Simulate a continuous-time threshold model on static networks.
Project description
Simulate a continuous-time threshold model on static networks using Gillespie’s stochastic simulation algorithm (SSA). The networks can be directed and/or weighted.
In contrast to the original discrete-time model, nodes whose aggregated inputs exceed their respective thresholds will not flip after the “next time step” because there are no time steps. Instead, a node whose threshold has been exceeded will enter an alert state from which it will enter the activated state with rate $gamma = 1$.
Install
pip install thresholdmodel
Example
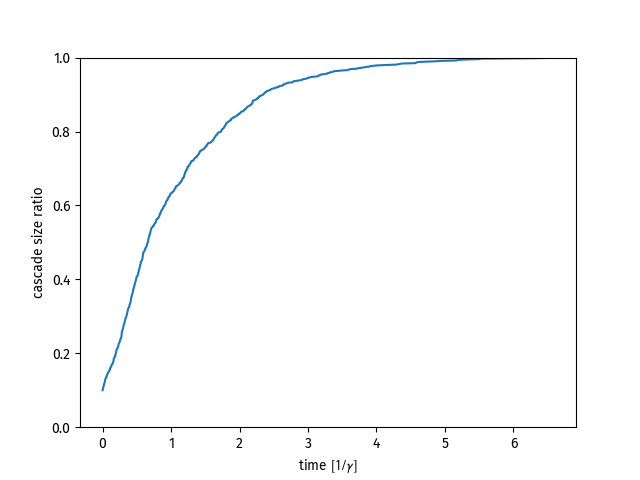
Simulate on an ER random graph.
import numpy as np
import networkx as nx
import matplotlib.pyplot as plt
from thresholdmodel import ThreshModel
N = 1000
k = 10
thresholds = 0.1
initially_activated = np.arange(100)
G = nx.fast_gnp_random_graph(N, k/(N-1.0))
Thresh = ThreshModel(G,initially_activated,thresholds)
t, cascade_size = Thresh.simulate()
plt.plot(t,cascade_size)
plt.show()API
Simulate
Given a networkx-Graph object G (can be a networkx.DiGraph, too), and values for initially_activated and thresholds, simulate like this
Thresh = ThreshModel(G,initially_activated,thresholds)
t, a = Thresh.simulate()t is a numpy.ndarray containing the times at which node activations happened. a is a numpy.ndarray containing the relative cascade size at the corresponding time in t. Note that the whole process is modeled as a Poisson process such that the time t will be given in units of the node activation rate gamma = 1.0. If you want to simulate for another node activation rate, simply rescale time as t /= gamma.
When the simulation is started with the save_activated_nodes=True flag, a list of activated nodes per time leap is saved in ThreshModel.activated_nodes.
t, a = Thresh.simulate(save_activated_nodes=True)
print(Thresh.activated_nodes)You can repeat a simulation with the same initial conditions by simply calling Thresh.simulate() again, all the necessary things will be reset automatically.
Set initially activated nodes
Set nodes 3, 5, and 8 to be activated initially.
initially_activated = [3, 5, 8] # this could also be a numpy arrayChoose 20% of all nodes randomly to be activated initially. When the simulation is restarted, the same nodes will be chosen as initial conditions.
initially_activated = 0.2Choose 35 randomly selected nodes to be activated initially. When the simulation is restarted, the same nodes will be chosen as initial conditions.
initially_activated = 35Set thresholds
Activation thresholds can be set for all nodes
thresholds = np.random.rand(G.number_of_nodes())Note that thresholds need to lie in the domain [0,1].
You can also set a universal threshold
thresholds = 0.1Here, 10% of a node’s neighbors need to be activated in order for the node to become active, too.
Directed networks
A node will become active if the sufficient number of nodes pointing towards the node are active. This means that a node’s in-degree will be the important measure to determine wether this particular node will become active.
Weighted networks
If you want to simulate on a weighted network, provide the weight keyword
Thresh = ThreshModel(G,initially_activated,thresholds,weight='weight')Similar to the networkx-documentation: weight (string, optional (default=None)) - The attribute name to obtain the edge weights. E.g.: G.edges[0,1]['weight'].
A focal node will become active when the cumulative edge weights of all activated nodes pointing towards the focal node will reach > threshold*in_degree.
Docstring
This is the model’s docstring.
>>> help(ThreshModel) Help on class ThreshModel in module thresholdmodel.model: class ThreshModel(builtins.object) | ThreshModel(G, initially_activated, thresholds, weight=None) | | A simple simulation class that runs | a threshold-model activation process | on a static network (potentially weighted and directed) | in continuous time using Gillespie's | stochastic simulation algorithm. | | The temporal dimension is fixed by assuming | that every node whose activation threshold | has been exceeded by neighboring inputs | is activated with constant and uniform | rate :math:`\gamma = 1`. | | Parameters | ========== | G : networkx.Graph, networkx.DiGraph | The network on which to simulate. | Nodes must be integers in the range | of ``[0, N-1]``. | initially_activated: float, int, or list of ints | Can be either of three things: | | 1. float of value ``0 < initially_activated < 1``. | In this case, ``initially_activated`` is | interpreted to represent a fraction of nodes | that will be randomly selected from the | set of nodes and set to be activated. | 2. int of value ``1 <= initially_activated < N-1``. | In this case, ``initially_activated`` nodes | will be randomly sampled from the node set | and set to be activated. | 3. list of ints. In this case, ``initially_activated`` | is interpreted to contain indices of nodes | that will be activated initially. | thresholds: float or iterable of floats | Can be either of two things: | | 1. float of value ``0 < thresholds <= 1``. | In this case, every node will have the same | activation threshold. | 2. iterable of values ``0 < thresholds <=1``. | In this case, the function expectes a list, | tuple, or array with length equal to the | number of nodes. Each entry `m` of this list | will be interpreted to be node `m`'s activation | threshold. | weight: str, default = None | A string that represents the weight keyword of a link. | If `None`, the network is assumed to be unweighted. | | Example | ======= | | >>> G = nx.fast_gnp_random_graph(1000,20/(1000-1)) | >>> model = TreshModel(G, 100, 0.1) | >>> t, cascade_size = model.simulate() | | Attributes | ========== | G : nx.Graph or nx.DiGraph | The network on which to simulate. | Nodes must be integers in the range | of ``[0, N-1]``. | N : int | The number of nodes in the network | weight: str | A string that represents the weight keyword of a link. | If `None`, the network is assumed to be unweighted. | in_deg : numpy.ndarray | Contains the in-degree of every node. | thresholds: numpy.ndarray | Each entry `m` of this array | represents node `m`'s activation | threshold. | initially_activated: numpy.ndarray | Each entry of this array contains | a node that will be activated initially. | time: numpy.ndarray | Contains every time point at which a node was | activates (after ``simulation()`` was called). | The temporal dimension is given by assuming | that every node whose activation threshold | has been exceeded by activation inputs | is activated with constant and uniform | rate :math:`\gamma = 1`. | cascade_size: numpy.ndarray | The relative size of the activation cascade | at the corrsponding time value in ``time`` | (relative to the size of the node set). | Only available after ``simulation()`` was called. | activated_nodes: list | A list of lists. | Each entry contains a list of integers representing | the nodes that have been activated | at the corrsponding time value in ``time``. | Each list entry will contain only a single node | for every other time than the initial time. | Only available after ``simulation()`` was called. | | Methods defined here: | | __init__(self, G, initially_activated, thresholds, weight=None) | Initialize self. See help(type(self)) for accurate signature. | | reset(self) | Reset the simulation. | | set_initially_activated(self, initially_activated) | Set the process's initial activation state. | | Parameters | ========== | initially_activated: float, int, or list of ints | Can be either of three things: | | 1. float of value ``0 < initially_activated < 1``. | In this case, ``initially_activated`` is | interpreted to represent a fraction of nodes | that will be randomly selected from the | set of nodes and set to be activated. | 2. int of value ``1 <= initially_activated < N-1``. | In this case, ``initially_activated`` nodes | will be randomly sampled from the node set | and set to be activated. | 3. list of ints. In this case, ``initially_activated`` | is interpreted to contain indices of nodes | that will be activated initially. | | set_thresholds(self, thresholds) | Set node activation thresholds. | | Parameters | ========== | thresholds: float or iterable of floats | Can be either of two things: | | 1. float of value ``0 < thresholds <= 1``. | In this case, every node will have the same | activation threshold. | 2. iterable of values ``0 < thresholds <=1``. | In this case, the function expectes a list, | tuple, or array with length equal to the | number of nodes. Each entry `m` of this list | will be interpreted to be node `m`'s activation | threshold. | | simulate(self, save_activated_nodes=False) | Simulate until all nodes that can be activated | have been activated. | | Parameters | ========== | save_activated_nodes: bool, default = False | If ``True``, write a list of activated nodes | to the class attribute ``activated_nodes`` | every time an activation event happens. | Such a list will contain only a single node | for every other time than the initial time. | | Returns | ======= | time : numpy.ndarray | Time points at which nodes were activated. | cascade_size : numpy.ndarray | The relative size of the activation cascade | at the corrsponding time value in ``time`` | (relative to the size of the node set).
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file thresholdmodel-0.0.3.tar.gz.
File metadata
- Download URL: thresholdmodel-0.0.3.tar.gz
- Upload date:
- Size: 32.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.6.1 requests/2.25.0 setuptools/50.3.2 requests-toolbelt/0.9.1 tqdm/4.54.1 CPython/3.8.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2d65e6966f0c528bccea80c2e92a6e524f64b0321026606653b03def42067f60
|
|
| MD5 |
af330af066cb674a659529384ec08fbd
|
|
| BLAKE2b-256 |
c2c1f5e6cb2516e66ebb851e2d1e9f4a826ce2fc5d3103f668c6d95595526f78
|