Merge barcoded or non barcoded fastq files generated by Nanopore runs
Project description






This is is the nanomerge pipeline from the Sequana project
- Overview:
merge fastq files generated by Nanopore run and generates raw data QC.
- Input:
individual fastq files generated by nanopore demultiplexing
- Output:
merged fastq files for each barcode (or unique sample)
- Status:
production
- Citation:
Cokelaer et al, (2017), ‘Sequana’: a Set of Snakemake NGS pipelines, Journal of Open Source Software, 2(16), 352, JOSS DOI doi:10.21105/joss.00352
Installation
You can install the packages using pip:
pip install sequana_nanomerge --upgrade
An optional requirements is pycoQC, which can be install with conda/mamba using e.g.:
conda install pycoQC
you will also need graphviz installed.
Usage
sequana_nanomerge --help
If you data is barcoded, they are usually in sub-directories barcoded/barcodeXY so you will need to use a pattern (–input-pattern) such as */*.gz:
sequana_nanomerge --input-directory DATAPATH/barcoded --samplesheet samplesheet.csv
--summary summary.txt --input-pattern '*/*fastq.gz'
otherwise all fastq files are in DATAPATH/ so the input pattern can just be *.fastq.gz:
sequana_nanomerge --input-directory DATAPATH --samplesheet samplesheet.csv
--summary summary.txt --input-pattern '*fastq.gz'
The –summary is optional and takes as input the output of albacore/guppy demultiplexing. usually a file called sequencing_summary.txt
Note that the different between the two is the extra */ before the *.fastq.gz pattern since barcoded files are in individual subdirectories.
In both bases, the command creates a directory with the pipeline and configuration file. You will then need to execute the pipeline:
cd nanomerge bash nanomerge.sh # for a local run
This launches a snakemake pipeline.
Concerning the sample sheet, whether your data is barcoded or not, it should be a CSV file
barcode,project,sample barcode01,main,A barcode02,main,B barcode03,main,C
For a non-barcoded run, you must provide a file where the barcode column can be set (empty):
barcode,project,sample ,main,A
or just removed:
project,sample main,A
Usage with apptainer:
With apptainer, initiate the working directory as follows:
sequana_nanomerge --use-apptainer
Images are downloaded in the working directory but you can store then in a directory globally (e.g.):
sequana_nanomerge --use-apptainer --apptainer-prefix ~/.sequana/apptainers
and then:
cd nanomerge sh nanomerge.sh
if you decide to use snakemake manually, do not forget to add apptainer options:
snakemake -s nanomerge.rules -c config.yaml --cores 4 --stats stats.txt --use-apptainer --apptainer-prefix ~/.sequana/apptainers --apptainer-args "-B /home:/home"
Requirements
This pipelines requires the following executable(s), which is optional:
pycoQC
dot
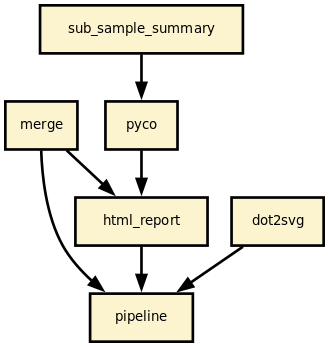
Details
This pipeline runs nanomerge in parallel on the input fastq files (paired or not). A brief sequana summary report is also produced.
Rules and configuration details
Here is the latest documented configuration file to be used with the pipeline. Each rule used in the pipeline may have a section in the configuration file.
Changelog
Version |
Description |
|---|---|
1.6.0 |
|
1.5.0 |
|
1.4.0 |
|
1.3.0 |
|
1.2.0 |
|
1.1.0 |
|
1.0.1 |
|
1.0.0 |
Stable release ready for production |
0.0.1 |
First release. |
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sequana_nanomerge-1.6.0.tar.gz.
File metadata
- Download URL: sequana_nanomerge-1.6.0.tar.gz
- Upload date:
- Size: 28.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
aa5af2efcdcb50e362a2b884657564363e6f73a03a80d52a2a9cfa7954abf968
|
|
| MD5 |
6b918361b399cfc2fa74e02082b477f5
|
|
| BLAKE2b-256 |
095acad5d869ce97038f55a787721bbf719522375884fcef293c6b0f13593ea6
|
File details
Details for the file sequana_nanomerge-1.6.0-py3-none-any.whl.
File metadata
- Download URL: sequana_nanomerge-1.6.0-py3-none-any.whl
- Upload date:
- Size: 28.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dc44be5e67007ee51458d93e90f0d0524742a9cb87ce04a5b24aac5eba7f3488
|
|
| MD5 |
f9afd6c6985855d416ef87c608108bad
|
|
| BLAKE2b-256 |
1dfc25b273bfb28327df6efd62888546e994dd5798eb5e597c8f74cdaef5e70c
|











