Open source framework for productionizing healthcare AI
Project description
Open-Source SDK for Healthcare AI Integration
HealthChain is an open-source Python SDK for connecting AI models to live clinical systems. The models aren't the bottleneck — the integration is. Get built-in FHIR support, real-time EHR connectivity, and production-ready deployment tooling without building it from scratch.
Installation
pip install healthchain
Quick Start
# Scaffold a FHIR Gateway project
healthchain new my-app -t fhir-gateway
cd my-app
# Run locally
healthchain serve

Edit app.py to add your model, and healthchain.yaml to configure compliance, security, and deployment settings.
See the CLI reference for all commands.
Core Features
The quickest way for AI developers and researchers to ship healthcare AI — everything you need out of the box, built to scale with you.
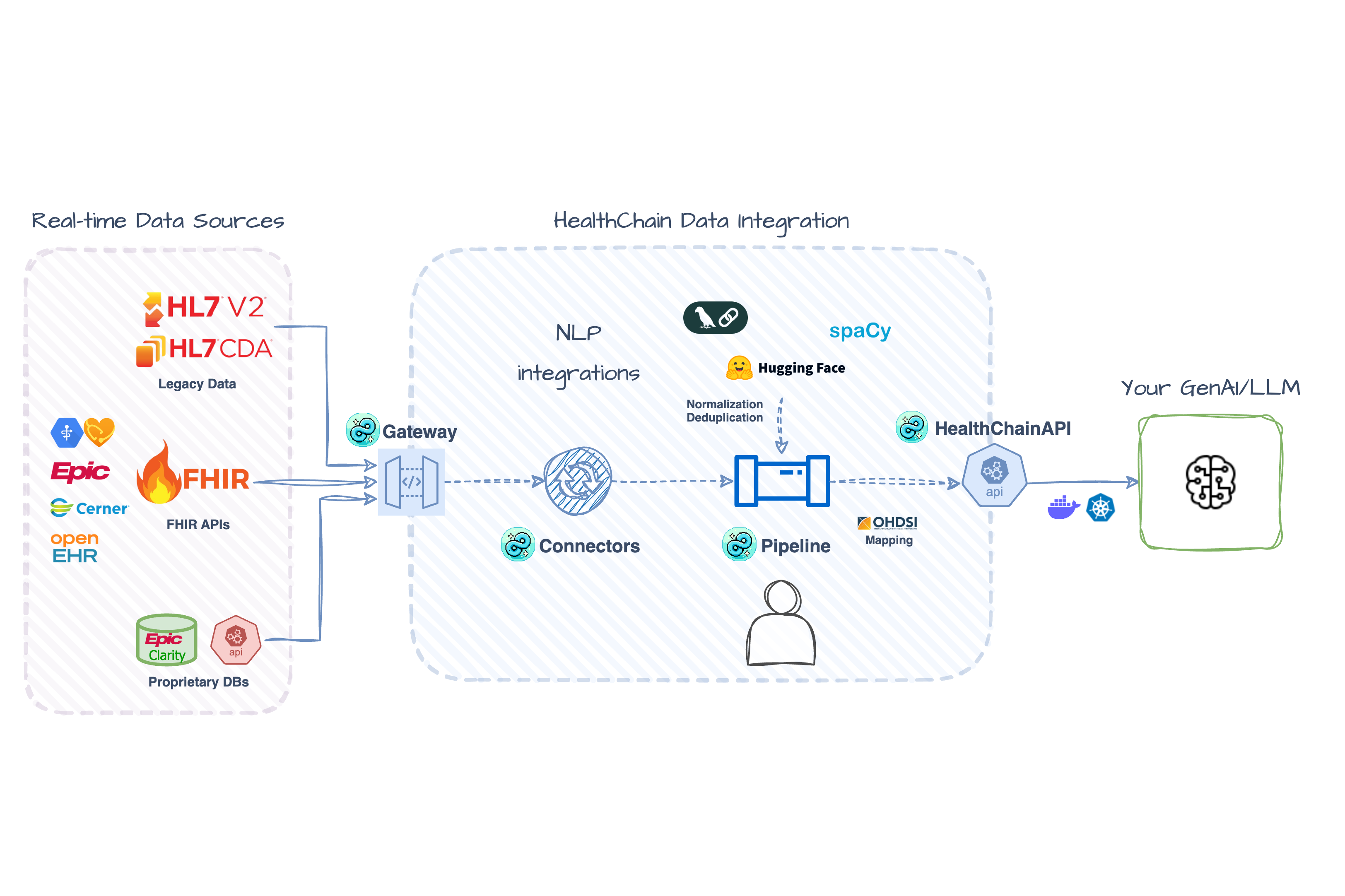
🔌 Multi-EHR Data Aggregation Aggregate patient data from multiple FHIR sources into unified records with built-in NLP processing and automatic deduplication Getting Started → |
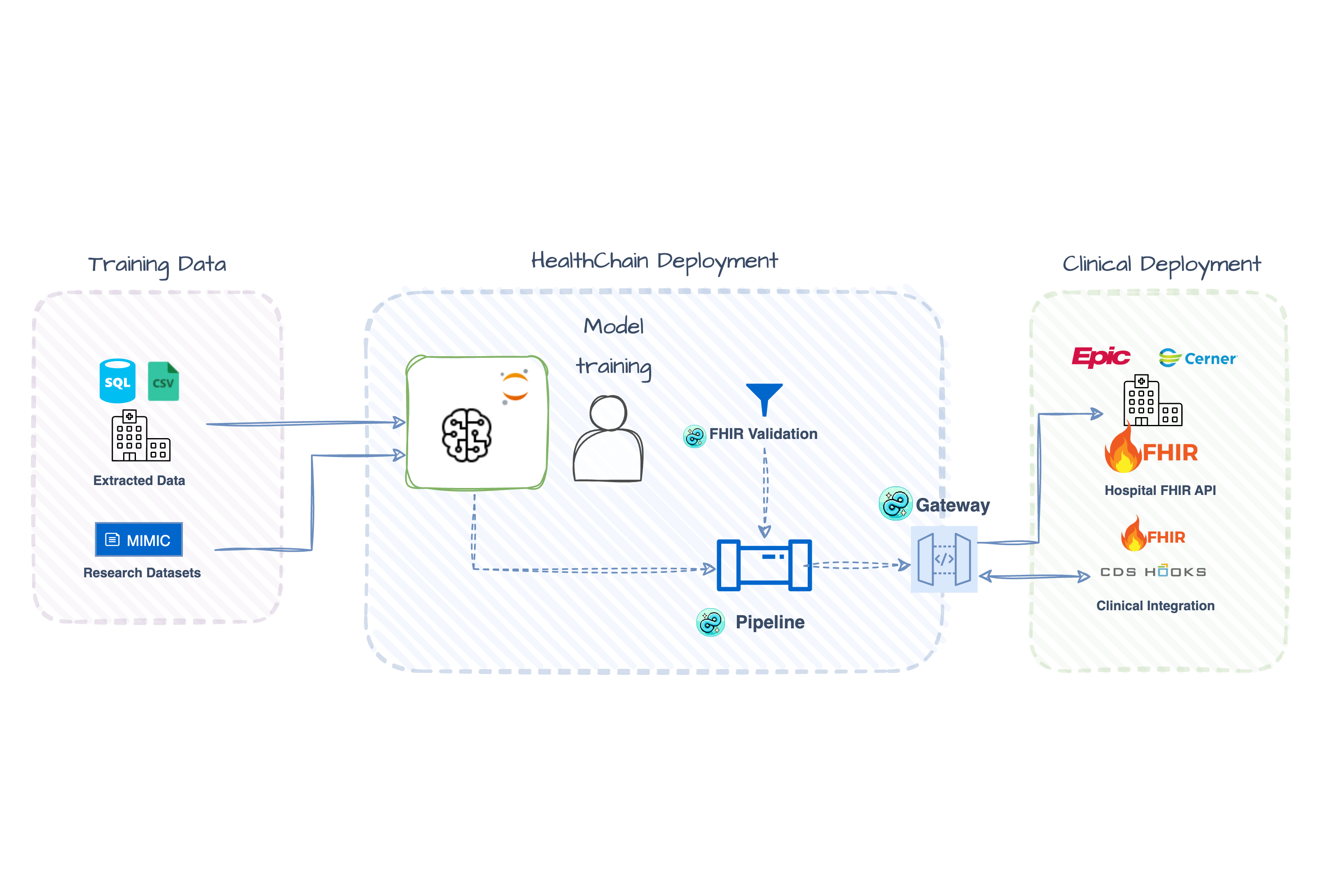
🚀 Deploy ML Models as Healthcare APIs Turn any trained model into a production-ready FHIR endpoint with OAuth2 authentication and type-safe healthcare data handling Getting Started → |
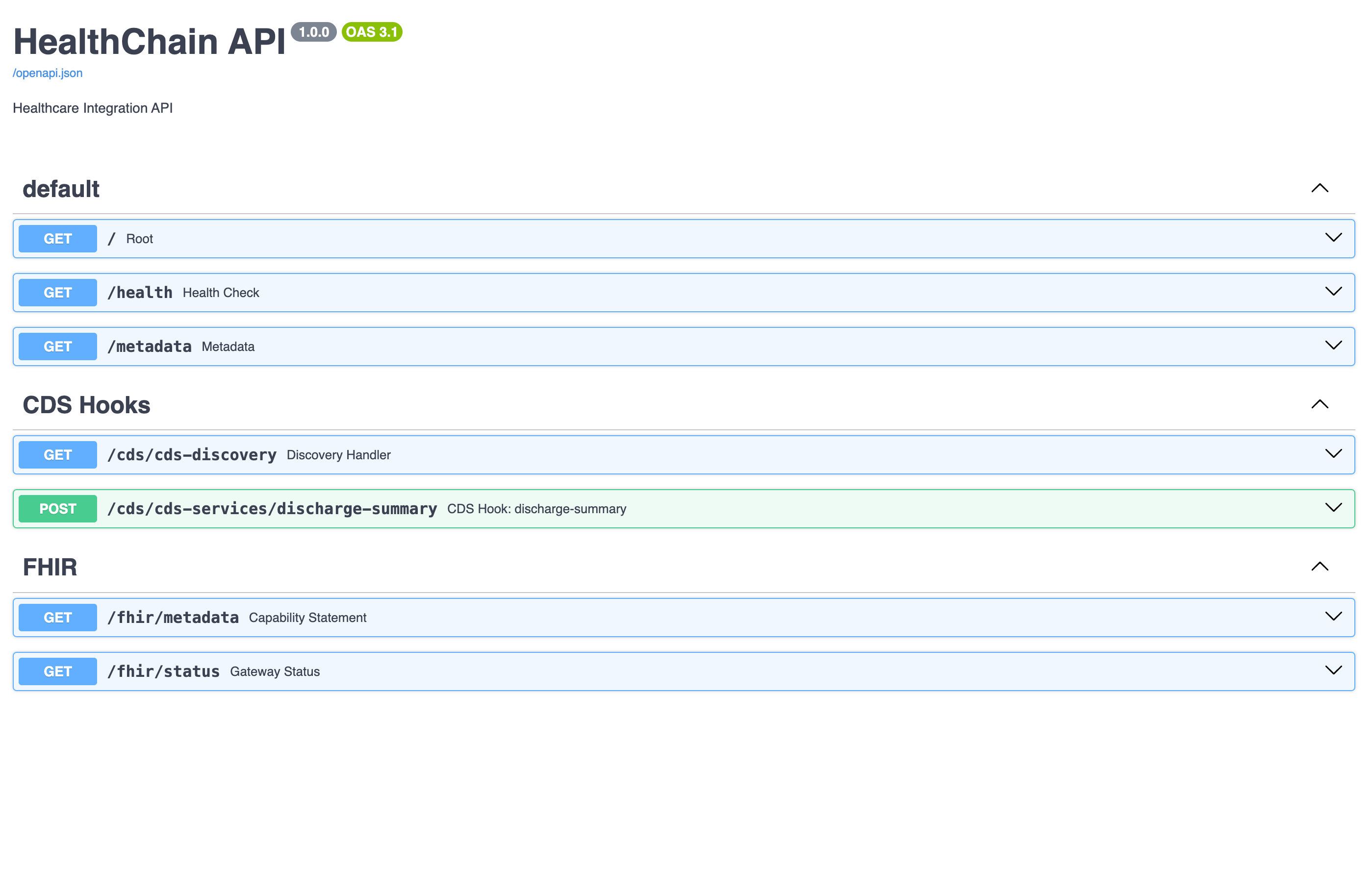
🔥 FHIR Development Utilities Type-safe FHIR resource creation, validation helpers, and sandbox environments — skip the boilerplate and work with healthcare data natively Getting Started → |
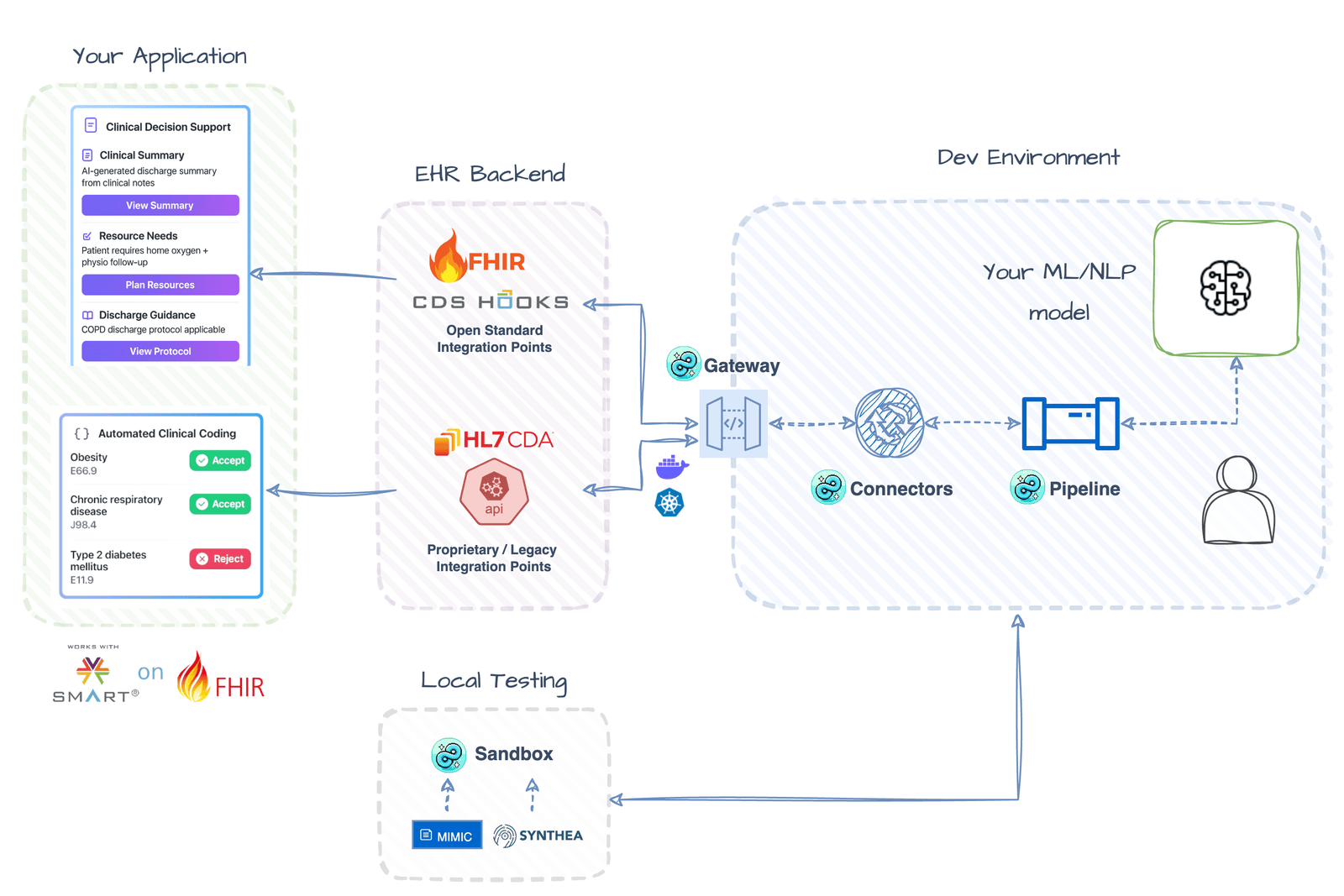
⚡️ Real-Time Clinical Workflow Integration Deploy AI models as CDS services that integrate directly into EHR workflows — alerts, recommendations, and automated coding at the point of care Getting Started → |
Why HealthChain?
Every serious healthcare AI project builds the same integration infrastructure from scratch. Whether you're deploying a logistic regression or a 70B parameter model, the wall between a trained model and a live clinical system is the same: real FHIR APIs, multi-site deployments, auditable governance. No off-the-shelf solution exists, and engineers who understand both AI and healthcare protocols are scarce and hard to retain.
HealthChain handles that complexity so you can focus on what actually matters: the model and the patient.
- Optimized for real-time - Connect to live FHIR APIs and integration points instead of stale data exports
- Automatic validation - Type-safe FHIR models prevent broken healthcare data
- Native LLM + ML support - Wire up any model, from LLMs to scikit-learn, and output results as FHIR
- Works with your existing stack - Integrates with FastAPI, LangChain, HuggingFace, and spaCy
- Production-ready foundations - Dockerized deployment, configurable security, and an architecture built for NHS and HIPAA environments
🏆 Recognition & Community
Featured & Presented:
- Featured in TLDR AI Newsletter (900K+ developers)
- Presented at NHS Python Open Source Conference (watch talk)
- Built from NHS AI deployment experience – read the origin story
🤝 Partnerships & Production Use
Exploring HealthChain for your product or organization? Get in touch to discuss integrations, pilots, or collaborations, or join our Discord to connect with the community.
Usage Examples
Creating a Gateway [Docs]
from healthchain.gateway import HealthChainAPI, FHIRGateway
from healthchain.fhir.r4b import Patient
# Create healthcare application
app = HealthChainAPI(title="Multi-EHR Patient Data")
# Connect to multiple FHIR sources
fhir = FHIRGateway()
fhir.add_source("epic", "fhir://fhir.epic.com/r4?client_id=epic_client_id")
fhir.add_source("cerner", "fhir://fhir.cerner.com/r4?client_id=cerner_client_id")
@fhir.aggregate(Patient)
def enrich_patient_data(id: str, source: str) -> Patient:
"""Get patient data from any connected EHR and add AI enhancements"""
bundle = fhir.search(
Patient,
{"_id": id},
source,
add_provenance=True,
provenance_tag="ai-enhanced",
)
return bundle
app.register_gateway(fhir)
# Available at: GET /fhir/transform/Patient/123?source=epic
# Available at: GET /fhir/transform/Patient/123?source=cerner
if __name__ == "__main__":
app.run(port=8888)
Building a Pipeline [Docs]
from healthchain.pipeline import Pipeline
from healthchain.pipeline.components.integrations import SpacyNLP
from healthchain.io import Document
# Create medical NLP pipeline
nlp_pipeline = Pipeline[Document]()
nlp_pipeline.add_node(SpacyNLP.from_model_id("en_core_web_sm"))
nlp = nlp_pipeline.build()
doc = Document("Patient presents with hypertension and diabetes.")
result = nlp(doc)
spacy_doc = result.nlp.get_spacy_doc()
print(f"Entities: {[(ent.text, ent.label_) for ent in spacy_doc.ents]}")
print(f"FHIR conditions: {result.fhir.problem_list}") # Auto-converted to FHIR Bundle
🛣️ What we're building towards
- 🔒 Production security and compliance — API authentication, audit logging, and configurable security for NHS/HIPAA deployments
- 📋 Governance as config — clinical safety, data access agreements, and compliance standards as a first-class deployment artifact in
healthchain.yaml - 🔌 Deeper EHR connectivity — more FHIR sources, live data patterns, and real-world integration examples from pilot deployments
- 📊 Observability — deployment telemetry and audit trails for healthcare systems
- 🤖 Agentic integrations — support for AI agent frameworks and agentic healthcare workflows
🤝 Contributing
HealthChain is built by and for the next generation of healthcare developers — researchers moving models from retrospective data into live systems, AI developers who don't want to spend months learning FHIR before they can ship anything. The best contributions come from people who have hit a real problem and have something specific to say about it.
Get started:
- Working with healthcare or research data? Contribute a cookbook — bring your use case, I'll personally support you through it
- Read CONTRIBUTING.md for guidelines
- Technical questions and ideas → GitHub Discussions
- Pilots and partnerships → email
🤗 Acknowledgements
This project builds on fhir.resources and CDS Hooks standards developed by HL7 and Boston Children's Hospital.
© 2024–2026 dotimplement ai. HealthChain is an open source project maintained by dotimplement ai.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file healthchain-0.14.4.tar.gz.
File metadata
- Download URL: healthchain-0.14.4.tar.gz
- Upload date:
- Size: 4.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
63b3623da8b6b947054fc1650a864a0ff2b057b7861772730a8aa8e67a10576c
|
|
| MD5 |
563bd3807d14779b3634fb6923834f40
|
|
| BLAKE2b-256 |
c2c8779ff2e7d3c969a3e235902ef6b0ad67372e23f156da08bb4b34b007dcaa
|
Provenance
The following attestation bundles were made for healthchain-0.14.4.tar.gz:
Publisher:
publish.yml on healthchainai/HealthChain
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
healthchain-0.14.4.tar.gz -
Subject digest:
63b3623da8b6b947054fc1650a864a0ff2b057b7861772730a8aa8e67a10576c - Sigstore transparency entry: 1451098655
- Sigstore integration time:
-
Permalink:
healthchainai/HealthChain@0f3710e6a93425be64b5fa9392d76bc5668baaad -
Branch / Tag:
refs/tags/0.14.4 - Owner: https://github.com/healthchainai
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@0f3710e6a93425be64b5fa9392d76bc5668baaad -
Trigger Event:
release
-
Statement type:
File details
Details for the file healthchain-0.14.4-py3-none-any.whl.
File metadata
- Download URL: healthchain-0.14.4-py3-none-any.whl
- Upload date:
- Size: 270.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d4fc902ded6527dc8078a820e2def0d4386418263f4df63451464f3dbe9c2a77
|
|
| MD5 |
cd450728d479f454f009d024f8cf8de0
|
|
| BLAKE2b-256 |
f178fa8fa3cfaf8626ed1480911a21eba076871a8dbafe1a41644d516d8c2084
|
Provenance
The following attestation bundles were made for healthchain-0.14.4-py3-none-any.whl:
Publisher:
publish.yml on healthchainai/HealthChain
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
healthchain-0.14.4-py3-none-any.whl -
Subject digest:
d4fc902ded6527dc8078a820e2def0d4386418263f4df63451464f3dbe9c2a77 - Sigstore transparency entry: 1451098732
- Sigstore integration time:
-
Permalink:
healthchainai/HealthChain@0f3710e6a93425be64b5fa9392d76bc5668baaad -
Branch / Tag:
refs/tags/0.14.4 - Owner: https://github.com/healthchainai
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@0f3710e6a93425be64b5fa9392d76bc5668baaad -
Trigger Event:
release
-
Statement type:






















