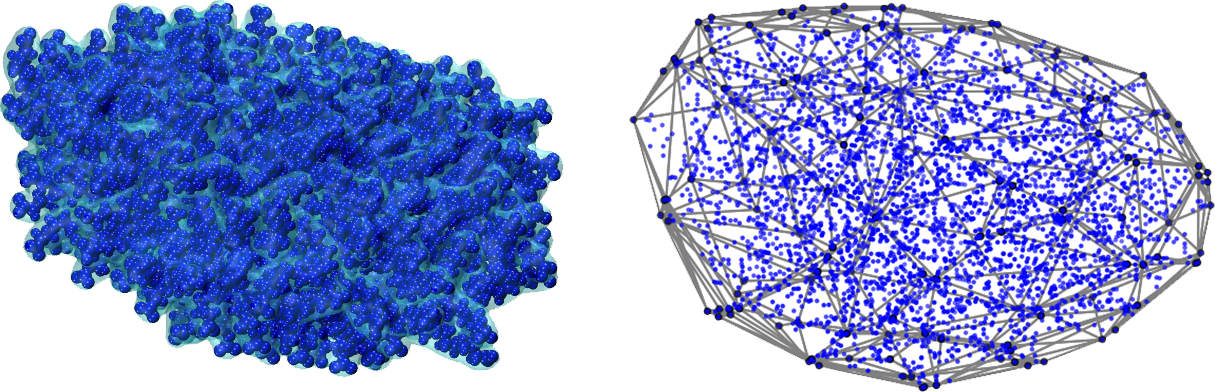
Python Utility for Characterizing Heterogeneous Interfaces and Kinetics creates a grid from an molecular dynamics structure file. The system in question can have an arbitrary structure. Volume and species densities can then calculated.
Project description
PUCHIK
Overview
Python Utility for Characterizing Heterogeneous Interfaces and Kinetics (PUCHIK), is a tool for analyzing molecular dynamics trajectories. It allows constructing an interface between two phases, enabling to calculate intrinsic density profiles, volumes, etc.
The interface construction works for spherical and rod-like nanoparticles equally well, making it a great tool to work with nanoparticles of almost every shape.
This package is built on top of MDAnalysis, SciPy, NumPy and PyGEL3D libraries.
This is an experimental branch that offers a functionality to use alpha shapes (with an optimal or user provided alpha value) instead of a convex hull. This is done with the help of AlphaShaper.exe which was compiled using the CGAL library. For now this only works on Windows. C++ source code is provided as well in case you would like to compile it for other platforms. The usage will be shown below.
Installation
You can install the PUCHIK package using pip:
pip install PUCHIK
Usage
The main class in this package is the "Interface" class. To set up a mesh, import it from PUCHIK:
from PUCHIK import Interface
You should provide it with a topology and optionally a trajectory files. PUCHIK uses MDAnalysis Readers to open a trajectory. You can find the supported formats here.
trj = '<path_to_trajectory>'
top = '<path_to_topology>'
m = Interface(trj, top)
Optionally, if you want to use the alpha shape functionality, change the use_alpha_shape property to True:
m.use_alpha_shape = True
Lastly, select the atom groups you want to consider, atom groups that comprise the interface, and run the calculate_density method:
m.select_atoms('all') # Consider every atom in the system
m.select_structure('<selection>') # resname of the nanoparticle
density_selection = 'resname TIP3'
m.calculate_density(density_selection)
Note that calculate_density uses every CPU core. You can specify the number of cores you want to use with the keyword argument cpu_count.
A more customized usage of the calculate_density method can be:
distances, densities = m.calculate_density(density_selection, start=10, end=1000, skip=2, norm_bin_count=12)
This version will start the calculation at the 10th frame and finish it at frame 1000 considering every 2nd frame. norm_bin_count specifies the number of divisions of the simulation box in each dimension to create a grid.
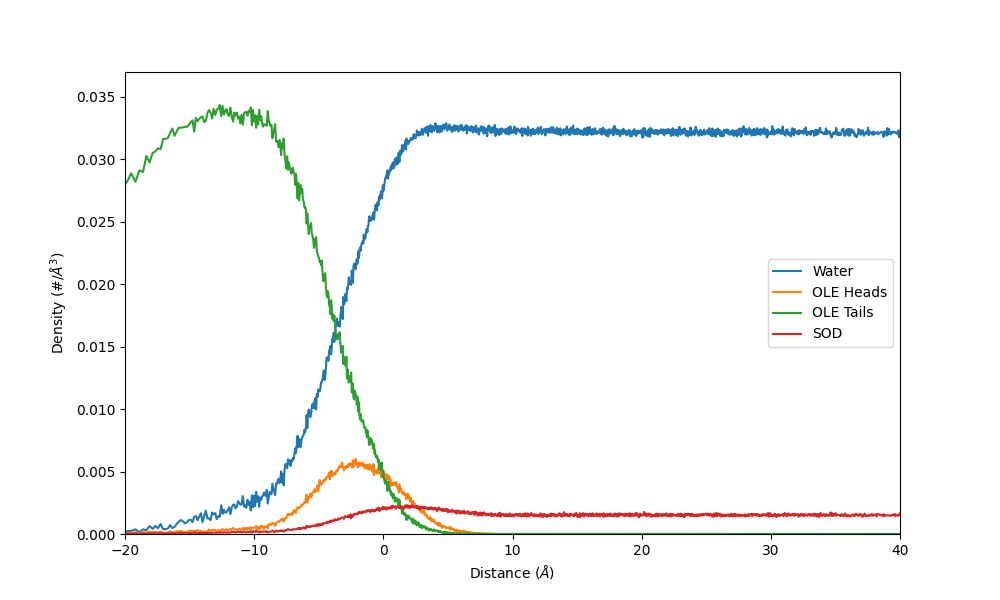
An example figure which shows the number density of different residues relative to the distance to the interface of a sodium oleate micelle:
Solubilized molecule count
PUCHIK also offers functionality for calculating the number of solubilized small molecules within a nanoparticle. This can be accomplished using the mol_count method. The signature of this method is identical to that of the calculate_density method. Here is an example usage:
sol = m.mol_count('resname TIP3 and type O', start=0, end=500)
Volume and area
As a direct consequence of using a convex hull, the volume of the hull can be easily extracted. The calculate_volume method does just that:
v = m.calculate_volume(start=500, end=1000, skip=2)
area keyword argument can be set to True to return the area of the hull as well:
v, a = m.calculate_volume(area=True, start=500, end=1000, skip=2)
Cite
If you've used this package in your project please cite the following paper:
Ishkhanyan, H., Santana-Bonilla, A., & Lorenz, C. D. (2025). PUCHIK: A Python Package To Analyze Molecular Dynamics Simulations of Aspherical Nanoparticles. Journal of Chemical Information and Modeling. https://doi.org/10.1021/ACS.JCIM.4C02128
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file puchik-1.2.5.tar.gz.
File metadata
- Download URL: puchik-1.2.5.tar.gz
- Upload date:
- Size: 755.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6c1144bdb712afd60262aeed41ad744a0a1b9e727dd9d0030be42f8c0f3bd9f9
|
|
| MD5 |
846c00ac963e0854d05afcde5c94db9a
|
|
| BLAKE2b-256 |
9947e6f8299b3de70a4a4fe2febc825a2bb0b1de4cc355d75f343e6f1f36bdba
|
File details
Details for the file puchik-1.2.5-cp310-cp310-win_amd64.whl.
File metadata
- Download URL: puchik-1.2.5-cp310-cp310-win_amd64.whl
- Upload date:
- Size: 759.6 kB
- Tags: CPython 3.10, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4264a0412ffdcba4bcdc20ef6ac9f6e2ea04c9f5f9735d350af020a83935f7de
|
|
| MD5 |
918ea25a6e627e39da93505c0e64879e
|
|
| BLAKE2b-256 |
5f06fd2164533557940e33ee9127aca9739d75ceb3574a8980f922244aced94e
|