Particle Swarm and Nelder-Mead Simplex optimization algorithms with Least Squares Regression and Bootstrap confidence intervals.
Project description
Swarm-Simplex-Bootstrap
Swarm-Simplex-Bootstrap is a python implementation of the Particle Swarm and Nelder-Mead Simplex minimization algorithms. Both algorithms make few assumptions about the function to be minimized (such as continuity or differentiability) so they are applicable to a wide variety of problems. Bounds, linear constraints, and nonlinear constraints are supported.
The emphasis of this library is paramterizing models using experimental data where the model parameters can be subject to bounds and constraints. Model parameterization is carried out by minimizing the Least Squares objective function and parameter uncertainty is estimated by Bootstrapping.
Table of Contents
- Installation
- Testing
- Project Structure, Versioning, and Documentation
- General Information
- Test Functions for Minimization
- Unconstrained Minimization with Nelder-Mead
- Bounds
- Bounded Minimization with Nelder-Mead
- Particle Swarm Minimization
- Particle Swarm Followed by Nelder-Mead Refinement
- Constraints
- Bounded and Constrained Minimization
- Model Regression
- Bootstrapping
Installation
ssb_optimize can be installed as a python package using pip. Dependencies include numpy, itertools, numbers, and multiprocessing.
python -m pip install numpy ssb_optimize
ssb_optimize was developed in Python 3.6 so I suspect the package will work with any Python 3 installation. That being said, it hasn't been tested with any other versions of Python (if somebody would like to help with this, please let me know).
Testing
To run unit tests, open a shell environment and run the following command in the top-level directory.
python -m unittest discover -v
Project Structure, Versioning, and Documentation
Project structure follows python-guide.org recommendations.
Docstring format follows Google style recoomendations.
Versioning for publishing to PyPI follows the "major.minor.patch" format based on https://semver.org/ recommendations.
- major version - when you make incompatible API changes,
- minor version - when you add functionality in a backwards-compatible manner, and
- patch version - when you make backwards-compatible bug fixes.
The Markdown cheat sheet is a useful reference for keeping documentation up to date.
General Information
Start off by importing the 'optimizer' module.
import optimizer as opt
Functions contained in the 'optimizer' module are summarized below.
- bounds_check: Check bounds list of size 'n' for consistency and return the list with basic problems corrected.
- constraints_check: Check constraints list for consistency and return the list with basic problems corrected.
- feasible_points_random: Generate an array of points using random sampling which satisfy the bounds and constraints.
- best_point: Return the point corresponding to the lowest evaluated value of 'func'.
- nelder_mead: Minimize a scalar function using the Nelder-Mead simplex algorithm.
- particle_swarm: Minimize a scalar function using the Particle Swarm algorithm.
- least_squares_objective_function: Returns the result of evaluation of the least squares objective function.
- least_squares_bootstrap: Returns a list of the results of repeated least squares fitting of func to random samples taken from x and fx.
The docstring for each function contains additional information about how each function works, arguments, return values, and error handling.
print(function.__doc__)
help(function)
All examples shown in this tutorial are provided in the 'optimizer.py' file for reference.
Test Functions for Minimization
Test functions used in this tutorial are detailed in the Test functions for optimization Wikipedia page. These test functions are used to demonstrate performance of minimization algorithms in situations relevant to real world applications.
Additional information was taken from the following reference.
Jamil, M., & Yang, X. S. (2013). A literature survey of benchmark functions for global optimisation problems. International Journal of Mathematical Modelling and Numerical Optimisation, 4(2), 150. https://doi.org/10.1504/ijmmno.2013.055204
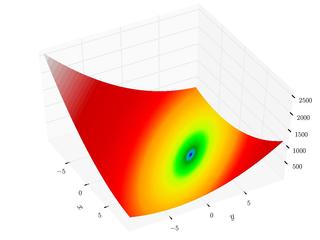
Booth Function
The Booth function is continuous, differentiable, non-separable, non-scalable, and unimodal. It has a smooth approach to the global minimum. This mimics the smooth approach to a minimum for many functions when near a local or global minimum.
Python implementation of the Booth function:
def booth(args):
""" Booth function
Global minimum: f(1.0,3.0) = 0.0
Search domain: -10.0 <= x, y <= 10.0
"""
return (args[0] + 2*args[1] - 7)**2 + (2*args[0] + args[1] - 5)**2
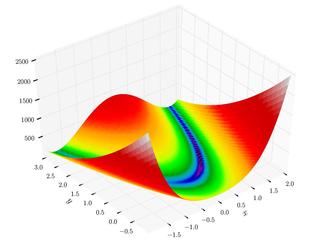
Rosenbrock Function
The Rosenbrock function is continuous, differentiable, non-separable, scalable, and unimodal. The global minimum is inside a long, narrow, parabolic shaped flat valley. To find the valley is trivial but converging to the global minimum is difficult.
Python implementation of the Rosenbrock function:
def rosenbrock(args):
"""Rosenbrock function
Global minimum: f(1,...,1) = 0.0
Search domain: -inf <= xi <= inf, 1 <= i <= n
"""
rosen = 0
for i in range(len(args) - 1):
rosen += 10.0*((args[i]**2) - args[i + 1])**2 + (1 - args[i])**2
return rosen
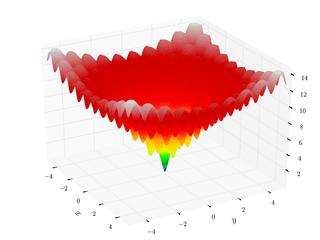
Ackley Function
The Ackley function is continuous, differentiable, non-separable, scalable, and multimodal. It has many local minima but only one one global minimum. Many minimizing algorithms will become trapped in one of the many local minimum during a search for the global minimum.
def ackley(args):
"""Ackley function
Global minimum: f(0,0) = 0.0
Search domain: -5.0 <= x, y <= 5.0
"""
first_sum = 0.0
second_sum = 0.0
for c in args:
first_sum += c ** 2.0
second_sum += np.cos(2.0 * np.pi * c)
n = float(len(args))
return -20.0*np.exp(-0.2*np.sqrt(first_sum/n)) - np.exp(second_sum/n) + 20.0 + np.e
Unconstrained Minimization with Nelder-Mead
The 'nelder_mead' algorithm is a very general direct search minimization method. The main weakness of the 'nelder_mead' algorithm for complex problems is that it can prematurely converge to a local minimum in search of a global minium.
Unconstrained Minimization with Nelder-Mead Simplex, Booth function example (successful convergence)
Input:
initial_pt = [2.0, 2.0]
func = booth
minimum = opt.nelder_mead(initial_pt, func)
print(minimum)
Return:
[1., 3.]
Unconstrained Minimization with Nelder-Mead Simplex, Rosenbrock function example (successful convergence)
Input:
initial_pt = [2.0, 2.0]
func = rosenbrock
minimum = opt.nelder_mead(initial_pt, func)
print(minimum)
Return:
[1., 1.]
Unconstrained Minimization with Nelder-Mead Simplex, Ackley function example (successful convergence)
Input:
initial_pt = [0.1, 0.1]
func = ackley
minimum = opt.nelder_mead(initial_pt, func)
print(minimum)
Return:
[2.39783206e-16, -1.75571593e-16]
Bounds
A bound specifies the minimum and maximum values allowed for each of the 'n' dimensions of the problem space. A set of bounds define a valid region of the problem space. If there are no bounds for a particular dimension of the problem space (i.e. infinity or -infinity), then specify 'None' for that element of the bound.
A valid bounds list is a list of bound tuples or bound lists.
- [(bound_tuple), ... ,(bound_tuple)]
- [[bound_list], ... ,[bound_list]]
Bounds tuples and lists are defined using the following syntax.
- (min, max), (None, max), (min, None), (None, None)
- [min, max], [None, max], [min, None], [None, None]
The 'bounds_check' function checks bounds lists for consistency and returns the list with basic problems corrected.
Bounds specification is optional for the 'nelder_mead' algorithm. However, bounds specification is required for the 'particle_swarm' algorithm because bounds are used to generate the initial particle swarm.
Input:
infinite_bounds = [(None, None), (None, None)]
finite_bounds = [(-5.0, 5.0), (-5.0, 5.0)]
inf_bounds_size = len(infinite_bounds)
fin_bounds_size = len(finite_bounds)
inf_bounds_checked = opt.bounds_check(inf_bounds_size, infinite_bounds)
fin_bounds_checked = opt.bounds_check(fin_bounds_size, finite_bounds)
print(inf_bounds_checked)
print(fin_bounds_checked)
Return:
[[-inf, inf],
[-inf, inf]]
[[-5., 5.],
[-5., 5.]]
Bounded Minimization with Nelder-Mead
Bounded Minimization with Nelder-Mead Simplex, Booth function example (successful convergence)
Input:
initial_pt = [2.0, 2.0]
func = booth
minimum = opt.nelder_mead(initial_pt, func, bounds=finite_bounds)
print(minimum)
Return:
[1., 3.]
Bounded Minimization with Nelder-Mead Simplex, Rosenbrock function example (successful convergence)
Input:
initial_pt = [2.0, 2.0]
func = rosenbrock
minimum = opt.nelder_mead(initial_pt, func, bounds=finite_bounds)
print(minimum)
Return:
[1., 1.]
Bounded Minimization with Nelder-Mead Simplex, Ackley function example (convergence failure)
This starting point is far from the global minimum for the Ackley function. This causes the 'nelder_mead' algorithm to become trapped in a local minimum.
Input:
initial_pt = [2.0, 2.0]
func = ackley
minimum = opt.nelder_mead(initial_pt, func, bounds=finite_bounds)
print(minimum)
Return:
[1.97445199, 1.97445199]
Bounded Minimization with Nelder-Mead Simplex, Ackley function example (successful convergence)
This starting point is close to the global minimum for the Ackley function. There are no local minima between this starting point and the global minimum. The 'nelder_mead' algorithm quickly converges to the global minimum.
Input:
initial_pt = [0.1, 0.1]
func = ackley
minimum = opt.nelder_mead(initial_pt, func, bounds=finite_bounds)
print(minimum)
Return:
[ 2.39783206e-16, -1.75571593e-16]
Particle Swarm Minimization
The 'particle_swarm' algorithm is an evolutionary minimization method. The main strength of the 'particle_swarm' algorithm is that it effectively identifies the global minimum in problem spaces that contain many other local minima. Though the algorithm can identify the global minimum for MOST problems problems, there is no guarantee that it will identify the global minimum for EVERY problem. The main weakness of the 'particle_swarm' algorithm is that it is not efficient at converging to a tight estimate of the global minimum (after the neighborhood of the global minimum is identified).
Bounds specification is required for this implementation of the 'particle_swarm' algorithm because bounds are used to generate the initial particle swarm.
Bounded Minimization with Particle Swarm, Booth function example (successful convergence)
Input:
func = booth
minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds)
print(minimum)
Return:
[0.99999974, 3.00000191]
Bounded Minimization with Particle Swarm, Rosenbrock function example (successful convergence)
Input:
func = rosenbrock
minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds)
print(minimum)
Return:
[0.99999155, 0.99998029]
Bounded Minimization with Particle Swarm, Ackley function example (successful convergence)
Input:
func = ackley
minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds)
print(minimum)
Return:
[-3.59268130e-11, 4.02149815e-10]
Particle Swarm Followed by Nelder-Mead Refinement
The 'particle_swarm' and 'nelder_mead' algorithms can be used together to efficiently minimize complex objective functions. The 'particle_swarm' algorithm is used first to find an estimate for the neighborhood of the global minimum (loose convergence critera is used). The 'particle_swarm' algorithm also generates an estimate for the initial size of the simplex in the 'nelder_mead' algorithm. The initial estimate of the global minimum and simplex size are then passed to the 'nelder_mead' algorithm. The 'nelder_mead' algorithm will converge to a tight estimate of the global minimum. Though this sequential procedure can identify the global minimum for MOST problems problems, there is no guarantee that it will identify the global minimum for EVERY problem.
Bounded Minimization with Combined Procedure, Booth function example (successful convergence)
Input:
func = booth
ps_minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds)
nm_minimum = opt.nelder_mead(ps_minimum, func, bounds=finite_bounds, initial_size=nelder_mead_initial_size)
print(nm_minimum)
Return:
[1., 3.]
Bounded Minimization with Combined Procedure, Rosenbrock function example (successful convergence)
Input:
func = rosenbrock
ps_minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds)
nm_minimum = opt.nelder_mead(ps_minimum, func, bounds=finite_bounds, initial_size=nelder_mead_initial_size)
print(nm_minimum)
Return:
[1., 1.]
Bounded Minimization with Combined Procedure, Ackley function example (successful convergence)
Input:
func = ackley
ps_minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds)
nm_minimum = opt.nelder_mead(ps_minimum, func, bounds=finite_bounds, initial_size=nelder_mead_initial_size)
print(nm_minimum)
Return:
[-2.48273518e-16, 1.10150570e-15]
Constraints
Constraints are functional requirements that define valid regions of the problem space. A complete constraint specification must include a constraint function, any optional arguments (args and kwargs), and constraint type (inequality type). More than one constraint can be specified to define a problem space.
A valid constraints list is a list of constraint dictionaries.
- [{const_dict}, ... ,{const_dict}]
- {'type': ineq_spec_string, 'func': callable_func, 'args': (args_tuple), 'kwargs': {kwargs_dict}}
The 'constraints_check' function checks constraints lists for consistency and returns the list with basic problems corrected.
Input:
def cubic(args):
"""Cubic curve"""
return (args[0] - 1.0) ** 3 - args[1] + 1.0
def line(args):
"""Line"""
return args[0] + args[1] - 2.0
def circle(args):
"""Circle"""
return args[0]**2 + args[1]**2 - 2.0
const_a = [{'type': '<=0', 'func': cubic},
{'type': '<=0', 'func': line}]
const_b = [{'type': '<=0', 'func': circle}]
const_a_checked = opt.constraints_check(const_a)
const_b_checked = opt.constraints_check(const_b)
print(const_a_checked)
print(const_b_checked)
Return:
[{'type': '<=0', 'func': <function func_a at 0x0352AC90>, 'args': (), 'kwargs': {}},
{'type': '<=0', 'func': <function func_b at 0x0352A348>, 'args': (), 'kwargs': {}}]
[{'type': '<=0', 'func': <function func_c at 0x0351FAE0>, 'args': (), 'kwargs': {}}]
Bounded and Constrained Minimization
It is straight forward to minimize a function after bounds and constraints have been specified. Both the 'nelder_mead' and 'particle_swarm' algorithms can be used with bounds and constraints. However, the combined procedure ('particle_swarm' followed by 'nelder_mead' refinement) is recommended.
Bounded and Constrained Minimization with Combined Procedure, Booth function example (successful convergence)
The minimum of the booth function subject to bounds and constraints is no longer the global minimum of the unconstrained booth function. The minimum lies on the edge of the constrained problem space.
Input:
func = booth
ps_minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds, constraints=const_a)
nm_minimum = opt.nelder_mead(ps_minimum, func, bounds=finite_bounds, constraints=const_a,
initial_size=nelder_mead_initial_size)
print(nm_minimum)
Return:
[-0.00337825, 2.00337825]
Bounded and Constrained Minimization with Combined Procedure, Booth function example (successful convergence)
The minimum of the booth function subject to bounds and constraints is no longer the global minimum of the unconstrained booth function. The minimum lies on the edge of the constrained problem space.
Input:
func = booth
ps_minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds, constraints=const_b)
nm_minimum = opt.nelder_mead(ps_minimum, func, bounds=finite_bounds, constraints=const_b,
initial_size=nelder_mead_initial_size)
print(nm_minimum)
Return:
[0.90574948, 1.08610215]
Bounded and Constrained Minimization with Combined Procedure, Rosenbrock function example (successful convergence)
The minimum of the rosenbrock function subject to bounds and constraints is still the global minimum of the unconstrained rosenbrock function. The global minimum lies right on the edge of the constrained problem space.
Input:
func = rosenbrock
ps_minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds, constraints=const_a)
nm_minimum = opt.nelder_mead(ps_minimum, func, bounds=finite_bounds, constraints=const_a,
initial_size=nelder_mead_initial_size)
print(nm_minimum)
Return:
[1., 1.]
Bounded and Constrained Minimization with Combined Procedure, Rosenbrock function example (successful convergence)
The minimum of the rosenbrock function subject to bounds and constraints is still the global minimum of the unconstrained rosenbrock function. The global minimum lies right on the edge of the constrained problem space.
Input:
func = rosenbrock
ps_minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds, constraints=const_b)
nm_minimum = opt.nelder_mead(ps_minimum, func, bounds=finite_bounds, constraints=const_b,
initial_size=nelder_mead_initial_size)
print(nm_minimum)
Return:
[1., 1.]
Bounded and Constrained Minimization with Combined Procedure, Ackley function example (successful convergence)
Input:
func = ackley
ps_minimum, nelder_mead_initial_size = opt.particle_swarm(func, bounds=finite_bounds, constraints=const_a)
nm_minimum = opt.nelder_mead(ps_minimum, func, bounds=finite_bounds, constraints=const_a,
initial_size=nelder_mead_initial_size)
print(nm_minimum)
Return:
[-2.20115198e-15, -1.85442232e-15]
Model Regression
The least squares objective function is the core of regression. This implementation of the least squares objective function facilitates weights as well as bootstrapping. The difference between 'fx' and 'func(x, theta)' is a measure of the goodness of fit. Minimizing this difference by adjusting 'theta' is how 'func' is regressed to fit the data set ('x' and 'fx').
Function to be Fit
def quadratic(x, a, b, c):
"""General quadratic function"""
return a*x**2 + b*x + c
'x' and 'fx' Vectors
# 'x' vector.
x = [0.0, 0.2, 0.4, 0.6, 0.8, 1.0, 1.2, 1.4, 1.6, 1.8, 2.0, 0.1, 0.3, 0.5, 0.7, 0.9, 1.1, 1.3, 1.5]
# 'fx' values
fx = [0.43, 0.27, 0.59, 0.62, 1.43, 2.04, 2.88, 3.96, 4.9, 6.27, 8.05, 0.3, 0.3, 0.48, 0.98, 1.51, 2.14, 3.1, 4.72]
Basic Least Squares Regression
Input:
func = quadratic
theta_initial_guess = [2.1, -0.4, 0.3]
theta = opt.nelder_mead(theta_initial_guess, opt.least_squares_objective_function, args=(func, x, fx))
print(theta)
Return:
[2.24925897, -0.67474865, 0.35668829]
Weighted Least Squares Regression
Weights can be added which allows additional influence to be attached to certain data points. Even weights for each term will yield the same result as unweighted least squares.
Input:
even_weight = [0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5, 0.5]
theta = opt.nelder_mead(theta_initial_guess, opt.least_squares_objective_function, args=(func, x, fx, even_weight))
print(theta)
Return:
[2.24925897, -0.67474865, 0.35668829]
Uneven weights will emphasize certain terms which impacts the regression result. Uneven weights are commonly encountered when fitting a model to experimental data where uncertainty is different for each measurement.
Input:
uneven_weight = [0.3, 0.4, 0.5, 0.3, 0.4, 0.5, 0.3, 0.4, 0.5, 0.3, 0.4, 0.5, 0.3, 0.4, 0.5, 0.3, 0.4, 0.5, 0.3]
theta = opt.nelder_mead(theta_initial_guess, opt.least_squares_objective_function, args=(func, x, fx, uneven_weight))
print(theta)
Return:
[2.24874079, -0.68614556, 0.36575071]
Bootstrapping
The 'least_squares_bootstrap' function drives repeated evaluation of the 'least_squares_objective_function' where input parameters for each evaluation are sampled from 'x', 'fx', and 'weight' with replacement. The bootstrapping technique uses the results (i.e. fitted model parameters) from each repeat evaluation to derive summary statistics which describe the overall result set (i.e. fitted model parameters with their uncertainties).
Input:
bootstrap_set = opt.least_squares_bootstrap(theta_initial_guess, func, x, fx,
weight=None,
bounds=None, constraints=None,
multiprocess=False,
samples=100, max_iter=1000)
print(bootstrap_set)
Return:
[[2.11986049, -0.40819355, 0.33375504]
[2.31068123, -0.80855933, 0.39918449]
[2.39058295, -0.95418734, 0.40050302]
[2.23447189, -0.67165168, 0.35552198]
[2.26151338, -0.59360129, 0.25014203]
[2.11059271, -0.41854009, 0.31450327]
[2.24066416, -0.55588869, 0.28528897]
[2.37982065, -0.87770428, 0.37037564]
[2.15489980, -0.60434848, 0.37946051]
[2.26765001, -0.74801637, 0.42275602]
[2.36032456, -0.93812904, 0.44158196]
[2.20161137, -0.54674909, 0.32233909]
[2.68675659, -1.20049127, 0.41312043]
[2.23992883, -0.73049307, 0.39298691]
[2.42285334, -1.02007571, 0.41370413]
[2.00304294, -0.14472707, 0.11966633]
[2.04840949, -0.23959399, 0.27271515]
[1.98548708, -0.24492065, 0.27279987]
[2.11948575, -0.36589710, 0.22541193]
[2.30513469, -0.81350200, 0.38391252]
[2.15453856, -0.53658969, 0.39341044]
[2.27455759, -0.75397887, 0.34604839]
[2.29431153, -0.76836397, 0.36190855]
[2.30590514, -0.81572687, 0.37263515]
[2.21988265, -0.65713741, 0.36846049]]
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file ssb_optimize-0.1.3.tar.gz.
File metadata
- Download URL: ssb_optimize-0.1.3.tar.gz
- Upload date:
- Size: 27.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/2.0.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.4.0 requests-toolbelt/0.9.1 tqdm/4.36.1 CPython/3.6.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5c0c3c1681d7dc64e3732ccd7d233ba29fb3e0bb88be676a2779a834ed5d3248
|
|
| MD5 |
6449f3a5c1f1273c50c2594e078b0aab
|
|
| BLAKE2b-256 |
927449015e6c6f3059dbd60a163970a40167f092f12eafacf22a389bc52e81f1
|
File details
Details for the file ssb_optimize-0.1.3-py3-none-any.whl.
File metadata
- Download URL: ssb_optimize-0.1.3-py3-none-any.whl
- Upload date:
- Size: 29.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/2.0.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.4.0 requests-toolbelt/0.9.1 tqdm/4.36.1 CPython/3.6.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dbbc04bf9c2accf6d8f334306a0eaa9da25e042bd7205c324025725416c29bc1
|
|
| MD5 |
0bb60946217e545d9578c6788a592e9e
|
|
| BLAKE2b-256 |
50c70b96024385016ec9b89f5c2e7a8e9ba02d1bf5f1343f775a39e5ed3ba103
|