Long read amplicon analysis
Project description




This is is the laa pipeline from the Sequana project
- Overview:
Perform amplicon analysis on Pacbio data sets including variant and phylogeny
- Input:
A set of CCS files from pacbio in FastQ formats
- Output:
variant calling, phylogney, consensus genomes, etc
- Status:
production but may change
- Citation:
Cokelaer et al, (2017), ‘Sequana’: a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI doi:10.21105/joss.00352
This pipeline was used in :
L’Honneur et al (polyomavirus, 2022) https://pubmed.ncbi.nlm.nih.gov/34979561/
Kali et al (rabies,2021), https://pubmed.ncbi.nlm.nih.gov/33444703/
Claireaux et al. (gene involved in HIV, 2022) https://pubmed.ncbi.nlm.nih.gov/35082297/
Installation
You must install Sequana first:
pip install sequana
Then, just install this package:
pip install sequana_laa
Usage
sequana_laa --help sequana_laa --input-directory DATAPATH
This creates a directory with the pipeline and configuration file. You will then need to execute the pipeline:
cd laa bash laa.sh # for a local run
This launches a snakemake pipeline.
Requirements
This pipelines requires the following executable(s):
vt
freebayes
igvtools
sequana
snpeff (optional)
samtools
bamtools
minimap2
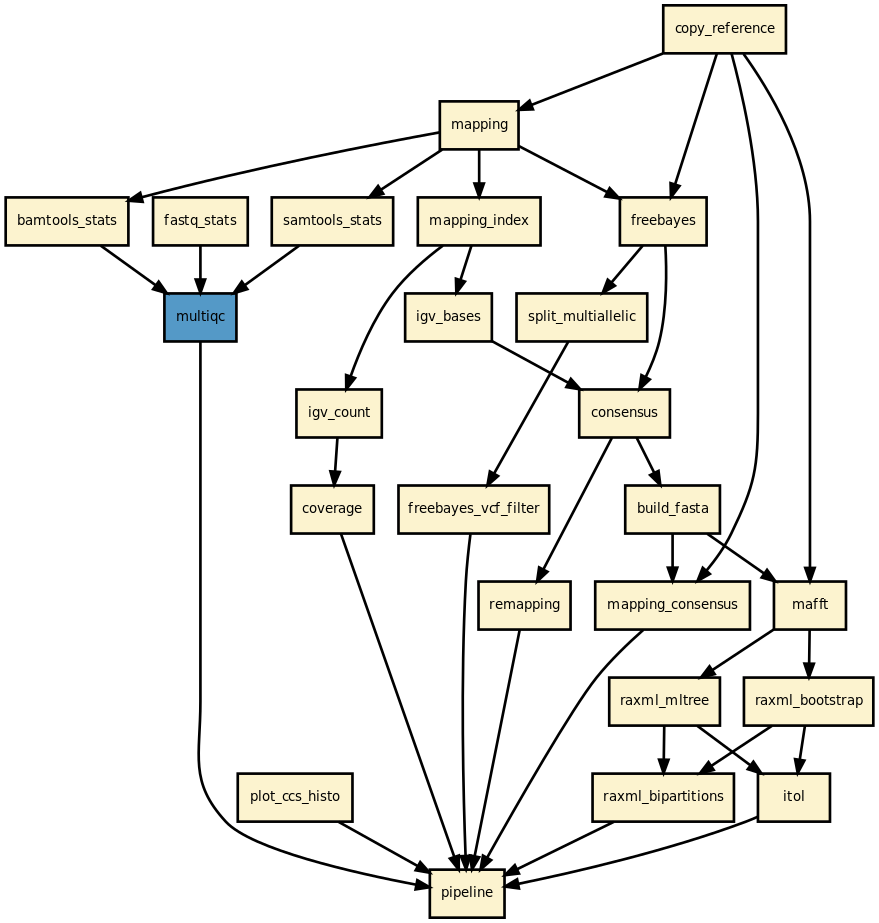
Details
This pipeline runs amplicon analysis on long reads data from pacbio sequencers.
Rules and configuration details
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
Changelog
Version |
Description |
|---|---|
0.12.0 |
|
0.11.0 |
add apptainer containers |
0.10.0 |
full integration with latest wrapeprs and apptainers from damona |
0.9.0 |
add singularity containers |
0.8.0 |
First release. |
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sequana_laa-0.12.0.tar.gz.
File metadata
- Download URL: sequana_laa-0.12.0.tar.gz
- Upload date:
- Size: 107.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
959ca54a6cec9ef32cf8879dacd7cbbe67fbbfe223fb2f2b0a759ecda85e86d6
|
|
| MD5 |
60c58bfa9197345875fdc43c7ec26712
|
|
| BLAKE2b-256 |
6722b099e28c97e580dd41f723224b92d4be6429c4ead9300c5f323a7c6e395e
|
File details
Details for the file sequana_laa-0.12.0-py3-none-any.whl.
File metadata
- Download URL: sequana_laa-0.12.0-py3-none-any.whl
- Upload date:
- Size: 108.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1eac5987d7aa46f30446e534d45758e6df31724bd9015e6fa2bba9f224555863
|
|
| MD5 |
0853a0f58ecfb64d4a178be16736a953
|
|
| BLAKE2b-256 |
0cbd4e7b98246ed5be36bbbe5f63c24b3a9004601136151ec8d7ea144e2f3758
|











