A collection of tools for analysing cmdspanpy output, written in Python
Project description
A Python library for analysing cmdstanpy output
This is a collection of functions for analysing output of cmdstanpy library. The main idea is to do a quick data analysis by calling a single function that makes:
-
traceplots of samples,
-
text and plots of the summaries of model parameters,
-
histograms and pair plots of posterior distributions of parameters.

The only known illustration of a tarpan made from life, depicting a five month old colt (Borisov, 1841). Source: Wikimedia Commons.
Setup
First, run:
pip install tarpan
Finally, install cmdstan by running:
install_cmdstan -v 2.21.0
Complete analysis: save_analysis
This is the main function of the library that saves summaries and trace/pair/tree plots in model_info directory. The function is useful when you want to generate all types of summaries and plots at once.
from tarpan.cmdstanpy.analyse import save_analysis
model = CmdStanModel(stan_file="your_model.stan")
fit = model.sample(data=your_data)
save_analysis(fit, param_names=['mu', 'sigma'])
If you don't need everything, you can call individual functions described below to make just one type of plot or a summary.
Analysis without cmdstanpy
Here is how to analyse values from Pandas' data frame columns:
from tarpan.shared.analyse import save_analysis
save_analysis(df, param_names=['mu', 'sigma'])
Summary: save_summary
Creates a summary of parameter distributions and saves it in text and CSV files.
from tarpan.cmdstanpy.summary import save_summary
model = CmdStanModel(stan_file="your_model.stan")
fit = model.sample(data=your_data)
save_summary(fit, param_names=['mu', 'tau', 'eta.1'])
-
Call
print_summaryto print summary to console instead of saving to a file.
The text summary format is such that the text can be pasted into Github/Gitlab/Bitbucket's Markdown file, like this:
| Name | Mean | Std | Mode | + | - | 68CI- | 68CI+ | 95CI- | 95CI+ | N_Eff | R_hat |
|---|---|---|---|---|---|---|---|---|---|---|---|
| mu | 8.05 | 5.12 | 7.53 | 4.63 | 4.59 | 2.93 | 12.16 | -1.84 | 18.74 | 1540 | 1.00 |
| tau | 6.41 | 5.72 | 2.36 | 5.41 | 2.35 | 0.00 | 7.76 | 0.00 | 17.07 | 1175 | 1.00 |
| eta.1 | 0.39 | 0.92 | 0.60 | 0.71 | 1.13 | -0.53 | 1.31 | -1.48 | 2.19 | 3505 | 1.00 |
Summary columns
-
Name, Mean, Std are the name of the parameter, its mean and standard deviation.
-
68CI-, 68CI+, 95CI-, 95CI+ are the 68% and 95% HPDIs (highest posterior density intervals). These values are configurable.
-
Mode, +, - is a mode of distribution with upper and lower uncertainties, which are calculated as distances to 68% HPDI.
-
N_Eff is Stan's number of effective samples, the higher the better.
-
R_hat is a Stan's parameter representing the quality of the sampling. This value needs to be around 1.00. After generating a model I usually immediately look at this R_hat column to see if the sampling was good.
Summary without cmdstanpy
Here is how to make summary of values from Pandas' data frame columns:
from tarpan.shared.summary import save_summary
save_summary(df, param_names=['mu', 'sigma'])
Tree plot: save_tree_plot
This function shows exactly the same information as save_summary, but in
the form a plot. The markers are the modes of the distributions, and the two error bars
indicate 68% and 95% HPDIs (highest posterior density intervals).
from tarpan.cmdstanpy.tree_plot import save_tree_plot
model = CmdStanModel(stan_file="your_model.stan")
fit = model.sample(data=your_data)
save_tree_plot([fit], param_names=['mu', 'sigma'])
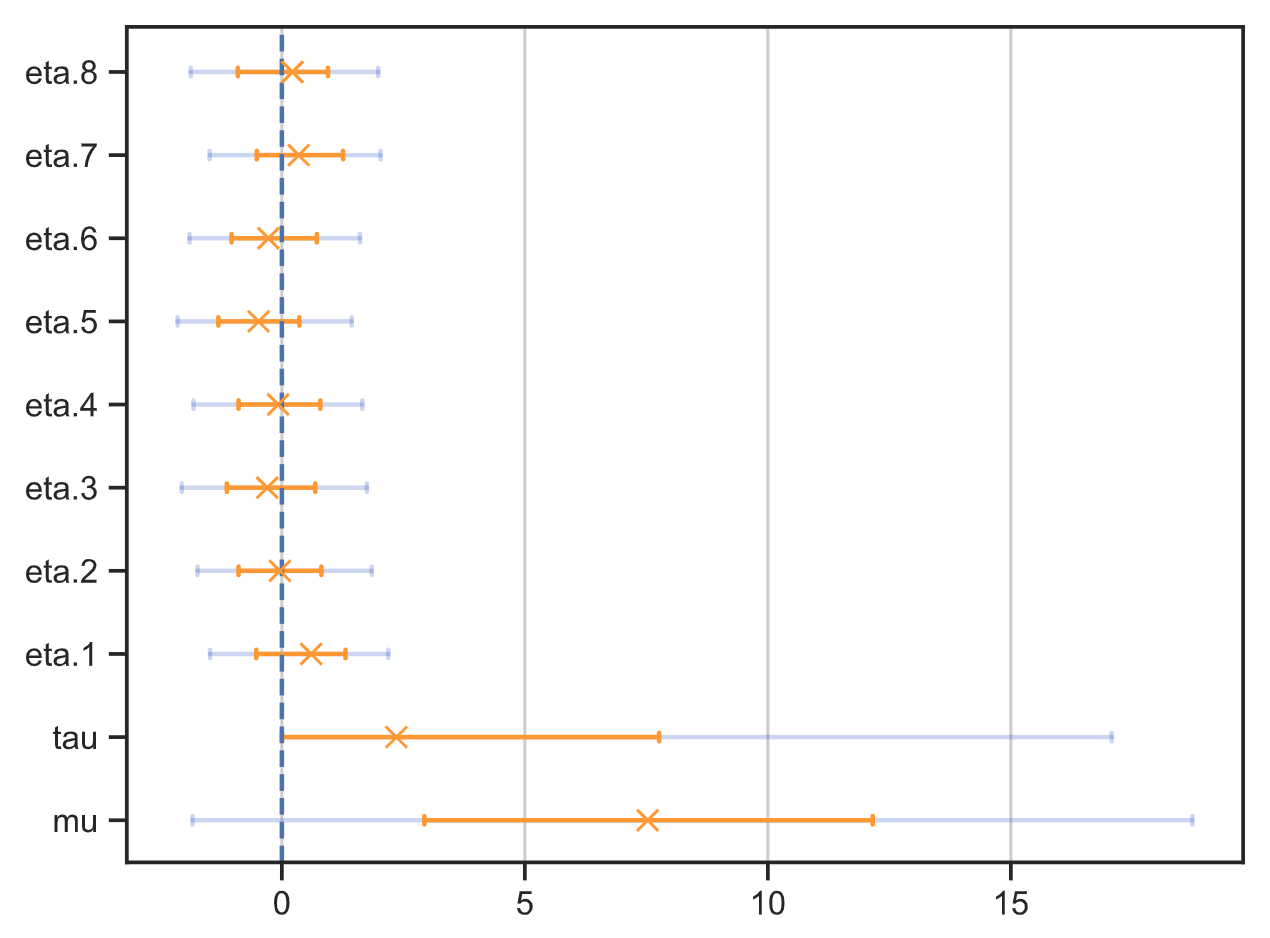
Tree plot without cmdstanpy
One can make a tree plot by supplying a Panda's data frame that shows summaries of values for the frame's columns:
from tarpan.shared.tree_plot import save_tree_plot
save_tree_plot([df], param_names=['mu', 'sigma'])
Comparing multiple models on a tree plot
Supply multiple fits in order to compare parameters from multiple models.
from tarpan.cmdstanpy.tree_plot import save_tree_plot
from tarpan.shared.tree_plot import TreePlotParams
# Sample from two models
model1 = CmdStanModel(stan_file="your_model1.stan")
fit1 = model1.sample(data=your_data)
model2 = CmdStanModel(stan_file="your_model2.stan")
fit2 = model2.sample(data=your_data)
# Supply legend labels (optional)
tree_params = TreePlotParams()
tree_params.labels = ["Model 1", "Model 2", "Exact"]
data = [{"mu": 2.2, "tau": 1.3}] # Add extra markers (optional)
save_tree_plot([fit1, fit2], extra_values=data, param_names=['mu', 'tau'],
tree_params=tree_params)
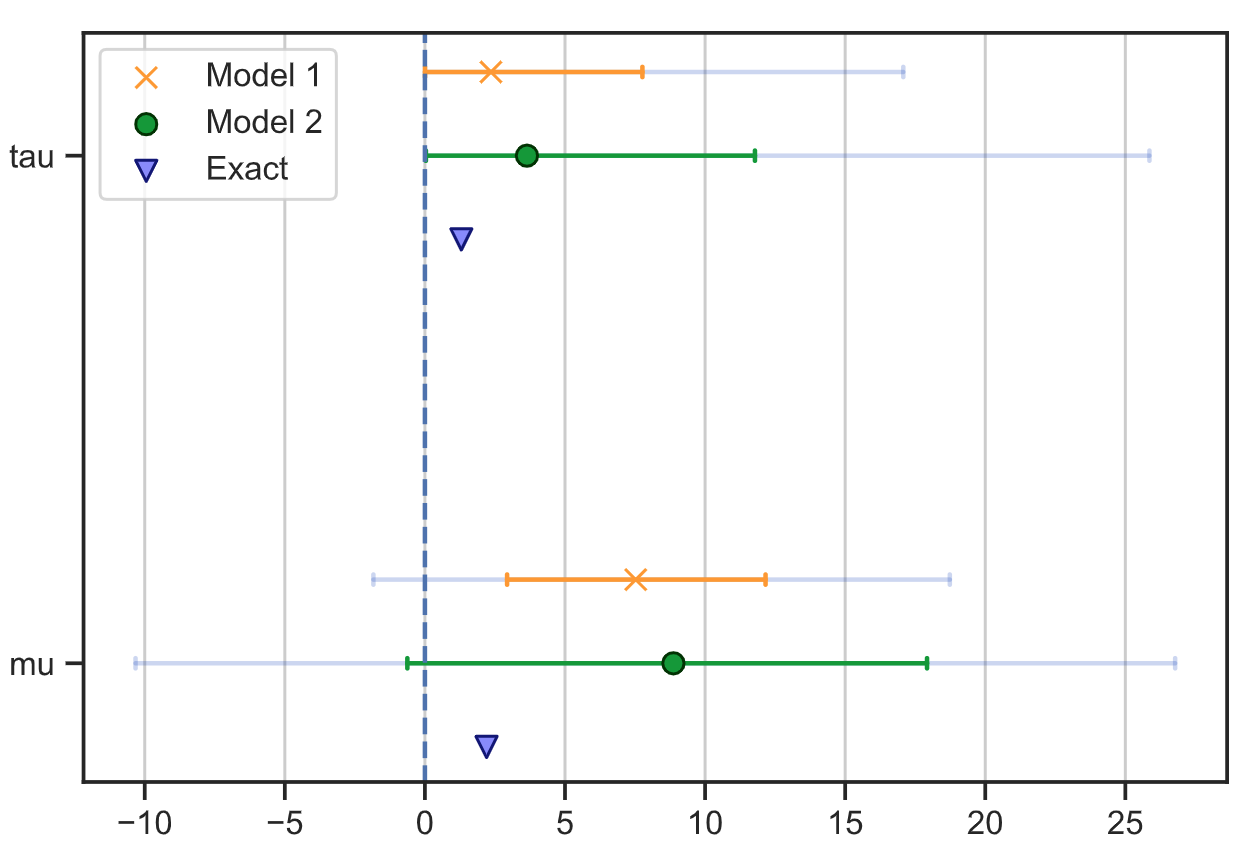
Comparing parameters of multiple models
Use save_compare_parameters function to compare parameters between different models:
| mu | tau | |
|---|---|---|
| Model 1 | 7.53 (+4.63, -4.59) | 2.36 (+5.41, -2.35) |
| Model 2 | 8.87 (+9.05, -9.50) | 3.64 (+8.14, -3.61) |
This table is a numerical version of the plot created by save_tree_plot. The values here are modes of the distributions and uncertainties are distances to 68% HPD intervals.
from tarpan.cmdstanpy.compare_parameters import save_compare_parameters
extra = [{"mu": 2.2, "theta": 1.3}] # Add extra values (optional)
save_compare_parameters([fit1, fit2], labels=['Model 1', 'Model 2', 'Extra'],
extra_values=extra,
param_names=["mu", "theta"])
Use save_compare_parameters without cmdstanpy
Here is how to compare parameters using Pandas data frames df1 and df2:
from tarpan.shared.compare_parameters import save_compare_parameters
extra = [{"mu": 2.2, "theta": 1.3}] # Add extra values (optional)
save_compare_parameters([df1, df2], labels=['Model 1', 'Model 2', 'Extra'],
extra_values=extra,
param_names=["mu", "theta"])
Trace plot: save_traceplot
The plot shows the values of parameters samples. Different colors correspond to samples form different chains. Ideally, the lines of different colors on the left plots are well mixed, and the right plot is fairly uniform.
from tarpan.cmdstanpy.traceplot import save_traceplot
model = CmdStanModel(stan_file="your_model.stan")
fit = model.sample(data=your_data)
save_traceplot(fit, param_names=['mu', 'tau', 'eta.1'])
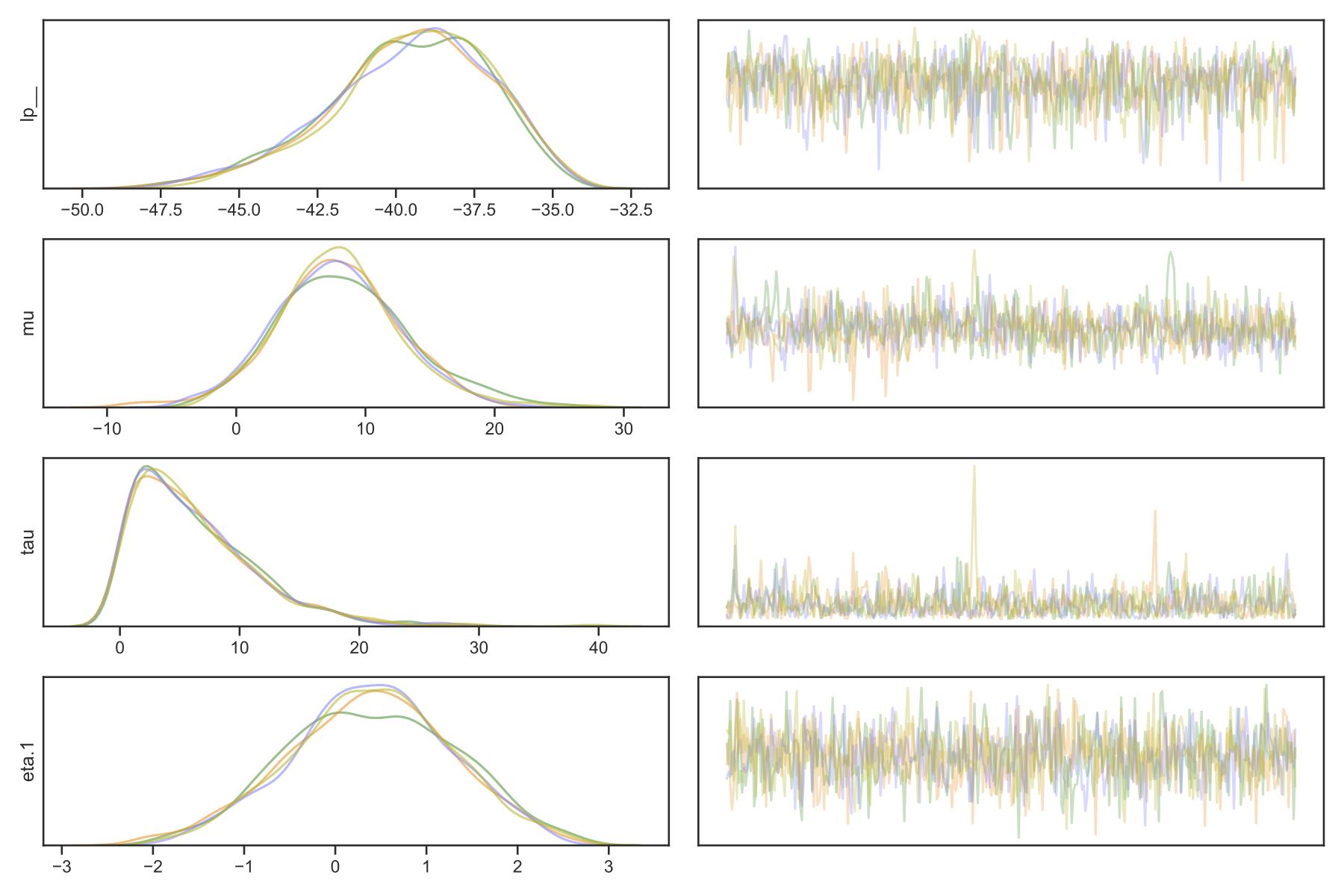
Pair plot: save_pair_plot
The plot helps to see correlations between parameters and spot funnel shaped distributions that can result in sampling problems.
from tarpan.cmdstanpy.pair_plot import save_pair_plot
model = CmdStanModel(stan_file="your_model.stan")
fit = model.sample(data=your_data)
save_pair_plot(fit, param_names=['mu', 'tau', 'eta.1'])
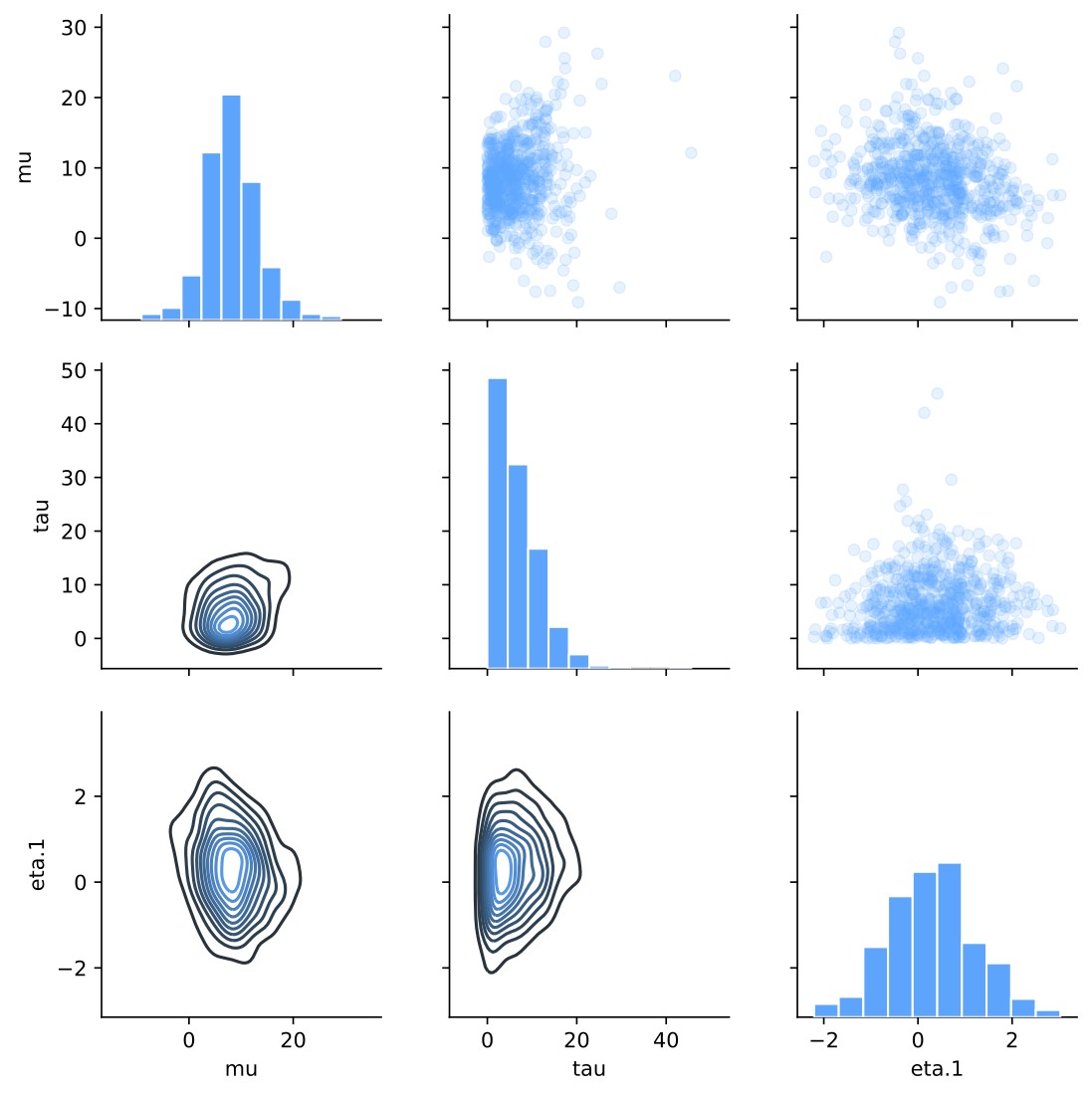
Pair plot without cmdstanpy
Here is how to make a pair plot of values from Pandas' data frame columns:
from tarpan.shared.pair_plot import save_pair_plot
save_pair_plot(df, param_names=['mu', 'sigma'])
Histogram: save_histogram
Show histograms of parameter distributions.
from tarpan.cmdstanpy.histogram import save_histogram
model = CmdStanModel(stan_file="your_model.stan")
fit = model.sample(data=your_data)
save_histogram(fit, param_names=['mu', 'tau', 'eta.1', 'theta.1'])
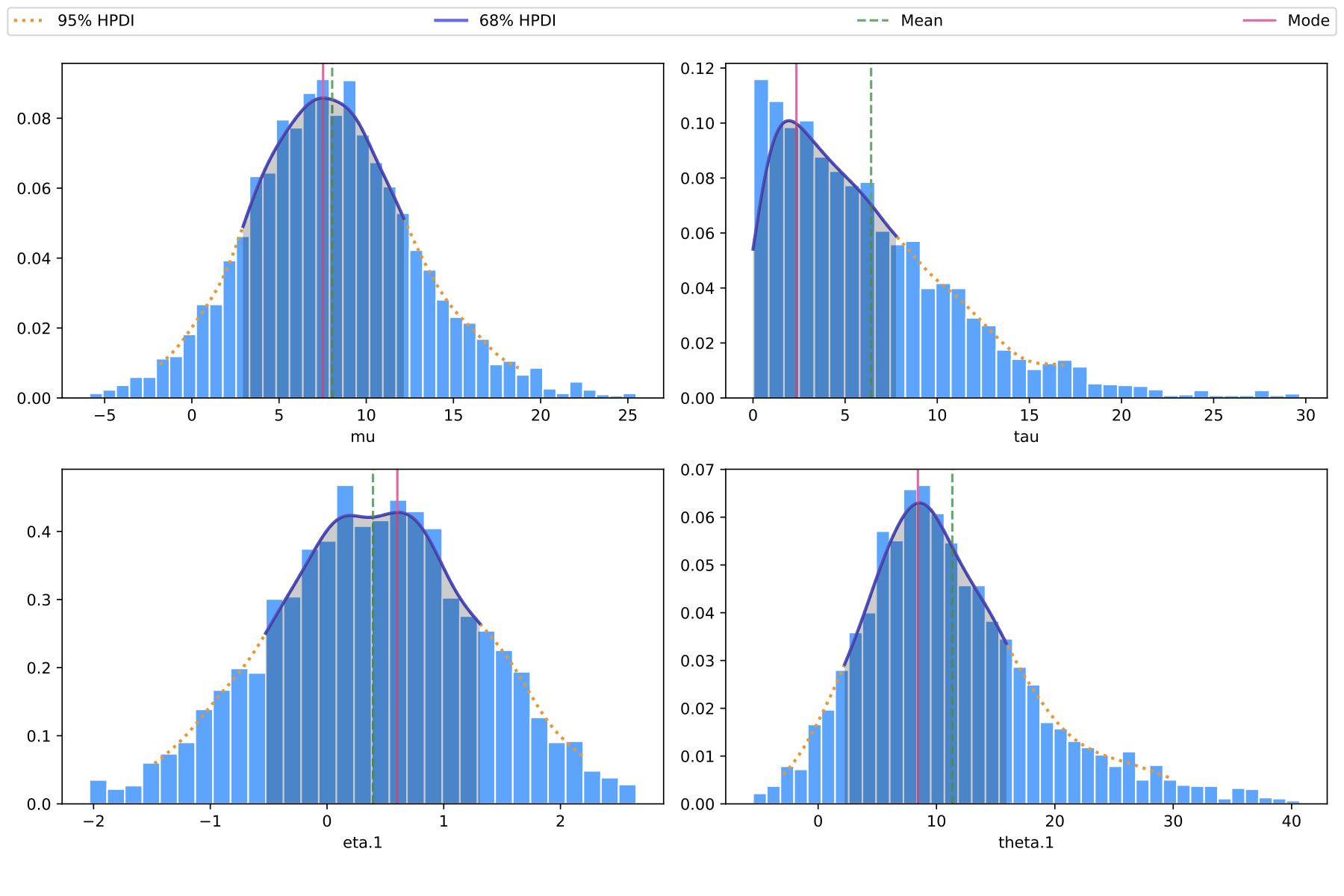
Histogram without cmdstanpy
Here is how to make histograms of values from Pandas' data frame columns:
from tarpan.shared.histogram import save_histogram
save_histogram(df, param_names=['mu', 'sigma'])
Comparing models
Run save_compare to compare multiple models using WAIC and PSIS methods in order to see which models are more compatible with the data.
from tarpan.cmdstanpy.compare import save_compare
model1 = CmdStanModel(stan_file="your_model1.stan")
fit1 = model1.sample(data=your_data)
model2 = CmdStanModel(stan_file="your_model2.stan")
fit2 = model2.sample(data=your_data)
models = {
"Model": fit1,
"Another model": fit2
}
save_compare(models=models, lpd_column_name="lpd_pointwise")
Here the lpd_column_name parameter takes the name of the array variable from the generated quantities block that contains log probability densities of all data points.
WAIC and PSIS plots
The save_compare function will create the plots showing WAIC and PSIS values (red round markers). The lower WAIC and PSIS values mean the model is more compatible with the data. The blue triangle marker show the difference between the model and the best model. The error bars correspond to standard errors.
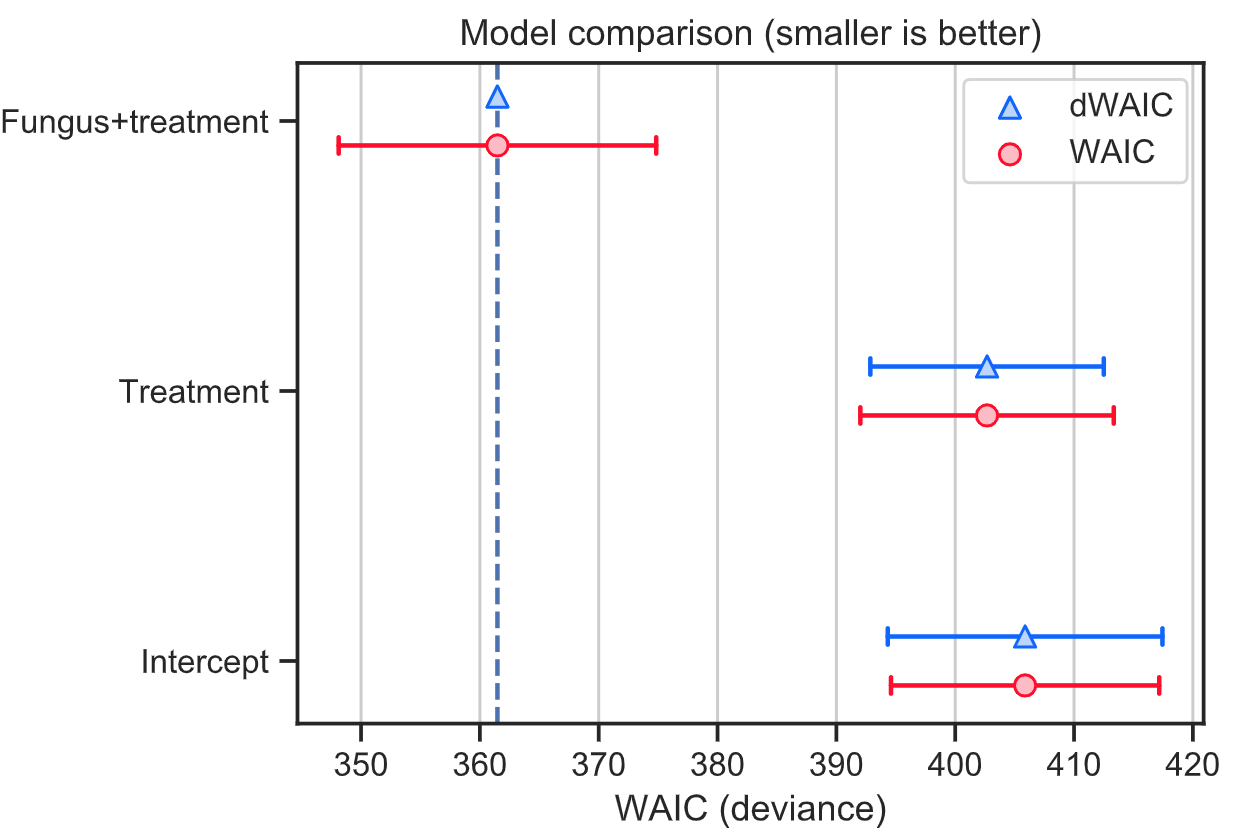
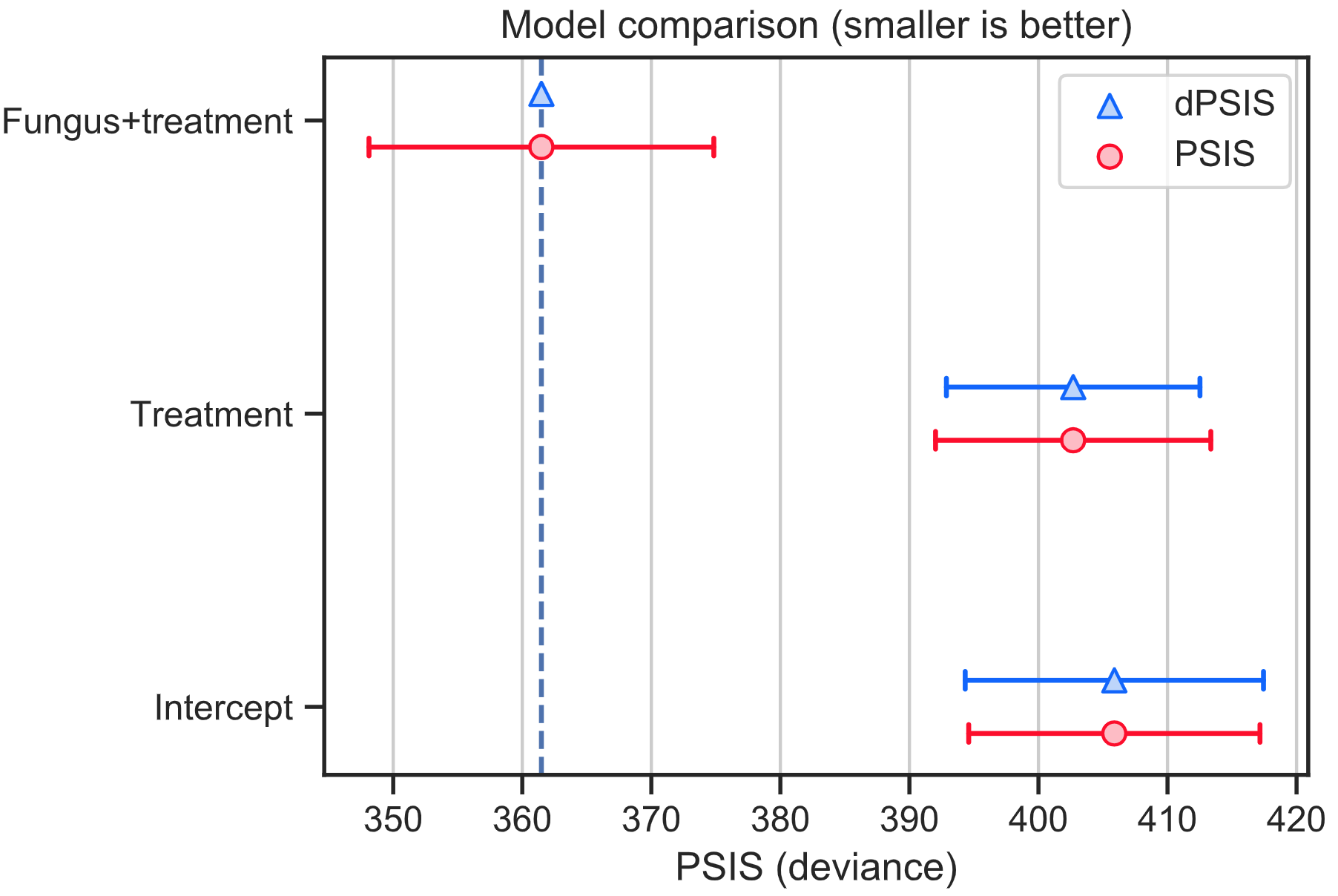
Numerical summaries for WAIC and PSIS
The save_compare function will also create text and csv files which are text versions of the above plots.
Summary columns are:
-
PSIS/WAIC, SE: PSIS and WAIC values and their standard errors.
-
dPSIS/dWAIC, dSE: The difference of PSIS and WAIC from the best model (i.e. model with lowest WAIC/PSIS) and the standard error of this difference.
-
pWAIC/pPSIS: the penalty (aka effective number of parameters). The purpose of number is to combat overfitting. Penalties are already included in the WAIC/PSIS numbers, so models with too many parameters will have larger penalties, and therefore, larger WAIC/PSIS values.
-
Weight: Very approximate measure of the relevance of the model, with higher numbers correspond to models that are more compatible with the data. Since this number is approximate and does not have uncertainty, it's better to use dWAIC/dPSIS with dSE to compare models.
-
MaxK: The maximum value of Pareto K parameter from the observations. If this value is above 0.5, and especially above 0.7, the PSIS/WAIC model comparisons might not be reliable.
WAIC summary
| WAIC | SE | dWAIC | dSE | pWAIC | Weight | |
|---|---|---|---|---|---|---|
| Fungus+treatment | 361.48 | 13.36 | 3.49 | 1.00 | ||
| Treatment | 402.68 | 10.66 | 41.20 | 9.82 | 2.53 | 0.00 |
| Intercept | 405.88 | 11.29 | 44.40 | 11.56 | 1.53 | 0.00 |
PSIS summary
| PSIS | SE | dPSIS | dSE | pPSIS | MaxK | Weight | |
|---|---|---|---|---|---|---|---|
| Fungus+treatment | 361.48 | 13.36 | 3.49 | 0.25 | 1.00 | ||
| Treatment | 402.69 | 10.67 | 41.21 | 9.82 | 2.54 | 0.33 | 0.00 |
| Intercept | 405.88 | 11.29 | 44.40 | 11.56 | 1.53 | 0.28 | 0.00 |
Plots of Pareto K values
The save_compare function creates plots of Pareto K values for data points. Points with Pareto K values higher than 0.7 are highlighted in red, with their indices shown below the markers. The red points are the ones that have large influence on the model. Having points above 0.7 could mean that WAIC and PSIS are failing and their results should be used with caution and large neon coloured disclaimers.
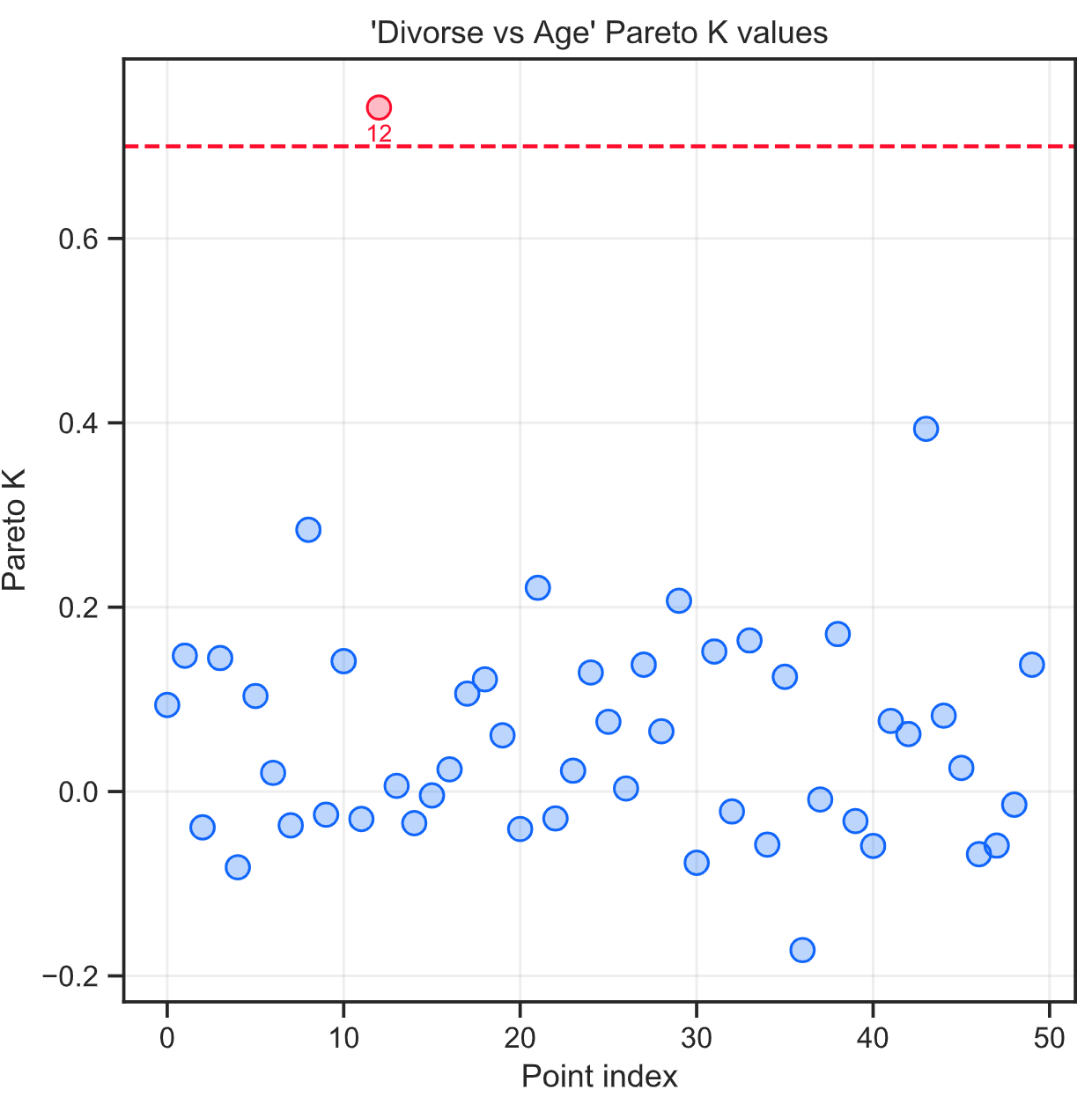
Saving cmdstan samples to disk
It saves a lot of time to sample the model and save the results to disk, so
they can be used on the next run instead of waiting for the sampling again.
This can be done with run function:
from tarpan.cmdstanpy.cache import run
# Your function that creates CmdStanModel, runs its `sample` method
# and returns the result.
#
# This function must take `output_dir` input parameter and pass it to `sample`.
#
# It may also have any other parameters you wish to pass from `run`.
def run_stan(output_dir, other_param):
model = CmdStanModel(stan_file="my_model.stan")
fit = model.sample(
data=data,
output_dir=output_dir # Pass to make CSVs in correct location
)
return fit # Return the fit
# Will run `run_stan` once, save model to disk and read it on next calls
fit = run(func=run_stan, other_param="some data")
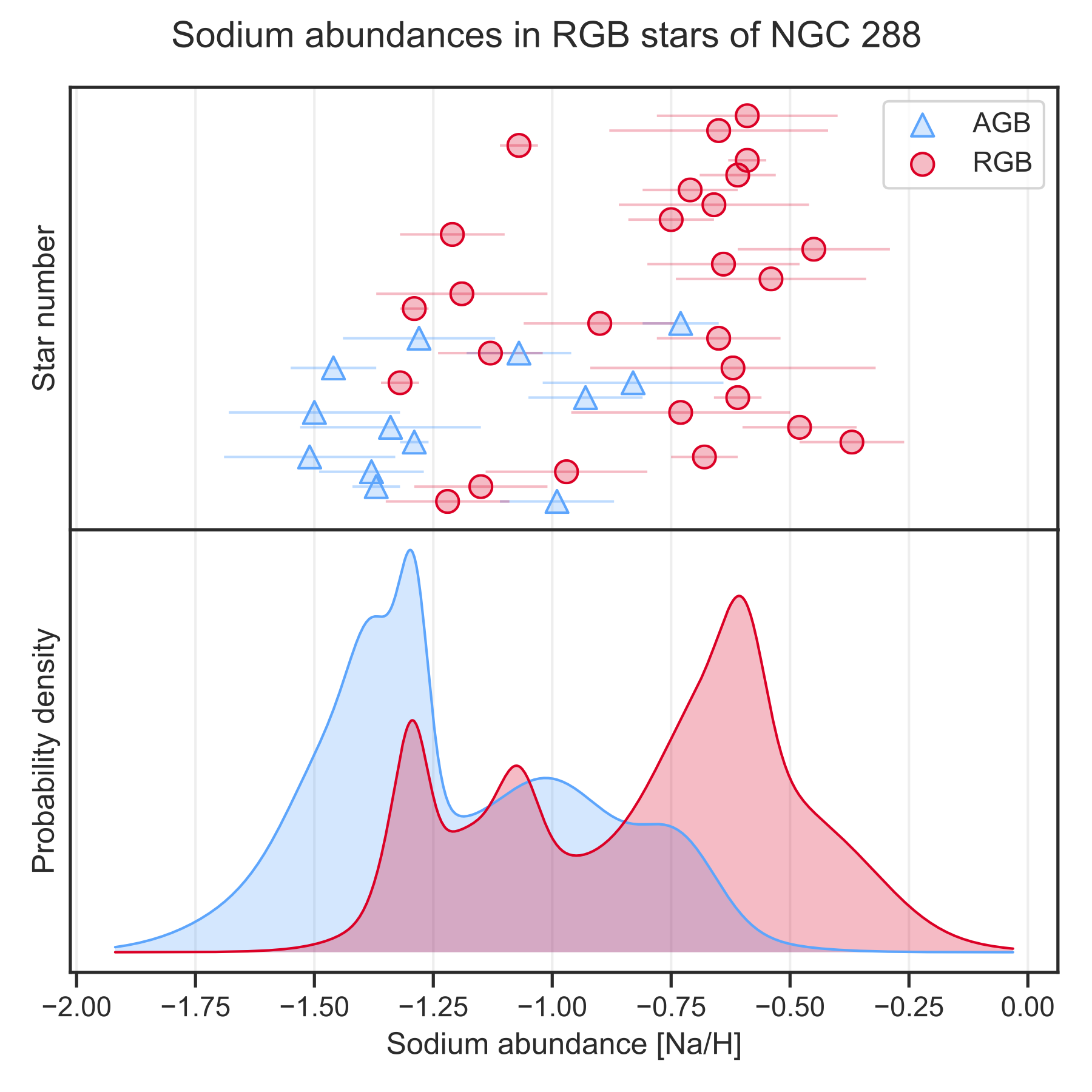
Scatter and KDE plot
The save_scatter_and_kde function saves a scatter and corresponding KDE (kernel density estimate) plot. The KDE plot takes into account uncertainties of individual values:
from tarpan.plot.kde import save_scatter_and_kde
values1 = [
-0.99, -1.37, -1.38, -1.51, -1.29, -1.34, -1.50, -0.93, -0.83,
-1.46, -1.07, -1.28, -0.73]
uncertainties1 = [
0.12, 0.05, 0.11, 0.18, 0.03, 0.19, 0.18, 0.12, 0.19,
0.09, 0.11, 0.16, 0.08]
values2 = [
-1.22, -1.15, -0.97, -0.68, -0.37, -0.48, -0.73, -0.61, -1.32,
-0.62, -1.13, -0.65, -0.90, -1.29, -1.19, -0.54, -0.64, -0.45,
-1.21, -0.75, -0.66, -0.71, -0.61, -0.59, -1.07, -0.65, -0.59]
uncertainties2 = [
0.13, 0.14, 0.17, 0.07, 0.11, 0.12, 0.23, 0.05, 0.04,
0.30, 0.11, 0.13, 0.16, 0.03, 0.18, 0.20, 0.16, 0.16,
0.11, 0.09, 0.20, 0.10, 0.08, 0.04, 0.04, 0.23, 0.19]
save_scatter_and_kde(values=[values1, values2],
uncertainties=[uncertainties1, uncertainties2],
title="Sodium abundances in RGB stars of NGC 288",
xlabel="Sodium abundance [Na/H]",
ylabel=["Star number", "Probability density"],
legend_labels=["AGB", "RGB"])
gaussian_kde function
The function returns the values for a KDE plot, taking into account uncertainties of individual values:
from tarpan.plot.kde import gaussian_kde
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0, 1, 100)
y = gaussian_kde(x, values, uncert)
plt.fill_between(x, y)
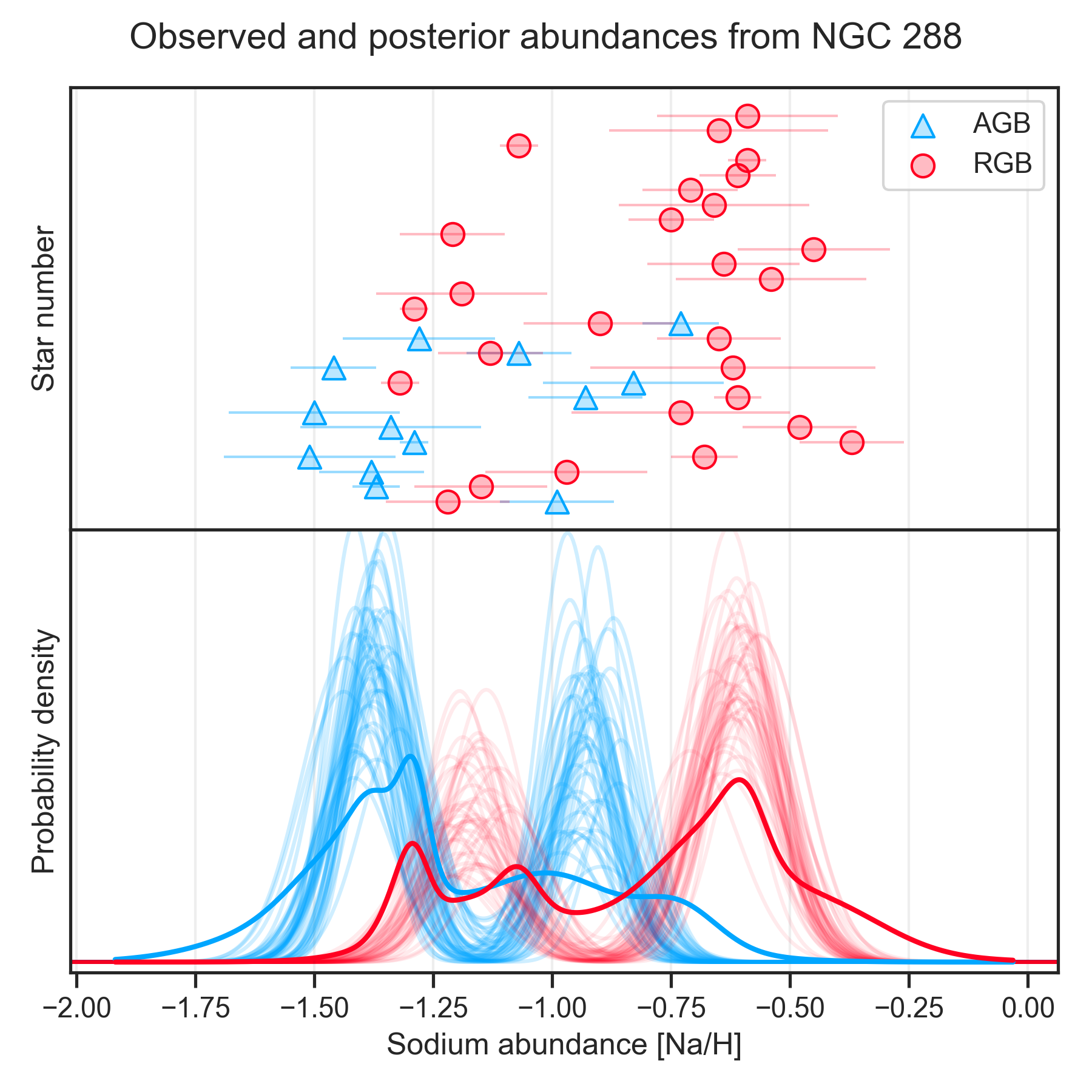
Make posterior-scatter-kde plot
The save_posterior_scatter_and_kde function makes a scatter-KDE plots
of the data, same as save_scatter_and_kde. In addition, it plots
the posterior distributions.
from tarpan.plot.posterior import save_posterior_scatter_and_kde
# Plot one sample from posterior distribution
def model_pdf(x, row):
mu = row['mu.1']
sigma = row['sigma']
return stats.norm.pdf(x, mu, sigma)
fig, axes = save_posterior_scatter_and_kde(
fits=[fit1, fit2], # Two models returned by model.sample function
pdf=model_pdf, # Function that plot posterior distribution
values=[data1["y"], data2["y"]],
uncertainties=[data1["uncertainties"], data2["uncertainties"]],
title="Sodium abundances in RGB stars of NGC 288",
xlabel="Sodium abundance [Na/H]",
ylabel=["Star number", "Probability density"],
legend_labels=["AGB", "RGB"])
Common questions
Run unit tests
pytest
The unlicense
This work is in public domain.
🐴🐴🐴
This work is dedicated to Tarpan, an extinct subspecies of wild horse.
Special thanks to Richard McElreath, who wrote Statistical Rethinking textbook, as well as Stan and arviz people.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file tarpan-0.3.11.tar.gz.
File metadata
- Download URL: tarpan-0.3.11.tar.gz
- Upload date:
- Size: 51.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.7.0 requests/2.25.1 setuptools/52.0.0.post20210125 requests-toolbelt/0.9.1 tqdm/4.56.1 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5148d493324a9eb066248586b4a194c15e5bd3164421d9fc2b44295874fcb666
|
|
| MD5 |
be6acfc0790238ce037e5776acefbcdb
|
|
| BLAKE2b-256 |
bbe0973c67f3971770aa61718a2080d19a82bfef60fcba1dabf59bdb16a3a124
|
File details
Details for the file tarpan-0.3.11-py3-none-any.whl.
File metadata
- Download URL: tarpan-0.3.11-py3-none-any.whl
- Upload date:
- Size: 63.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.7.0 requests/2.25.1 setuptools/52.0.0.post20210125 requests-toolbelt/0.9.1 tqdm/4.56.1 CPython/3.8.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a916c1051edf8968245ceef5835b35a262c0bfad5827b6a2f6121d321c9c7972
|
|
| MD5 |
5fa807afa06454ea2117024e4688ea2d
|
|
| BLAKE2b-256 |
640d983776722f5fe19c8f6eccbb7e7727107f06e3d531009503cb3315fbe85d
|











