Unsupervised clustering algorithm for 2D data (neurons by time)
Project description
Rastermap
Rastermap is a discovery algorithm for neural data. The algorithm was written by Carsen Stringer and Marius Pachitariu. For support, please open an issue. Please see install instructions below. Check out the paper and the tutorial video for more info. Rastermap runs in python 3.8+ and has a graphical user interface (GUI) for running it easily.
If you use Rastermap or analysis code in this repo in your work, please cite the paper: Stringer C., Zhong L., Syeda A., Du F., Kesa M., & Pachitariu M. (2024). Rastermap: a discovery method for neural population recordings. Nature Neuroscience. https://doi.org/10.1038/s41593-024-01783-4.
Table of Contents
Example notebooks
- rastermap_largescale.ipynb
shows how to use it with large-scale data from mouse cortex (> 200 neurons)
- rastermap_singleneurons.ipynb
shows how to use it with small to medium sized data (< 200 neurons), in this case recorded from rat hippocampus
- rastermap_zebrafish.ipynb
shows how to use it with large-scale data from zebrafish
- rastermap_widefield.ipynb
shows how to use it with widefield imaging data, or other types of datasets that are too large to fit into memory
- rastermap_interactive.ipynb
allows running Rastermap in an interactive way without a local installation
- tutorial.ipynb
is a guided tutorial for integrating rastermap and facemap to visualize behavioral representations. See the student/teacher versions here.
Also all notebooks to analyze the data and create the figures in the paper are here. All data available here.
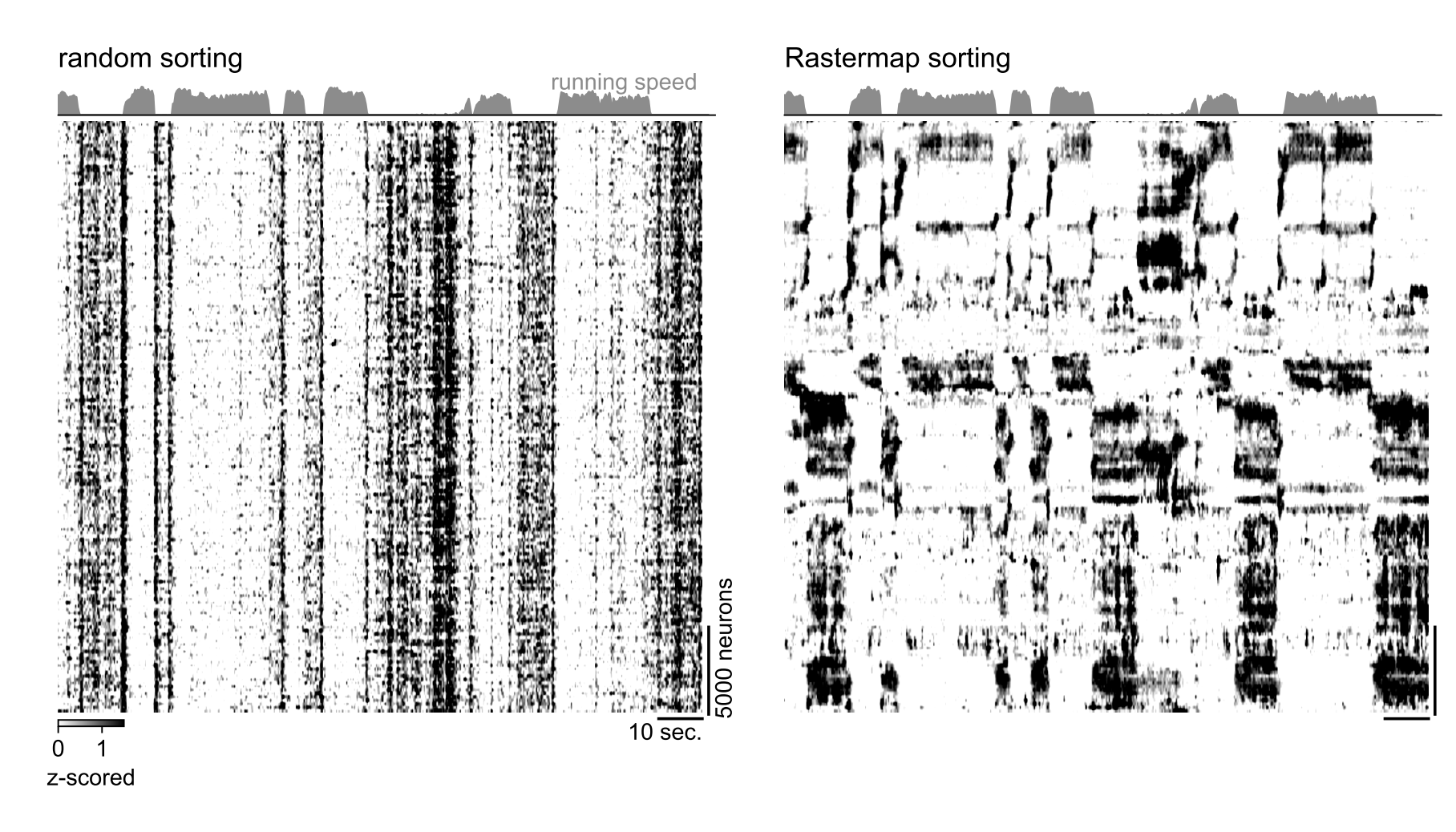
Here is what the output looks like for a segment of a mesoscope recording in a mouse during spontaneous activity (3.2Hz sampling rate), compared to random neural sorting:
Installation
Local installation (< 2 minutes)
System requirements
Linux, Windows and Mac OS are supported for running the code. For running the graphical interface in Mac, you will need a Mac OS later than Yosemite. At least 8GB of RAM is recommended to run the software. 16GB-32GB may be required for larger datasets. The software has been heavily tested on Windows 10 and Ubuntu 20.04 and less well-tested on Mac OS. Please open an issue if you have problems with installation.
Instructions
We recommend to install a miniforge (conda-based) distribution of Python. Note you might need to use an anaconda prompt (windows) if you did not add anaconda to the path. Open an anaconda prompt / command prompt with python 3 in the path, then:
pip install rastermap
For the GUI
pip install rastermap[gui]
Rastermap has only a few dependencies so you may not need to make a special environment for it
(e.g. it should work in a suite2p or facemap environment), but if the pip install above does not work,
please follow these instructions:
- Open an anaconda prompt / command prompt which has
condafor python 3 in the path - Create a new environment with
conda create --name rastermap python=3.9. We recommend python 3.9, but python 3.8, 3.9 and 3.11 will also work. - To activate this new environment, run
conda activate rastermap - (option 1) To install rastermap with the GUI, run
python -m pip install rastermap[gui]. If you're on a zsh server, you may need to use ' ':python -m pip install 'rastermap[gui]'. - (option 2) To install rastermap without the GUI, run
python -m pip install rastermap. To upgrade rastermap (package here), run the following in the environment:
pip install rastermap --upgrade
If you have an older rastermap environment you can remove it with conda env remove -n rastermap before creating a new one.
Note you will always have to run conda activate rastermap before you run rastermap. If you want to run jupyter notebooks in this environment, then also pip install notebook.
Dependencies
This package relies on the awesomeness of numpy, scipy, numba, scikit-learn, PyQt6, PyQt6.sip and pyqtgraph. You can pip install or conda install all of these packages. If having issues with PyQt6, then make an Anaconda environment and try to install within it conda install pyqt. On Ubuntu you may need to sudo apt-get install libegl1 to support PyQt6. Alternatively, you can use PyQt5 by running pip uninstall PyQt6 and pip install PyQt5. If you already have a PyQt version installed, Rastermap will not install a new one.
Using rastermap
GUI
The quickest way to start is to open the GUI from a command line terminal. You might need to open an anaconda prompt if you did not add anaconda to the path. Then run:
python -m rastermap
To start using the GUI, save your data into an npy file that is just a matrix that is neurons x timepoints. Then "File > Load data matrix" and choose this file (or drag and drop your file). You can also try it out with our demo data available here.
Next click "Run > Run rastermap" and click run. See the parameters section to learn about the parameters.
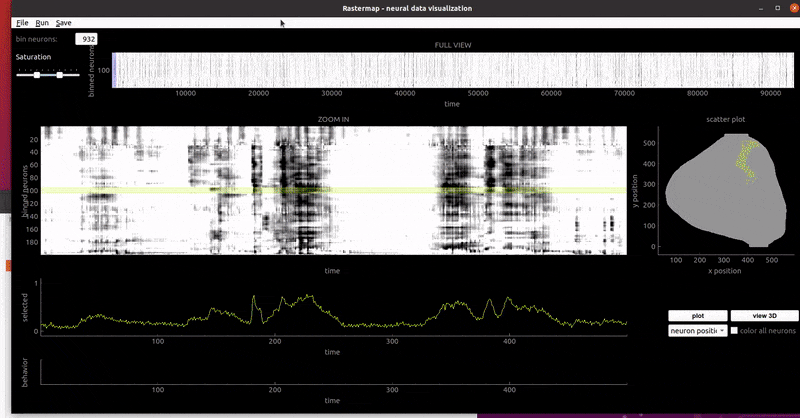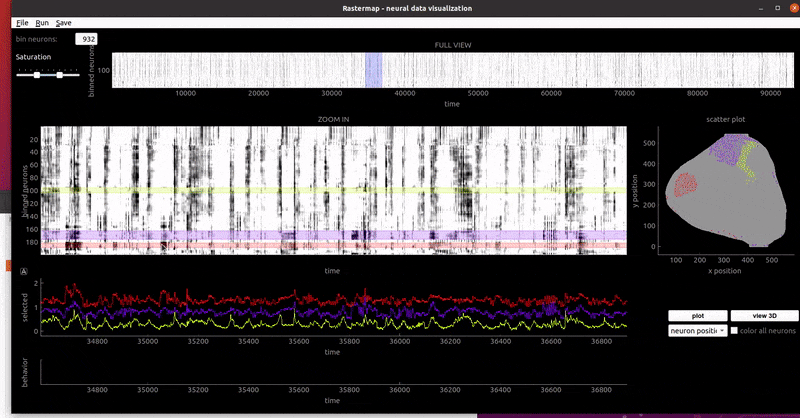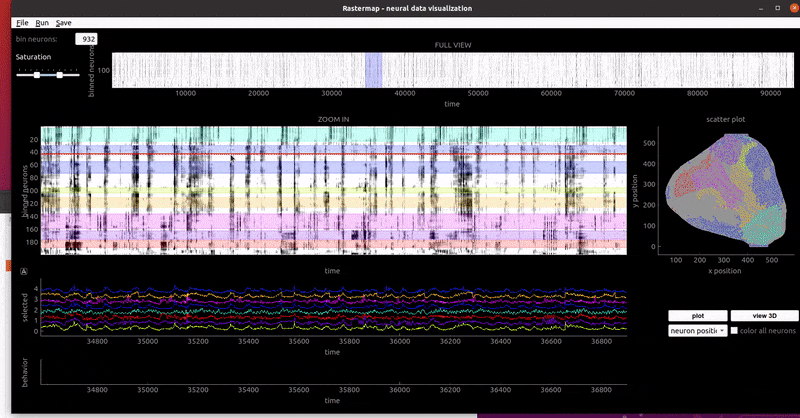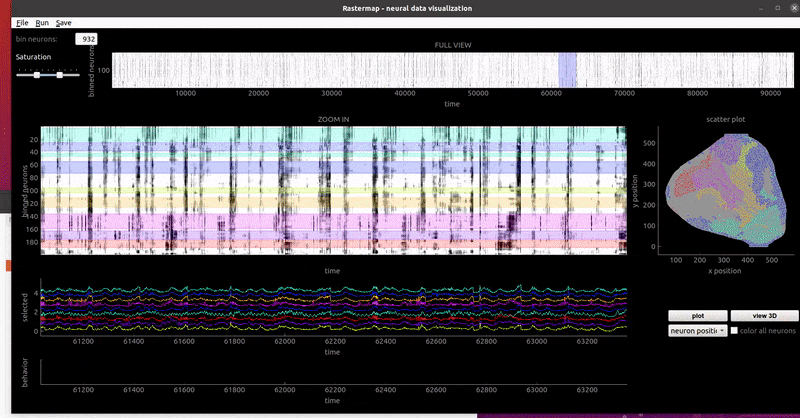
The GUI will start with a highlighted region that you can drag to visualize the average activity of neurons in a given part of the plot. To draw more regions, you right-click to start a region, then right-click to end it. The neurons' activity traces then show up on the botton of the GUI, and if the neuron positions are loaded, you will see them colored by the region color. You can delete a region by holding CTRL and clicking on it. You can save the ROIs you've drawn with the "Save > Save processed data" button. They will save along with the embedding so you can reload the file with the "Load processed data" option.
NOTE: If you are using suite2p "spks.npy", then the GUI will automatically use the "iscell.npy" file in the same folder to subsample your recording with the chosen neurons, and will automatically load the neuron positions from the "stat.npy" file.
GUI examples:
zebrafish:
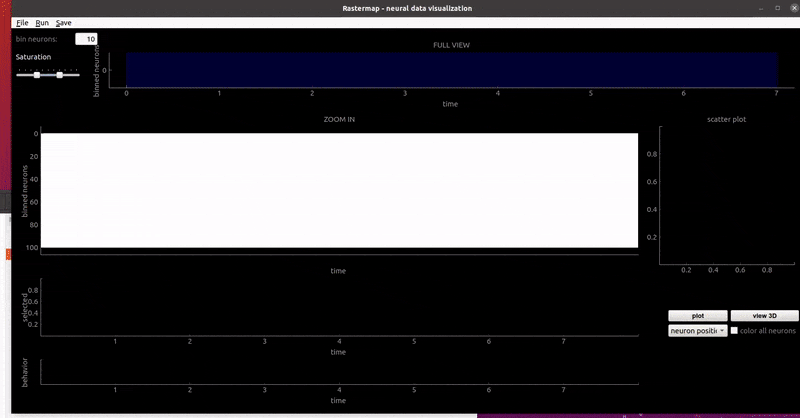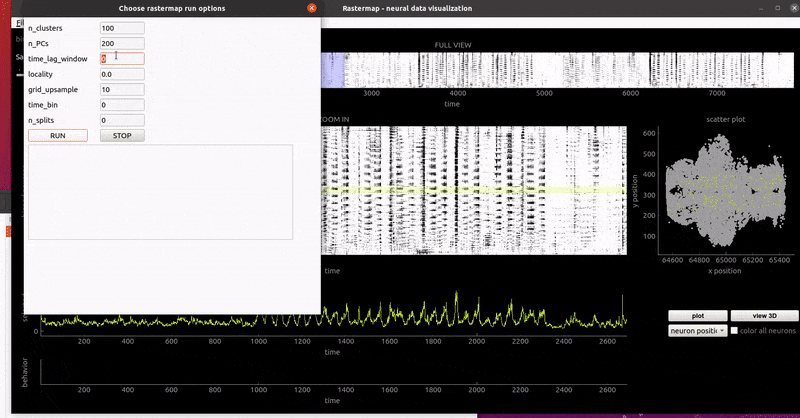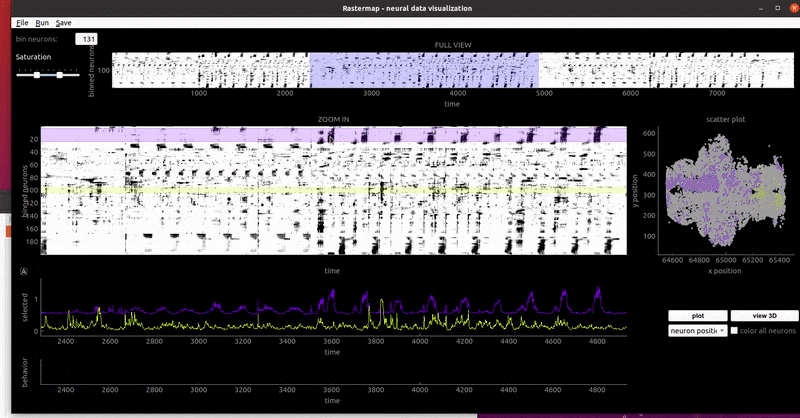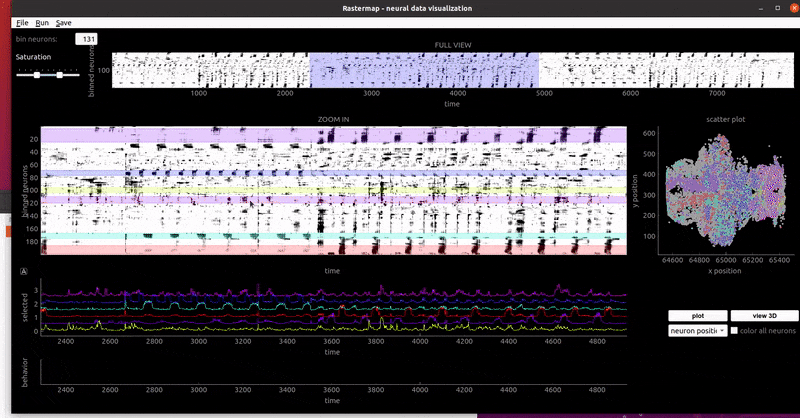
mouse sensorimotor activity:
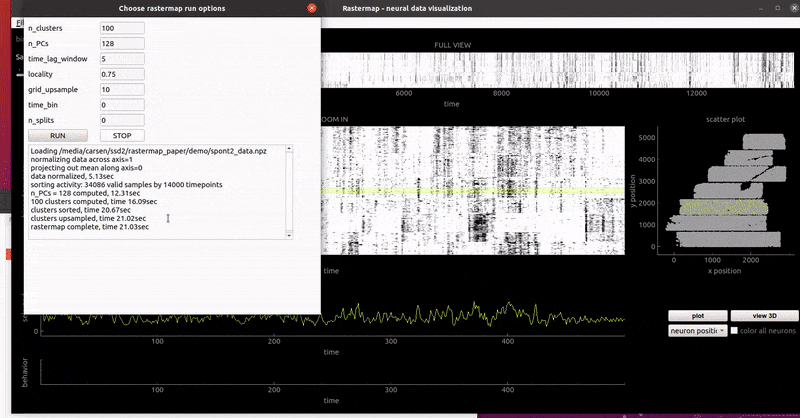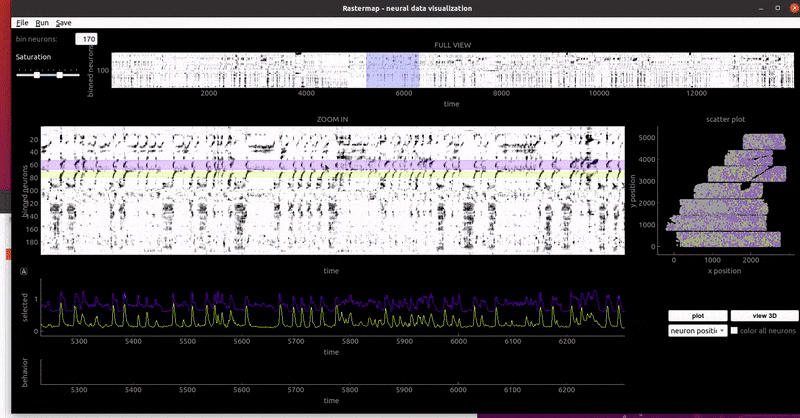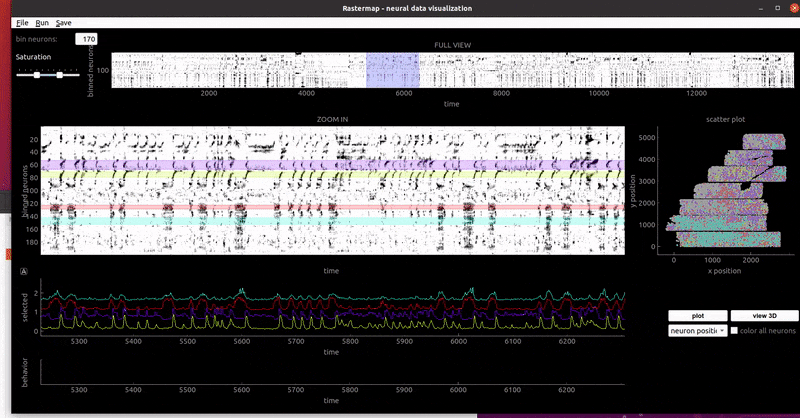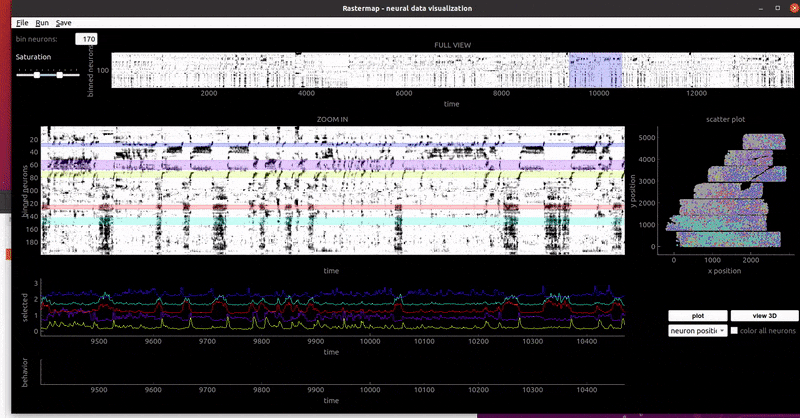
rat hippocampus:
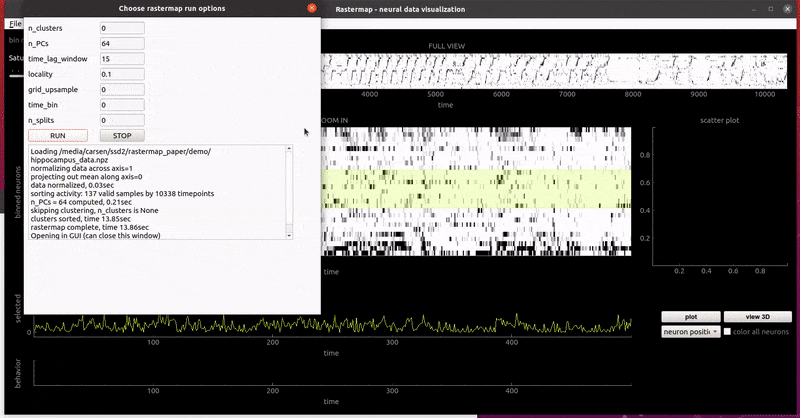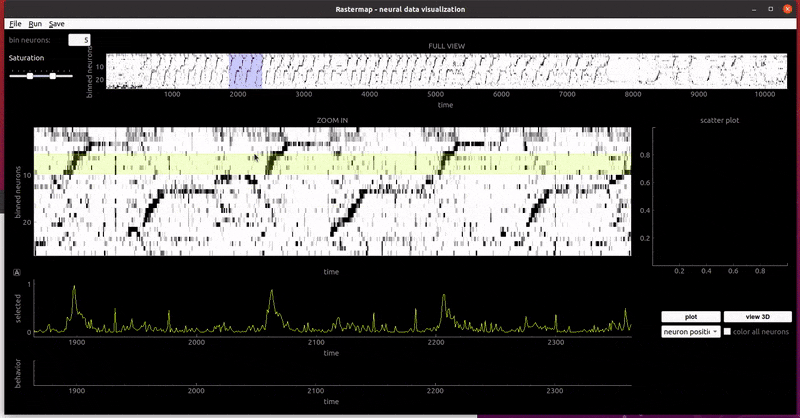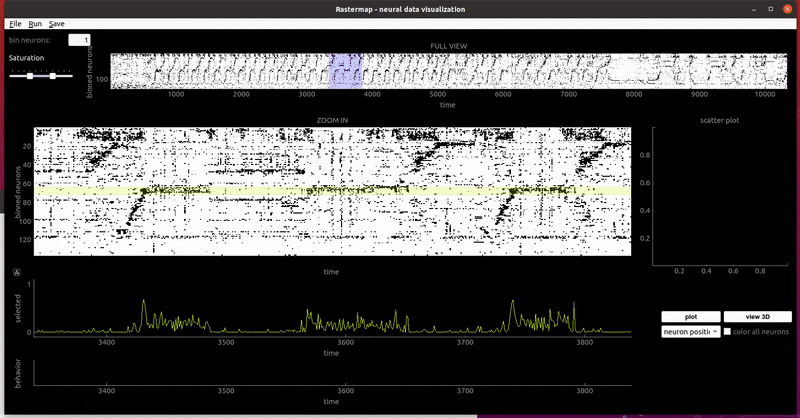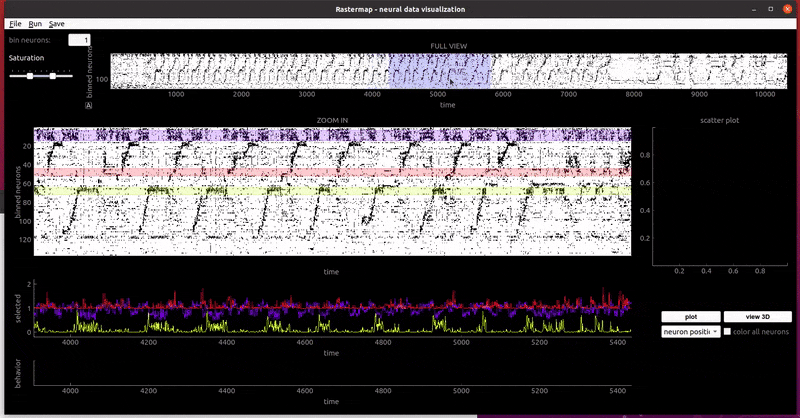
mouse widefield:
In a notebook
For this, pip install notebook and pip install matplotlib. See example notebooks for full examples.
Short example code snippet for running rastermap:
import numpy as np
import matplotlib.pyplot as plt
from rastermap import Rastermap
# spks is neurons by time
spks = np.load("spks.npy").astype("float32")
# fit rastermap
model = Rastermap(n_PCs=200, n_clusters=100,
locality=0.75, time_lag_window=5).fit(spks)
y = model.embedding # neurons x 1
isort = model.isort
# visualize binning over neurons
X_embedding = model.X_embedding
# plot
fig = plt.figure(figsize=(12,5))
ax = fig.add_subplot(111)
ax.imshow(X_embedding, vmin=0, vmax=1.5, cmap="gray_r", aspect="auto")
If you are using google colab, you can mount your google drive and use your data from there with the following command, you will then see your files in the left bar under drive:
from google.colab import drive
drive.mount('/content/drive')
From the command line
Save an "ops.npy" file with the parameters and a "spks.npy" file with a matrix of neurons by time, and run
python -m rastermap --S spks.npy --ops ops.npy
From MATLAB
If you have an existing MATLAB analysis pipeline (and are using MATLAB version R2021b or newer), you can use MATLAB's Python interface to call Rastermap. First you need to tell MATLAB where your Python enviroment with Rastermap is (more details: https://www.mathworks.com/help/matlab/matlab_external/install-supported-python-implementation.html). Create or modify the "PYTHONHOME" environmental variable in your OS to point to the Rastermap environment root folder. Then in MATLAB run the following statement, modified for the specific path to your Rastermap environment pythonw executable:
pyenv('Version','C:\Users\admin\.conda\envs\rastermap\pythonw.exe', 'ExecutionMode', 'OutOfProcess')
Then you should be able to see your environment details in MATLAB after typing "pyenv" (the intepreter will not actually load until you try to run a Python statement). An example function to call Rastermap and return the sort order back as a MATLAB array is:
function [sortIdx] = rastermapSort(shankDataToSort)
% wrapper to convert to numpy array, call Rastermap, and then convert back to MATLAB array
data = shankDataToSort.zScoredFiringRates;
dataNdArray = py.numpy.array(data);
pyrun("from rastermap import Rastermap") %load interpreter, import main function
rmModel = pyrun("model = Rastermap(locality=0.5, time_lag_window=50).fit(spks)", "model", spks=dataNdArray);
sortIdx = int16(py.memoryview(rmModel.isort.data)) + 1; %back to MATLAB array, 1-indexing
end
Inputs
Most of the time you will input to Rastermap().fit a matrix of neurons by time. For more details, these are all the inputs to the function:
- data : array, shape (n_samples, n_features) (optional, default None) this matrix is usually neurons/voxels by time, or None if using decomposition, e.g. as in widefield imaging
- Usv : array, shape (n_samples, n_PCs) (optional, default None) singular vectors U times singular values sv
- Vsv : array, shape (n_features, n_PCs) (optional, default None) singular vectors U times singular values sv
- U_nodes : array, shape (n_clusters, n_PCs) (optional, default None) cluster centers in PC space, if you have precomputed them
- itrain : array, shape (n_features,) (optional, default None) fit embedding on timepoints itrain only
If you have a spike_times.npy and spike_clusters.npy, create your time-binned data
matrix with, where the bin size st_bin is in milliseconds (assuming your spike times are in seconds):
from rastermap import io
# bin spike times into neurons by time matrix
data = io.load_spike_times("spike_times.npy", "spike_clusters.npy", st_bin=100)
You can also load these matrices into the GUI with the File > Load spike_times... option.
Settings
These are inputs to the Rastermap class initialization, the settings are sorted in order of importance
(you will probably never need to change any other than the first few):
- n_clusters : int, optional (default: 100) number of clusters created from data before upsampling and creating embedding (any number above 150 will be slow due to NP-hard sorting problem, max is 200)
- n_PCs : int, optional (default: 200) number of PCs to use during optimization
- time_lag_window : int, optional (default: 0) number of time points into the future to compute cross-correlation, useful to set to several timepoints for sequence finding
- locality : float, optional (default: 0.0) how local should the algorithm be -- set to 1.0 for highly local + sequence finding, and 0.0 for global sorting
- grid_upsample : int, optional (default: 10) how much to upsample clusters, if set to 0.0 then no upsampling
- time_bin : int, optional (default: 0) binning of data in time before PCA is computed, if set to 0 or 1 no binning occurs
- mean_time : bool, optional (default: True) whether to project out the mean over data samples at each timepoint, usually good to keep on to find structure
- n_splits : int, optional (default: 0) split, recluster and sort n_splits times (increases local neighborhood preservation for high-dim data); results in (n_clusters * 2**n_splits) clusters
- run_scaled_kmeans : bool, optional (default: True) run scaled_kmeans as clustering algorithm; if False, run kmeans
- verbose : bool (default: True) whether to output progress during optimization
- verbose_sorting : bool (default: False) output progress in travelling salesman
- keep_norm_X : bool, optional (default: True) keep normalized version of X saved as member of class
- bin_size : int, optional (default: 0) binning of data across n_samples to return embedding figure, X_embedding; if 0, then binning based on data size, if 1 then no binning
- symmetric : bool, optional (default: False) if False, use only positive time lag cross-correlations for sorting (only makes a difference if time_lag_window > 0); recommended to keep False for sequence finding
- sticky : bool, optional (default: True) if n_splits>0, sticky=True keeps neurons in same place as initial sorting before splitting; otherwise neurons can move each split (which generally does not work as well)
- nc_splits : int, optional (default: None) if n_splits > 0, size to split n_clusters into; if None, nc_splits = min(50, n_clusters // 4)
- smoothness : int, optional (default: 1) how much to smooth over clusters when upsampling, number from 1 to number of clusters (recommended to not change, instead use locality to change sorting)
Outputs
The main output you want is the sorting, isort, which is assigned to the Rastermap class, e.g.
model = Rastermap().fit(spks)
isort = model.isort
You may also want to color the neurons by their positions which are in embedding, e.g.
y = model.embedding[:,0]
plt.scatter(xpos, ypos, cmap="gist_rainbow", c=y, s=1)
Here is the list of all variables assigned from fit:
- embedding : array, shape (n_samples, 1) embedding of each neuron / voxel
- isort : array, shape (n_samples,) sorting along first dimension of input matrix - use this to get neuron / voxel sorting
- igood : array, shape (n_samples, 1) neurons/voxels which had non-zero activity and were used for sorting
- Usv : array, shape (n_samples, n_PCs) singular vectors U times singular values sv
- Vsv : array, shape (n_features, n_PCs) singular vectors U times singular values sv
- U_nodes : array, shape (n_clusters, n_PCs) cluster centers in PC space
- Y_nodes : array, shape (n_clusters, 1) np.arange(0, n_clusters)
- X_nodes : array, shape (n_clusters, n_features) cluster activity traces in time
- cc : array, shape (n_clusters, n_clusters) sorted asymmetric similarity matrix
- embedding_clust : array, shape (n_samples, 1) assignment of each neuron/voxel to each cluster (before upsampling)
- X : array, shape (n_samples, n_features) normalized data stored (if keep_norm_X is True)
- X_embedding : array, shape (n_samples//bin_size, n_features) normalized data binned across samples (if compute_X_embedding is True)
The output from the GUI and the command line is a file that ends with _embedding.npy. This file contains:
- filename: str, path to file that rastermap was run on
- save_path: str, folder with filename
- embedding : array, shape (n_samples, 1) embedding of each neuron / voxel
- isort : array, shape (n_samples,) sorting along first dimension of input matrix - use this to get neuron / voxel sorting
- user_clusters: list, list of user drawn clusters in GUI
- ops: dict, dictionary of options used to run rastermap
License
Copyright (C) 2023 Howard Hughes Medical Institute Janelia Research Campus, the labs of Carsen Stringer and Marius Pachitariu.
This code is licensed under GPL v3 (no redistribution without credit, and no redistribution in private repos, see the license for more details).
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file rastermap-1.0.tar.gz.
File metadata
- Download URL: rastermap-1.0.tar.gz
- Upload date:
- Size: 169.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e71fb30346737ecde596cae27ab8be066988d1bf8d1b54601bbb99e72021e2e8
|
|
| MD5 |
d466d4f3d80a5d925ca51c8553c29188
|
|
| BLAKE2b-256 |
31dbe0ed0068934fb588a8e934ebeafb39adbcf9d709a56cb3cb83b3e72be959
|
File details
Details for the file rastermap-1.0-py3-none-any.whl.
File metadata
- Download URL: rastermap-1.0-py3-none-any.whl
- Upload date:
- Size: 90.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.12.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c7ec55d625bc88cf97df50d5a2e068869e5790091ea595120f497cc7fab9370a
|
|
| MD5 |
5f765bf73314add3909de1d5c3190eed
|
|
| BLAKE2b-256 |
c0c801a9eafcc8e2c074c494824f33cc688f6ad4ac98964b6ca5d8c2368246b0
|























