A testbed for simulation-optimization experiments.
Project description
About the Project
SimOpt is a testbed of simulation-optimization problems and solvers. Its purpose is to encourage the development and constructive comparison of simulation-optimization (SO) solvers (algorithms). We are particularly interested in the finite-time performance of solvers, rather than the asymptotic results that one often finds in related literature.
For the purposes of this project, we define simulation as a very general technique for estimating statistical measures of complex systems. A system is modeled as if the probability distributions of the underlying random variables were known. Realizations of these random variables are then drawn randomly from these distributions. Each replication gives one observation of the system response, i.e., an evaluation of the objective function or stochastic constraints. By simulating a system in this fashion for multiple replications and aggregating the responses, one can compute statistics and use them for evaluation and design.
Several papers have discussed the development of SimOpt and experiments run on the testbed:
- Eckman et al. (2024) studies feasibility metrics for stochastically constrained simulation-optimization problems in preparation for introducing related metrics in SimOpt.
- Shashaani et al. (2024) conducts a large data-farming experiment over solver factors to learn relationships between their settings and a solver's finite-time performance.
- Eckman et al. (2023) is the most up-to-date publication about SimOpt and describes the code architecture and how users can interact with the library.
- Eckman et al. (2023) introduces the design of experiments for comparing solvers; this design has been implemented in the latest Python version of SimOpt. For detailed description of the terminology used in the library, e.g., factors, macroreplications, post-processing, solvability plots, etc., see this paper.
- Eckman et al. (2019) describes in detail changes to the architecture of the MATLAB version of SimOpt and the control of random number streams.
- Dong et al. (2017) conducts an experimental comparison of several solvers in SimOpt and analyzes their relative performance.
- Pasupathy and Henderson (2011) describes an earlier interface for MATLAB implementations of problems and solvers.
- Pasupathy and Henderson (2006) explains the original motivation for the testbed.
Code
Python
- The
master branchcontains the source code for the latest stable release of the testbed - The
development branchcontains the latest code for the testbed, but may contain more bugs than the master branch
Matlab
⚠️ MATLAB versions of this testbed are no longer supported
- The
matlab branchcontains a previous stable version of the testbed written in MATLAB
Documentation
Full documentation for the source code can be found on our readthedocs page.
Getting Started
Requirements
- Miniconda or Anaconda
- If you already have a compatible IDE (such as VS Code), we've found that Miniconda will work fine at 1/10 of the size of Anaconda. Otherwise, you may need the Spyder IDE that comes with the full Anaconda distribution.
- It is highly recommended to check the box during installation to add Python/Miniconda/Anaconda to your system PATH.
- If you know you have Python installed but are getting a
Command not founderror when trying to use Python commands, then you may need to add Python to your PATH.
- VS Code (optional)
- This is a lightweight IDE that is compatible with Miniconda.
- Git (optional)
- If you don't have Git installed, you can download the code as a zip file instead
Downloading Source Code
There are two ways to download a copy of the source code onto your machine:
-
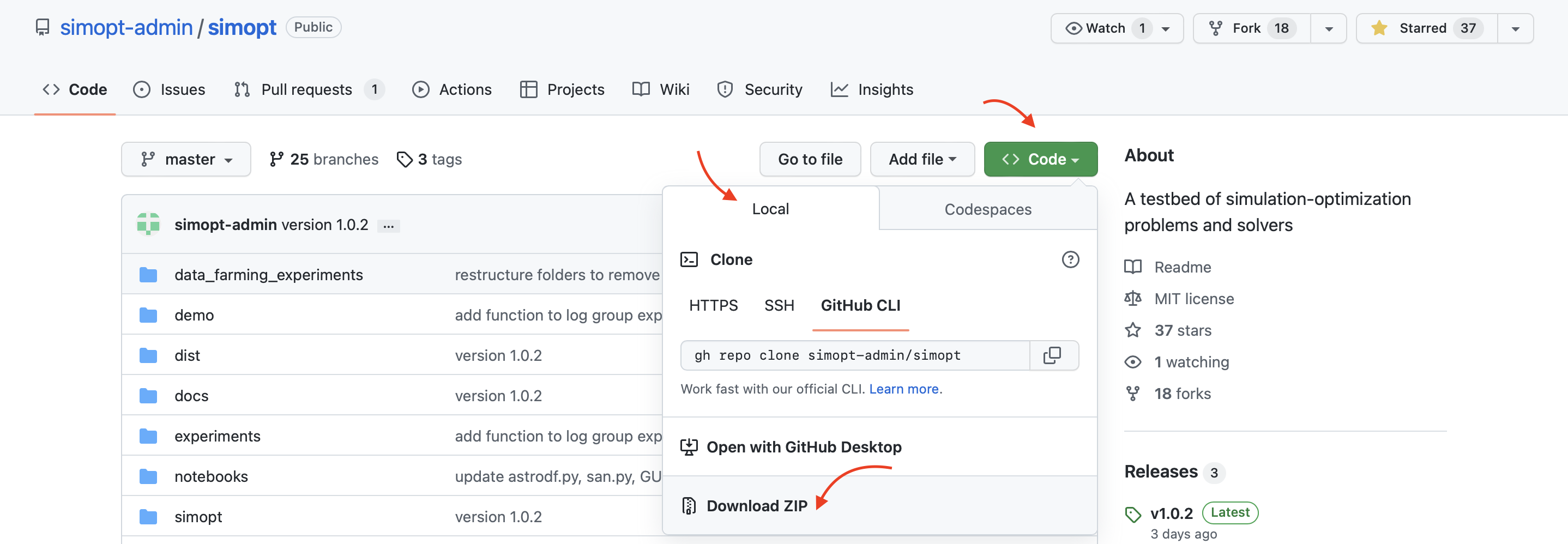
Download the code in a zip file by clicking the green
<> Codebutton above repo contents and clicking theDownload ZIPoption, then unzip the code to a folder on your computer. This does not requiregitto be installed but makes downloading updates to the repository more challenging. -
Clone the branch you'd like to download to a folder on your computer. This requires
gitto be installed but makes downloading updates to the repository much easier.
If you do not need the source code for SimOpt, you may install the library as a Python package instead. See the Package and Basic Example sections for more details about this option.
The notebooks folder includes several useful Jupyter notebooks and scripts that are easy to customize. You can either run the scripts as standalone programs or open the notebooks in JupyterLab or VS Code. A description of the contents is provided below:
| File | Description |
|---|---|
demo_model.py |
Run multiple replications of a simulation model and report its responses |
demo_problem.py |
Run multiple replications of a given solution for an SO problem and report its objective function values and left-hand sides of stochastic constraints |
demo_problem_solver.py |
Run multiple macroreplications of a solver on a problem, save the outputs to a .pickle file in the experiments/outputs folder, and save plots of the results to .png files in the experiments/plots folder |
demo_problems_solvers.py |
Run multiple macroreplications of groups of problem-solver pairs and save the outputs and plots |
demo_data_farming_model.py |
Create a design over model factors, run multiple replications at each design point, and save the results to a comma separated value (.csv) file in the data_farming_experiments folder |
demo_san-sscont-ironorecont_experiment |
Run multiple solvers on multiple versions of (s, S) inventory, iron ore, and stochastic activiy network problems and produce plots |
Environment Setup
After downloading the source code, you will need to configure the conda environment to run the code. This can be done by running the following command in the terminal:
Windows (Command Prompt)
setup_simopt.bat
Windows (PowerShell)
cmd /c setup_simopt.bat
MacOS/Linux
chmod +x setup_simopt.sh && ./setup_simopt.sh
This script will create a new conda environment called simopt and install all necessary packages. To activate the environment, run the following command in the terminal:
conda activate simopt
If you wish to update the environment with the latest compatible packages, you can simply rerun the setup script.
Graphical User Interface (GUI) - User Guide
Main Menu
To open the GUI, run python -m simopt. This will launch the main menu. From the menu, you will be given 2 options:
- Open the
Data Farm Modelswindow - Open the
Simulation Optimization Experimentswindow (read more)
Simulation Optimization Experiments
Overview
From the GUI, you can create a specified problem-solver pair or a problem-solver group, run macroreplications, and generate plots. The main page provides ways to create or continue working with experiments:
- Create an individual problem-solver pair with customized problem and solver factors.
- Load a .pickle file of a previously created problem-solver pair.
- Create a problem-solver group.
At the bottom of the main page, there is a workspace containing all problem-solver pairs and problem-solver groups. The first tab lists the problem-solver pairs ready to be run or post-replicated, the second tab lists the problem-solver groups made from the cross-design or by generating a problem-solver group from partial set of problem-solver pair in the first tab, and the third tab lists those problem-solver pairs that are ready to be post-normalized and prepared for plotting.
1. Creating a problem-solver pair
This is the main way to add problem-solver pairs to the queue in the workspace.
- First, select a solver from the "Solver" dropdown list. Each of the solvers has an abbreviation for the type of problems the solver can handle. Once a solver is selected, the "Problem" list will be sorted and show only the problems that work with the selected solver.
- Change factors associated with the solver as necessary. The first factor is a customizable name for the solver that use can specify.
- All solvers with unique combinations of factors must have unique names, i.e., no two solvers can have the same name, but different factors. If you want to use the same solver twice for a problem but with different solver factors, make sure you change the name of the solver accordingly. For example, if you want to create two problem-solver pairs with the same problem and solver but with or without CRN for the solver, you can change the name of the solver of choice for each pair to reflect that. This name will appear in the queue within the workspace below.
- Select a problem from the "Problem" dropdown list. Each problem has an abbreviation indicating which types of solver is compatible to solve it. The letters in the abbreviation stand for:
| Objective | Constraint | Variable | Direct Gradient Observations |
|---|---|---|---|
| Single (S) | Unconstrained (U) | Discrete (D) | Available (G) |
| Multiple (M) | Box (B) | Continuous (C) | Not Available (N) |
| Deterministic (D) | Mixed (M) | ||
| Stochastic (S) |
- Change factors associated with the problem and model as necessary.
- All problems with unique factors must have unique names, i.e., no two problems can have the same name, but different factors. If you want to use the same problem twice for a solver but with different problem or model factors, make sure you change the name of the problem accordingly. This name will appear in the queue within the workspace below.
- The number of macroreplications can be modified in the top-left corner. The default is 10.
- Select the "Add problem-solver pair" button, which only appears when a solver and problem is selected. The problem-solver pair will be added in the "Queue of problem-solver pairs."
2. Loading a problem-solver pair from a file
Instead of creating a problem-solver pair from scratch, you can load one from a *.pickle file:
- In the top left corner, click "Load a problem-solver pair". Your file system will pop up, and you can navigate to and select an appropriate *.pickle file. The GUI will throw an error if the selected file is not a *.pickle file.
- Once a problem-solver pair object is loaded, it will be added to the "Queue of problem-solver pairs".
- The Run and Post-Process buttons will be updated to accurately reflect whether the problem-solver pair has already been run and/or post-processed.
3. Creating a problem-solver group
Currently, problem-solver groups can only be created within the GUI or command line; they cannot be loaded from a file.
You can create a problem-solver group and add a new item to the "Queue of problem-solver groups" in two ways. The first is a quick grouping of problems and solvers that are compatible with their default factors: of problems and solvers with their default factors.
- Click the "Create a problem-solver group" button.
- Check the compatibility of the Problems and Solvers being selected. Note that solvers with deterministic constraint type can not handle problems with stochastic constraints (e.g., ASTRO-DF cannot be run on FACSIZE-2).
- Specify the number of macroreplications - the default is 10.
- Click "Confirm Cross-Design problem-solver group."
- The pop-up window will disappear, and the problem-solver pairs frame will automatically switch to the "Queue of problem-solver groups".
- To exit out of the problem-solver group pop-up without creating a problem-solver group, click the red "x" in the top-left corner of the window.
The second is converting a list of problem-solver pairs into a problem-solver group by a cross-design:
- Select the problem-solver pairs of interest from the "Queue of problem-solver pairs".
- Clicking the "Convert to a problem-solver group" button. This will complete the cross-design for the partial list and create a new row in the "Queue of problem-solver groups".
Running a problem-solver pair or a problem-solver group
To run a problem-solver pair or a problem-solver group, click the "Run" button in the "Queue of problem-solver pairs" or "Queue of problem-solver groups". Once the problem-solver pair or problem-solver group has been run, the "Run" button becomes disabled. Note: Running a problem-solver pair can take anywhere from a couple of seconds to a couple of minutes depending on the problem-solver pair and the number of macroreplications.
Post-Processing and Post-Normalization
Post-processing happens before post-normalizing and after the run is complete. You can specify the number of post-replications and the (proxy) optimal solution or function value. After post-normalization is complete, the Plotting window appears. To exit out of the Post-Process/Normalize pop-up without post-processing or post-normalizing, click the red "x" in the top-left corner of the window.
- problem-solver pair
problem-solver pairs can be post-processed from the "Queue of problem-solver pairs" tab by clicking "Post-Process." Adjust Post-Processing factors as necessary. Only problem-solver pairs that have already been run can be post-processed. After post-processing, click the "Post-Normalize by Problem" tab to select which problem-solver pairs to post-normalize together.
- Only problem-solver pairs with the same problem can be post-normalized together.
- Once all problem-solver pairs of interest are selected, click the "Post-Normalize Selected" button at the bottom of the GUI (this button only appears when in the Post-Normalize tab).
- In the new pop-up form, update any values necessary and click "Post-Normalize" when the problem-solver pairs are ready to be post-normalized.
- problem-solver group
problem-solver groups are post-processed and post-normalized at the same time. In the "Queue of problem-solver groups" tab, click the "Post-Process" button for the specific problem-solver group, then change any values necessary, then click "Post-Process".
Plotting
The Plotting page is identical for both problem-solver pairs and problem-solver groups. Currently, multiple problem-solver pairs with the same problem can be plotted together, and any problem-solver pair from a single problem-solver group can be plotted together:
- On the left side, select one or more problems from the problem list.
- Select solvers from the solver list.
- On the right side, select a plot type and adjust plot parameters and settings. There are 5 settings common to most plot types: Confidence Intervals, Number of Bootstrap Samples, Confidence Level, Plot Together, and Print Max HW. The type of plots that are currently available in the GUI are: Mean Progress Curve, Quantile Progress Curve, Solve Time CDF, Scatter Plot, CDF Solvability, Quantile Solvability, CDF Difference Plot, Quantile Difference Plot, Terminal Box/Violin, and Terminal Scatter.
- Click "Add."
- All plots will show in the plotting queue, along with information about their parameters and where the file is saved.
- To view one plot, click "View Plot." All plots can be viewed together by clicking "See All Plots" at the bottom of the page.
- To return to the main page, click the red "x" in the top-left corner of the window.
Package
The simoptlib package is available to download through the Python Packaging Index (PyPI) and can be installed from the terminal with the following command:
python -m pip install simoptlib
Basic Example
After installing simoptlib, the package's main modules can be imported from the Python console (or in code):
import simopt
from simopt import models, solvers, experiment_base
The following snippet of code will run 10 macroreplications of the Random Search solver ("RNDSRCH") on the Continuous Newsvendor problem ("CNTNEWS-1"):
myexperiment = simopt.experiment_base.ProblemSolver("RNDSRCH", "CNTNEWS-1")
myexperiment.run(n_macroreps=10)
The results will be saved to a .pickle file in a folder called experiments/outputs. To post-process the results, by taking, for example 200 postreplications at each recommended solution, run the following:
myexperiment.post_replicate(n_postreps=200)
simopt.experiment_base.post_normalize([myexperiment], n_postreps_init_opt=200)
A .txt file summarizing the progress of the solver on each macroreplication can be produced:
myexperiment.log_experiment_results()
A .txt file called RNDSRCH_on_CNTNEWS-1_experiment_results.txt will be saved in a folder called experiments/logs.
One can then plot the mean progress curve of the solver (with confidence intervals) with the objective function values shown on the y-axis:
simopt.experiment_base.plot_progress_curves(
experiments=[myexperiment],
plot_type=simopt.experiment_base.PlotType.MEAN,
normalize=False,
)
The Python scripts in the notebooks folder provide more guidance on how to run common experiments using the library.
One can also use the SimOpt graphical user interface by running the following from the terminal:
python -m simopt
Contributing
You can contribute problems and solvers to SimOpt (or fix other coding bugs) by forking the repository and initiating pull requests in GitHub to request that your changes be integrated.
Authors
The core development team currently consists of
- David Eckman (Texas A&M University)
- Sara Shashaani (North Carolina State University)
- Shane Henderson (Cornell University)
- Cen Wang (Texas A&M University)
- Emily Liu (Cornell University)
Previous maintainer:
- William Grochocinski (North Carolina State University)
Citation
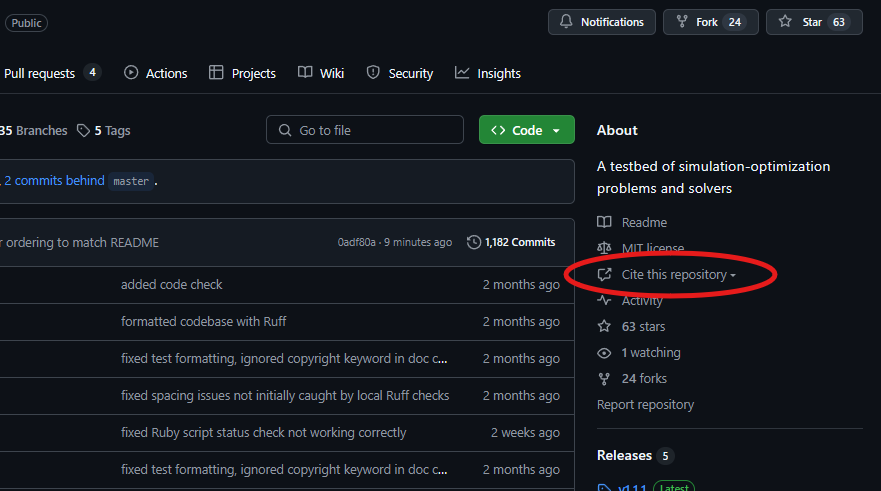
To cite this work, please use the CITATION.cff file or use the built-in citation generator:
Acknowledgments
An earlier website for SimOpt (http://www.simopt.org) was developed through work supported by the following grants:
- National Science Foundation
Recent work on the development of SimOpt has been supported by the following grants
- National Science Foundation
- Air Force Office of Scientific Research
- FA9550-12-1-0200
- FA9550-15-1-0038
- FA9550-16-1-0046
- Army Research Office
- W911NF-17-1-0094
Any opinions, findings and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation (NSF).
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file simoptlib-1.2.4.tar.gz.
File metadata
- Download URL: simoptlib-1.2.4.tar.gz
- Upload date:
- Size: 346.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.11.0 {"installer":{"name":"uv","version":"0.11.0","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":{"name":"CachyOS Linux","version":null,"id":null,"libc":null},"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":null}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f8bed9fc27b0e88673475dd0218cab4277b0fef8ef2481f049564056a3dc93d4
|
|
| MD5 |
84c43fcaa7cf94edeac05baddcb28ac0
|
|
| BLAKE2b-256 |
ea1a0ebc356c800f1705ad478605419c52dc64a6f3de209bd2d0a3ee43ac85eb
|
File details
Details for the file simoptlib-1.2.4-py3-none-any.whl.
File metadata
- Download URL: simoptlib-1.2.4-py3-none-any.whl
- Upload date:
- Size: 392.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.11.0 {"installer":{"name":"uv","version":"0.11.0","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":{"name":"CachyOS Linux","version":null,"id":null,"libc":null},"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":null}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1cbc4ed60910ee3db1887e11d28914a6dee66764927b8bd5541d2285febff67e
|
|
| MD5 |
3c812e4435d931429f6df1f68d9fbea3
|
|
| BLAKE2b-256 |
e4f61d8a02c931935b8841f782411b605a71c108101c8a1c441970ce45521753
|