Numerical integration of stochastic differential equations (SDE)
Project description
Overview
sdeint is a collection of numerical algorithms for integrating Ito and Stratonovich stochastic ordinary differential equations (SODEs). It has simple functions that can be used in a similar way to scipy.integrate.odeint() or MATLAB’s ode45.
There already exist some python and MATLAB packages providing Euler-Maruyama and Milstein algorithms, and a couple of others. So why am I bothering to make another package?
It is because there has been 25 years of further research with better methods but for some reason I can’t find any open source reference implementations. Not even for those methods published by Kloeden and Platen way back in 1992. So I will aim to gradually add some improved methods here.
This is prototype code in python, so not aiming for speed. Later can always rewrite these with loops in C when speed is needed.
Warning: this is an early pre-release. Wait for version 1.0. Bug reports are very welcome!
functions
These work with scalar or vector equations. They will choose an algorithm for you. Or you can use a specific algorithm directly:
specific algorithms:
For more information and advanced options for controlling random value generation and repeated integrals see the documentation for each function.
utility functions:
Examples:
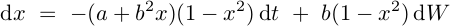
import numpy as np
import sdeint
a = 1.0
b = 0.8
tspan = np.linspace(0.0, 5.0, 5001)
x0 = 0.1
def f(x, t):
return -(a + x*b**2)*(1 - x**2)
def g(x, t):
return b*(1 - x**2)
result = sdeint.itoint(f, g, x0, tspan)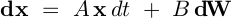
import numpy as np
import sdeint
A = np.array([[-0.5, -2.0],
[ 2.0, -1.0]])
B = np.diag([0.5, 0.5]) # diagonal, so independent driving Wiener processes
tspan = np.linspace(0.0, 10.0, 10001)
x0 = np.array([3.0, 3.0])
def f(x, t):
return A.dot(x)
def G(x, t):
return B
result = sdeint.itoint(f, G, x0, tspan)References for these algorithms:
TODO
Fast, parallel GPU implementation in C++, wrapped with this python interface.
Rewrite Iwik() and Jwik() so they don’t waste so much memory.
Fix stratKP2iS(). In the unit tests it is currently less accurate than itoEuler() and this is likely due to a bug.
Implement the Ito version of the Kloeden and Platen two-step implicit alogrithm.
Add more strong stochastic Runge-Kutta algorithms. Perhaps starting with Burrage and Burrage (1996)
Currently prioritizing those algorithms that work for very general d-dimensional systems with arbitrary noise coefficient matrix, and which are derivative free. Eventually will add special case algorithms that give a speed increase for systems with certain symmetries. That is, 1-dimensional systems, systems with scalar noise, diagonal noise or commutative noise, etc. The idea is that itoint() and stratint() will detect these situations and dispatch to the most suitable algorithm.
Some time in the dim future, implement support for stochastic delay differential equations (SDDEs).
See also:
nsim: Framework that uses this sdeint library to enable massive parallel simulations of SDE systems (using multiple CPUs or a cluster) and provides some tools to analyze the resulting timeseries. https://github.com/mattja/nsim For parallel simulation this will be obsoleted by the GPU implementation in development.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file sdeint-0.3.0.tar.gz.
File metadata
- Download URL: sdeint-0.3.0.tar.gz
- Upload date:
- Size: 40.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.10.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
edee046344abc9350518cc4421e0cd983967cae150d382518bfd84e0306ca64a
|
|
| MD5 |
a943fb2ec20b742b0437314e0b1ba0f7
|
|
| BLAKE2b-256 |
a06f8b6eecefd4d5fba57f58edba98f0050a5823e5c42db918600f370274012d
|











