Database-free identification of cell-cell interaction genes from spatial transcriptomics
Project description
CellNeighborEX v2: Identifying Context-Specific Cell-Cell Interaction Genes Without Ligand-Receptor Databases from Spatial Transcriptomics
CellNeighborEX v2 is a database-free computational framework for identifying cell–cell interaction (CCI)–associated genes from spatial transcriptomics (ST) data. Most existing CCI inference approaches rely on predefined ligand–receptor databases, which limits their ability to capture diverse interaction mechanisms, particularly in low-resolution ST platforms such as Visium.
CellNeighborEX v2 addresses this limitation by detecting deviations between observed and expected gene expression at the spot-population level, enabling direct identification of genes influenced by neighboring cell types. These deviations are evaluated using a hybrid statistical framework based on permutation testing and cell-type pair abundance.
Unlike conventional ligand–receptor–based methods, CellNeighborEX v2 can capture a broad spectrum of intercellular communication mechanisms, including both paracrine signaling and contact-dependent interactions.
Across multiple datasets—including hippocampus, liver cancer, colorectal cancer, ovarian cancer, and lymph node infection—CellNeighborEX v2 accurately recapitulates previously reported CCIs and reveals additional context-specific interactions not represented in existing databases.
By expanding the analytical potential of low-resolution ST data, CellNeighborEX v2 provides a powerful tool for exploring the molecular language of intercellular communication in complex tissues.
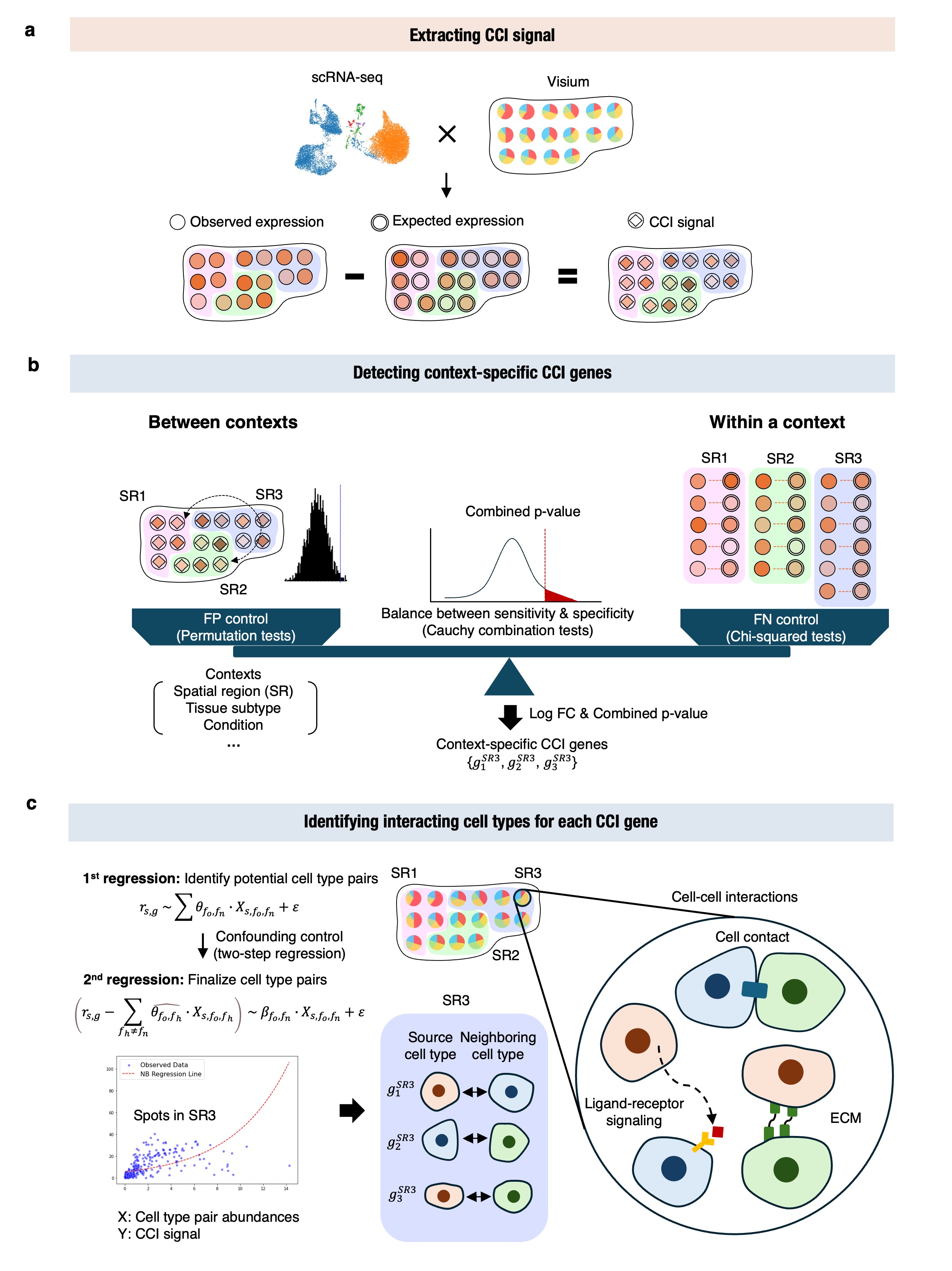
Workflow
The figure below shows the workflow of CellNeighborEX v2:
Installation
CellNeighborEX v2 requires Python version >= 3.9 and < 3.11. We recommend using conda environment to avoid dependency conflicts. The dependencies are listed in requirements.txt.
# Create conda environment “CellNeighborEX-env”
conda create -n CellNeighborEX-env python=3.9
conda activate CellNeighborEX-env
# Navigate into the directory where requirements.txt is located. Then, install dependencies
pip install -r requirements.txt
# Install CellNeighborEX v2 from PyPI
pip install CellNeighborEX
Source code
GitHub repository:
https://github.com/hkim240/CellNeighborEX-v2
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cellneighborex-2.0.0.tar.gz.
File metadata
- Download URL: cellneighborex-2.0.0.tar.gz
- Upload date:
- Size: 3.9 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.8.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
25d14514a3452d9993ccf450d2f582df864a243ef0e5370b9610d0db547f9126
|
|
| MD5 |
0d7a1c94b482f9837e0e643164972da0
|
|
| BLAKE2b-256 |
50d906f4c487703b0fbe8c0ddd62b417344482b6acf885cc712828278c2a1378
|
File details
Details for the file cellneighborex-2.0.0-py3-none-any.whl.
File metadata
- Download URL: cellneighborex-2.0.0-py3-none-any.whl
- Upload date:
- Size: 4.0 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.8.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1e2c18eafbed3820a7072a1b6a854abb15bb136ab6b535f01fd120fb168af126
|
|
| MD5 |
e00e21edb2abda9ba6f87b7a5b28f68f
|
|
| BLAKE2b-256 |
06879d1c227e863a513501a2bd34c6bb26f649b9d09dd9c28d5e7aa1a7fa1362
|