Automation framework for the scientific method in R&D
Project description

Usage |
|
Release |
|
Development |
|
Miscellaneous |
Quick Links
Quick Start — Install and run your first experiment
What is SIERRA? — Overview and architecture
Features at a Glance — Supported platforms and capabilities
Why SIERRA? — Motivation and comparison with alternatives
Limitations — What SIERRA is not designed for
Citing — How to cite SIERRA in your research
Troubleshooting — Common issues and how to get help
Contributing — How to contribute to SIERRA
What is SIERRA?
Running large-scale computational experiments is mostly engineering: writing scripts, managing configuration files, wrangling outputs, and rebuilding the same pipeline for every new project.
SIERRA automates this entire workflow.
SIERRA (reSearch pIpEline for Reproducibility, Reusability, and Automation) is a command-line tool and plugin framework that automates the full experimental workflow — generating experiment inputs, executing experiments across heterogeneous computing environments, processing results, and producing analysis artifacts such as plots, videos, and comparative summaries.
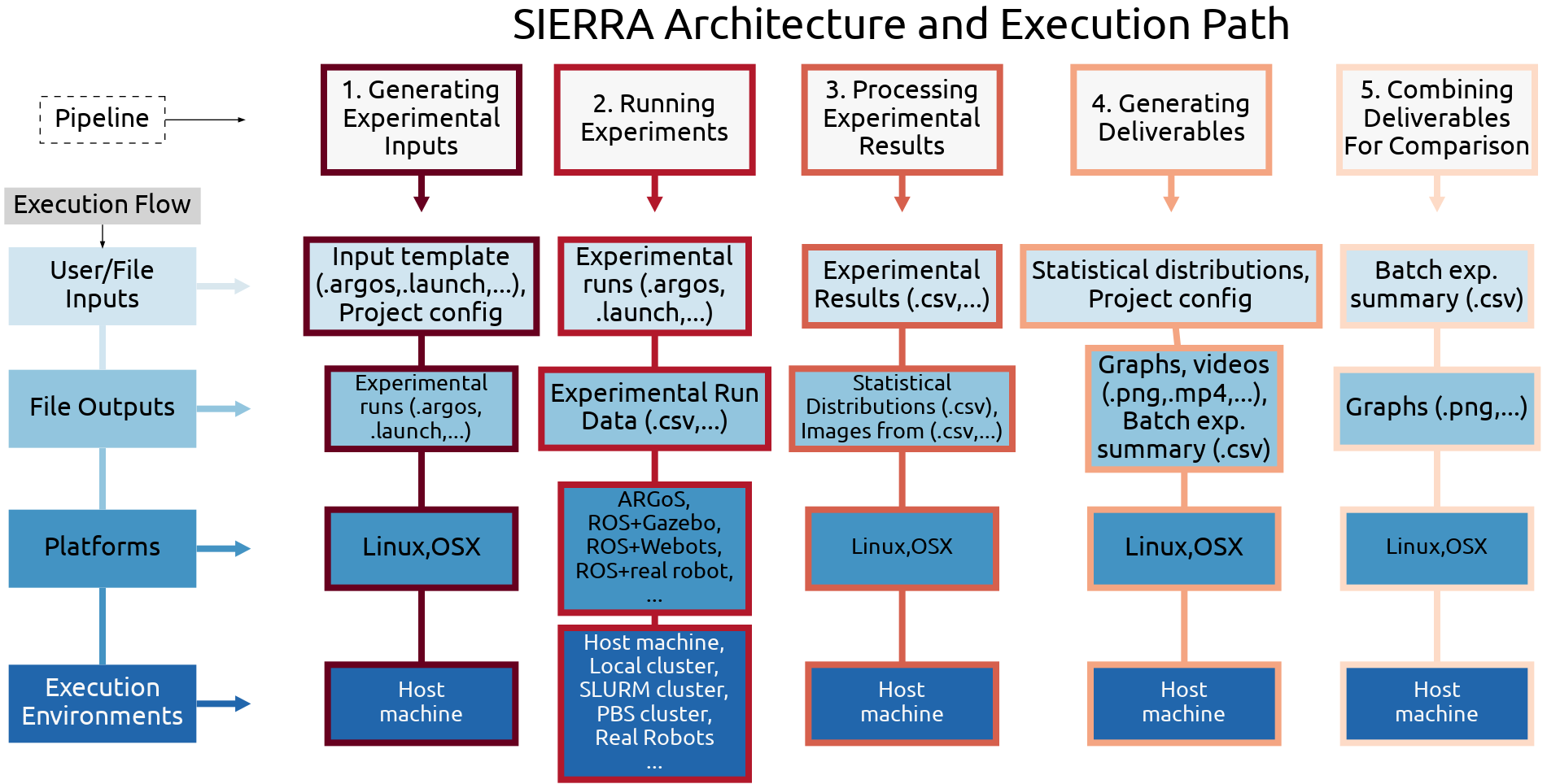
SIERRA architecture, organized by pipeline stage (left to right).
Pipeline Stages
SIERRA organizes experiments into a fixed pipeline:
Input generation — Create experiment configurations from templates
Execution — Run experiments locally, on clusters, etc.
Postprocessing — Parse raw outputs into structured datasets
Product generation — Generate plots, summaries, and derived artifacts
SIERRA is built around three design goals:
Reproducibility — every result is fully described by its path on disk, which encodes the project, controller, scenario, and batch criteria used to produce it. Re-running the same invocation always produces the same layout.
Automation — experiment generation, execution, post-processing, and product generation are handled by the pipeline; no scripting required.
Reusability — the plugin architecture means a project written for one simulator or execution environment requires minimal changes to run on another.
Features at a Glance
All categories below are extensible via plugins.
Feature |
Details |
|---|---|
Supported simulators |
ARGoS, ROS1+Gazebo, ROS1+Robot |
Execution environments |
Local machine, HPC clusters (SLURM, PBS), AWS Batch, Prefect server |
Parameter sweeps |
Numeric, categorical, or mixed combinations |
Output formats |
CSV, Arrow, GraphML, graphs (PNG, HTML), video (mp4) |
Model framework |
Overlay analytical models on empirical results |
Reproducibility |
Fully archived, citable experiment configurations |
Python version |
3.9+ |
Quick Start
Install:
pip3 install sierra-researchMinimal Example
Run a small parameter sweep locally using the built-in sample project.
sierra \
--sierra-root=$HOME/exp \
--project=sierra_sample_project \
--platform=platform.argos \
--exec-env=hpc.local \
--template-input-file=exp/argos/template.argos \
--scenario=LowBlockCount.10x10x1 \
--batch-criteria population_size.Log8 \
--n-runs=2 \
--pipeline 1 2 3 4This generates all experiment inputs, runs them locally, and processes results into graphs. See the getting started guide for a full walkthrough including expected output.
Why SIERRA?
SIERRA changes the paradigm of running experiments from manual and procedural to declarative and automated. Instead of:
"I need to write a script to run the experiment, another script to
process the data and a 3rd script to generate the graphs I want."you describe:
"Here is the environment and/or simulator I want to use, the deliverables
I want to generate, and the data I want on them — GO."SIERRA acts as a backend compiler for research: turning a description of what you want into a fully executed experiment with processed results.
Key advantages:
Deep parameter sweep support — numeric, categorical, or any combination thereof, with no boilerplate. A sweep that would require ~200 lines of brittle, non-reusable bash takes one SIERRA invocation.
Broad platform coverage — mix and match simulators (ARGoS, ROS1+Gazebo), execution environments (local, SLURM, PBS), and output formats with little to no code changes between them.
Maximum reusability — designed so that no copy-pasting is ever needed across projects or platforms.
Rich model framework — run analytical models, generate synthetic data, and overlay it on empirical results automatically on the same figure.
Research-domain focus — built for the scientific workflow, with native concepts for experiments, runs, batch criteria, and replication.
Why SIERRA over Prefect, Dagster, or Airflow?
General-purpose workflow tools like Prefect, Dagster, and Airflow require you to build your own research pipeline from scratch. SIERRA provides a battle-tested pipeline with first-class support for the patterns that matter most in R&D: parameter sweeps, simulator integration, reproducible archiving, and model-vs-empirical comparison.
The trade-off: SIERRA is more opinionated and less feature-complete than those frameworks for general data engineering workloads. For most research use cases, that gap doesn’t matter.
Limitations
SIERRA is purpose-built for the scientific experiment workflow. It is not the right tool if:
You need a general-purpose DAG runner for arbitrary data engineering tasks — use Prefect, Dagster, or Airflow instead.
Your experiments are highly irregular and cannot be expressed as systematic variations on a template.
You need ROS2—ROS1 is supported; ROS2 support is planned.
If you are unsure whether SIERRA fits your use case, check the use cases page or open a discussion thread.
Citing
If you use SIERRA and have found it helpful, please cite the following paper:
@inproceedings{Harwell2022a-SIERRA,
author = {Harwell, John and Lowmanstone, London and Gini, Maria},
title = {SIERRA: A Modular Framework for Research Automation},
year = {2022},
isbn = {9781450392136},
publisher = {International Foundation for Autonomous Agents and Multiagent Systems},
booktitle = {Proceedings of the 21st International Conference on Autonomous Agents and Multiagent Systems},
pages = {1905–1907}
}To cite a specific version of SIERRA (recommended for reproducibility), use the DOI badge in the Miscellaneous row of the badge table above.
Troubleshooting
If you encounter problems using SIERRA, please open an issue or post in the GitHub Discussions forum. Please include your SIERRA version, OS, and a minimal reproducible example where possible.
Contributing
Contributions of all sizes are welcome — bug fixes, documentation improvements, new plugins, or larger features.
A good first contribution is adding a storage plugin or a simple processor plugin — both are typically under 100 lines and have clear contracts defined in the plugin developer guide.
If you have an idea to discuss before diving in, open a discussion thread at any point. See the contributing guide for the full procedure.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sierra_research-1.5.9.tar.gz.
File metadata
- Download URL: sierra_research-1.5.9.tar.gz
- Upload date:
- Size: 6.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
05d04d08dae31658c5e47cbbf89adb541c045a4c2209dc186450039fd2dcf9d8
|
|
| MD5 |
c0a6aa3ee3eaf38886da2bed9febe314
|
|
| BLAKE2b-256 |
49525beb3f43f5a4581f7c51730e3435b36f49cfac1c3a2df0de82136e341d6e
|
Provenance
The following attestation bundles were made for sierra_research-1.5.9.tar.gz:
Publisher:
publish.yml on jharwell/sierra
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
sierra_research-1.5.9.tar.gz -
Subject digest:
05d04d08dae31658c5e47cbbf89adb541c045a4c2209dc186450039fd2dcf9d8 - Sigstore transparency entry: 1085560110
- Sigstore integration time:
-
Permalink:
jharwell/sierra@5b099d80d53a07c376a964870289674aa9fcb0d1 -
Branch / Tag:
refs/tags/1.5.9 - Owner: https://github.com/jharwell
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@5b099d80d53a07c376a964870289674aa9fcb0d1 -
Trigger Event:
release
-
Statement type:
File details
Details for the file sierra_research-1.5.9-py3-none-any.whl.
File metadata
- Download URL: sierra_research-1.5.9-py3-none-any.whl
- Upload date:
- Size: 291.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
39d1fca56155c2db92b83965aaa207b480de92290a113235c7bbfaf8a762ecf7
|
|
| MD5 |
ea376b2a4e318a177d221f1352e8d854
|
|
| BLAKE2b-256 |
4383224cf56dbf12dfcf9b3142cef49631675fd6f9d3d91f4219e23d618eeb7e
|
Provenance
The following attestation bundles were made for sierra_research-1.5.9-py3-none-any.whl:
Publisher:
publish.yml on jharwell/sierra
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
sierra_research-1.5.9-py3-none-any.whl -
Subject digest:
39d1fca56155c2db92b83965aaa207b480de92290a113235c7bbfaf8a762ecf7 - Sigstore transparency entry: 1085560177
- Sigstore integration time:
-
Permalink:
jharwell/sierra@5b099d80d53a07c376a964870289674aa9fcb0d1 -
Branch / Tag:
refs/tags/1.5.9 - Owner: https://github.com/jharwell
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@5b099d80d53a07c376a964870289674aa9fcb0d1 -
Trigger Event:
release
-
Statement type:























