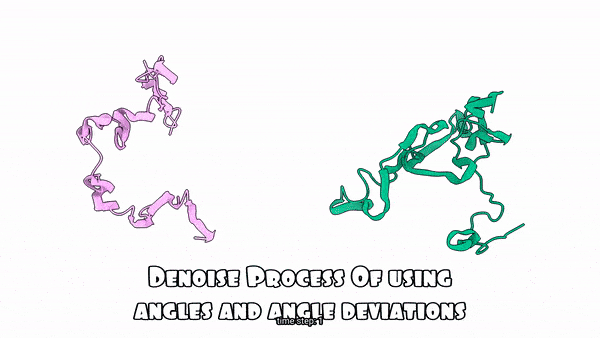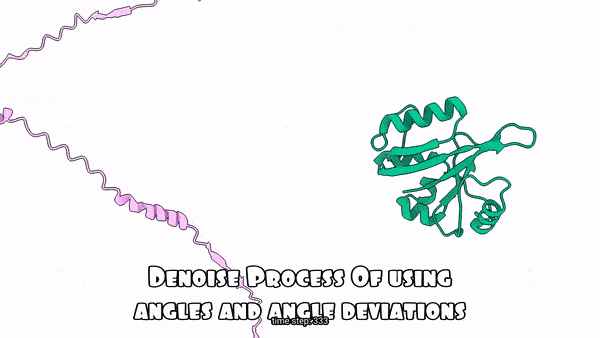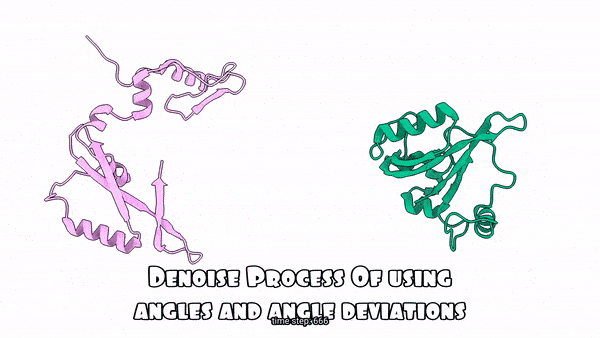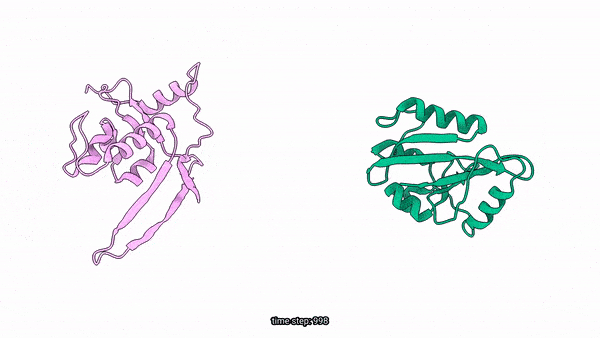This is a Transformer-Based diffusion model for Efficient Protein Conformational Ensemble Generation
Project description
Angular Deviation Diffuser
Overview
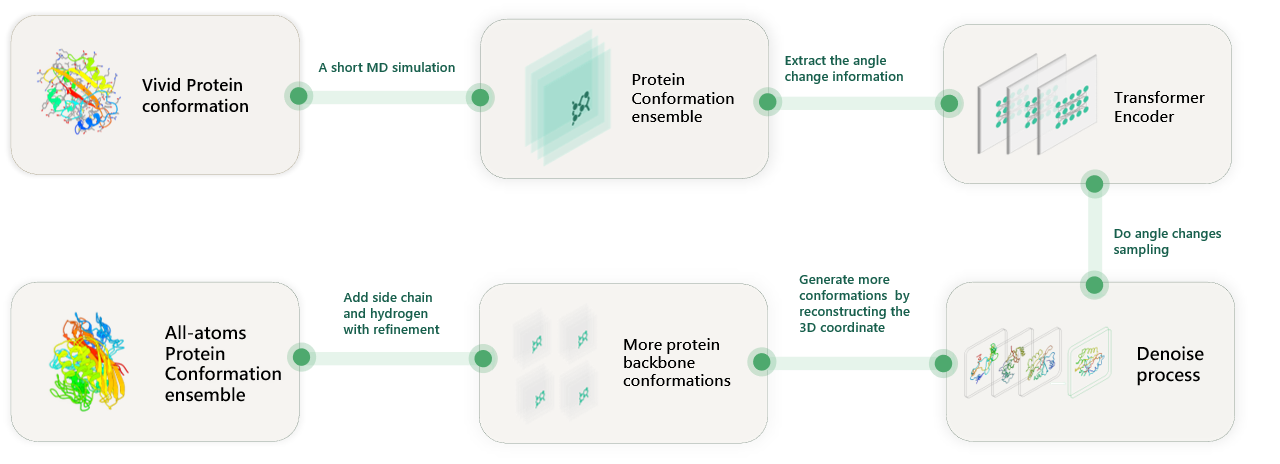
Angular Deviation Diffuser is a transformer-based diffusion model designed for efficiently generating conformational ensembles of protein backbones by using angular deviations as data flow. It aims to overcome the limitations of traditional molecular dynamics (MD) simulations by providing a fast and computationally efficient approach for sampling protein conformational landscapes. This model leverages the concepts of SE(3) symmetry, angular deviations, and diffusion processes to produce dynamic ensembles that closely match those generated through MD simulations, thereby offering a new way to study protein structure and function.
Overview of this work
Computational results
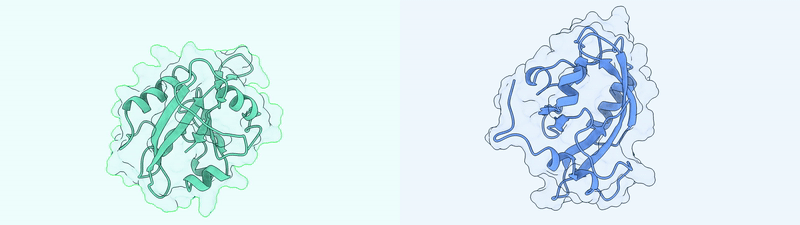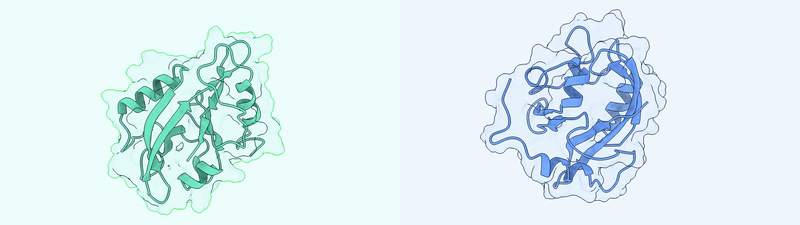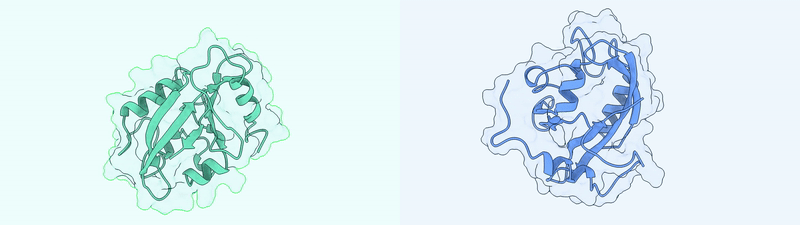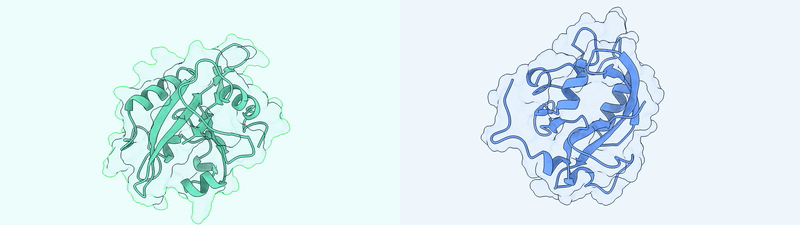
Generated Conformations Example (Dark State and Light State)
Using absolute angles vs. Using angle deviations for the denoising process(Sampling Process)
Background
Protein dynamics are essential for understanding biological functionality, as proteins exist not only in a single static structure but also in multiple dynamic conformational states. MD simulations are the gold standard for studying these dynamics, but they are resource-intensive and limited in their ability to fully explore all possible conformational states. The Angular Deviation Diffuser addresses these limitations by utilizing advanced deep learning techniques, specifically a diffusion model integrated with SE(3) invariance, to efficiently generate accurate protein conformations.
Features
- Angular Deviation-Based Diffusion: Uses angular deviations rather than absolute angles for data representation, improving stability and efficiency.
- Transformer Backbone: Utilizes a transformer architecture for learning protein dynamics from training data, capturing the conformational space effectively.
- SE(3) Symmetry Integration: Ensures the generated conformations respect the inherent rotational and translational symmetry of molecular systems.
- Efficient Ensemble Generation: Capable of generating diverse conformational ensembles in significantly less time compared to traditional MD simulations.
Installation
To install and use Angular Deviation Diffuser, follow these steps:
Prerequisites
- Conda: Ensure that Conda is installed to manage the environment and dependencies.
Steps
-
Create and Activate Conda Environment:
conda create -n angular_deviation_diffusion python==3.8 conda activate angular_deviation_diffusion
-
Install Angular Deviation Diffuser:
pip install Angular_Deviation_Diffuser python -c 'import pyrosetta_installer; pyrosetta_installer.install_pyrosetta()'
Usage
The following sections provide detailed guidance on how to use the package for generating protein conformations:
1. Extract Six Types of Angles
Extract the backbone angles (ϕ, ψ, ω, θ₁, θ₂, θ₃) from a given PDB file.
from Angular_Deviation_Diffuser import extract_six_angles
angles = extract_six_angles.get_angle_from_pdb("your_pdb_file.pdb")
Replace "your_pdb_file.pdb" with the appropriate PDB file name. This function returns an angle matrix containing all six types of backbone angles.
2. Reconstruct 3D Coordinates and Generate a PDB File
Reconstruct the 3D atomic coordinates using the six angle types and generate the corresponding PDB file.
from Angular_Deviation_Diffuser import reconstruct_coordinate
# Given an L x 6 angle matrix, reconstruct the Cartesian coordinates of the atoms.
# Replace 'angles' with the actual angle data in a numpy array type.
coor = reconstruct_coordinate.angles2coord(angles)
# Save the reconstructed coordinates to a PDB file
reconstruct_coordinate.coor_to_pdb(coor, "reconstructed_structure.pdb")
Replace "reconstructed_structure.pdb" with the desired output PDB file name.
3. Training the Model
Train the transformer-based diffusion model using the angular deviation data obtained from the previous step.
python training.py --data_dir 'path/to/training_data'
Replace 'path/to/training_data' with the path to your training dataset. The training set can be downloaded from here.
4. Generating Conformations
Generate a diverse ensemble of protein backbone conformations using the trained model.
from Angular_Deviation_Diffuser import sampling
# Generate conformations with refinement
sampling.generate_conformations_with_refinement(batch_size=10, total_samples=10)
The pre-trained model weights can be downloaded from here.
5. Adding Side Chains with Refinement
Utilize PyRosetta to add side chains to the generated backbone structures and refine them.
from Angular_Deviation_Diffuser import refine
# Refine the generated backbone structure
refine.refine_conformations('reconstructed_structure.pdb', "refined_conformation.pdb")
Replace 'reconstructed_structure.pdb' and "refined_conformation.pdb" with the appropriate file names for input and output.
Online Training and Sampling
Google Colab:https://colab.research.google.com/drive/1paTyFVRMzD4b75DeFjYyXlMV38d1eE1I#scrollTo=Dg41j6Feaj6f.
License
This project is licensed under the MIT License. See the LICENSE file for more information.
Acknowledgements
We are grateful to our research team for their invaluable contributions and support throughout the development of this model.
If you have any issues or questions, please feel free to open an issue in the repository or contact us directly.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file angular_deviation_diffuser-1.0.9.tar.gz.
File metadata
- Download URL: angular_deviation_diffuser-1.0.9.tar.gz
- Upload date:
- Size: 16.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.8.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4761c298bc2b1f3ee7f9c85f096d6b3c288ed84ea49fa3c1e5424987b6aa1eb5
|
|
| MD5 |
cc3fc976e321aa19222358d1e5a731fd
|
|
| BLAKE2b-256 |
71eafe73eda4dc32291f9b46b347cb3dce8c881cc92e339b6c516dcf9d4342fc
|
File details
Details for the file Angular_Deviation_Diffuser-1.0.9-py3-none-any.whl.
File metadata
- Download URL: Angular_Deviation_Diffuser-1.0.9-py3-none-any.whl
- Upload date:
- Size: 15.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.8.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
421ce8a75d2c3bbc02a450476ed721546d3fddf1fb5aa0b6a40fd76f880f8e47
|
|
| MD5 |
656d31a3762bb8407fed169740b86319
|
|
| BLAKE2b-256 |
4ac09c19f7df460de499a9682c3f726769dacfcf1c4df516b9b453b6898a2d39
|