4c05783ce19
|
|
| BLAKE2b-256 |
ca77981a451114a08455b275fbb9e11dafbdfd9b669ddde93f2a0d5dafbb1e81
|
File details
Details for the file pysamosa-1.0.0-cp38-cp38-musllinux_1_1_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp38-cp38-musllinux_1_1_x86_64.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.8, musllinux: musl 1.1+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e57bb370a0699e71fdc8b3535693a8b7c1194ce2d2f46a821062efa327ccc9f8
|
|
| MD5 |
e5d10aba66a1cf89f717a856d393d6ed
|
|
| BLAKE2b-256 |
a35442174d8b54845431ec5ee20077793c5748a15500a4343926ff682c202aed
|
File details
Details for the file pysamosa-1.0.0-cp38-cp38-musllinux_1_1_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp38-cp38-musllinux_1_1_i686.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.8, musllinux: musl 1.1+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cd6688937ab6498244fa7eb4488083b9c4e10e8cc7a54972884bf516feb13d73
|
|
| MD5 |
407626d054bda4bc1fa27c4106919920
|
|
| BLAKE2b-256 |
d49923b5247780cc0d325d7e1532ee3db011202f12786683cbeca9ed072f3344
|
File details
Details for the file pysamosa-1.0.0-cp38-cp38-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp38-cp38-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.8, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c4e78edceb9a94be72befc8cdcc6032a2b67d8374acfff04410ab9d14d3563b5
|
|
| MD5 |
4cd0269c05ecfc7815de88a1bb690602
|
|
| BLAKE2b-256 |
dd53ece380567768e1d2439f75dc9b15cf0a18b980ba6222390866f63f5cf464
|
File details
Details for the file pysamosa-1.0.0-cp38-cp38-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp38-cp38-manylinux_2_5_i686.manylinux1_i686.manylinux_2_17_i686.manylinux2014_i686.whl
- Upload date:
- Size: 4.3 MB
- Tags: CPython 3.8, manylinux: glibc 2.17+ i686, manylinux: glibc 2.5+ i686
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0a7799c3ed02e4f31c377489251260c6a8841c5115030824064e1f3455afeae6
|
|
| MD5 |
1c20c2e89cce98e37d26379361ed7aa3
|
|
| BLAKE2b-256 |
055d9f3c6a62e2cb5e103a50ec6c621cd215a60376aed7a7ad030a9459c29d0e
|
File details
Details for the file pysamosa-1.0.0-cp38-cp38-macosx_10_9_x86_64.whl.
File metadata
- Download URL: pysamosa-1.0.0-cp38-cp38-macosx_10_9_x86_64.whl
- Upload date:
- Size: 4.0 MB
- Tags: CPython 3.8, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/4.0.2 CPython/3.11.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7827fd160b46679cbfadc105c0b977d5d01d7b88dbabaab438f73b473deee31e
|
|
| MD5 |
9794a66b7bb3b548b3737c326c2e343d
|
|
| BLAKE2b-256 |
b3453c5e18ac6125d862bae5f59f4905cf3751cbfac84cb74531ad8498b09d24
|
















