Genomic decomposition and reconstruction of non-tumor diploid subclones
Project description
CLEMENT
- Genomic decomposition and reconstruction of non-tumor diploid subclones (2023)
- CLonal decomposition via Expectation-Maximization algorithm established in Non-Tumor setting
- Support multiple diploid sample
- Biallelic variants (Homo, 1/1) can degrade the performance of CLEMENT.
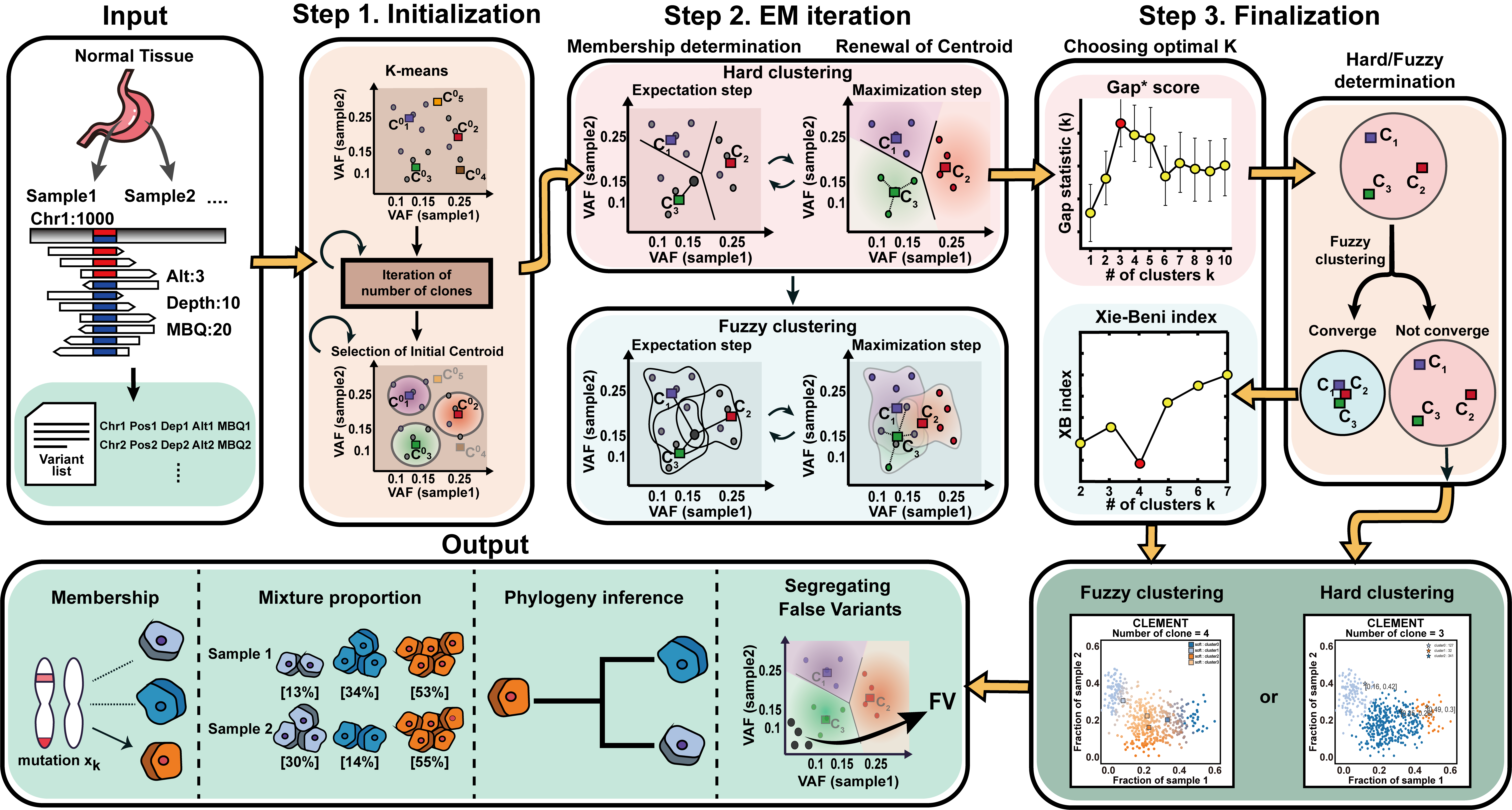
Overview of CLEMENT workflow and core algorithms
Installation
Dependencies
- python 3.6.x
- matplotlib 3.5.2
- seaborn 0.11.2
- numpy 1.21.5
- pandas 1.3.4
- scikit-learn 1.0.2
- scipy 1.7.3
- palettable 3.3.0
Install from github
-
git clone https://github.com/Yonsei-TGIL/CLEMENT.git
cd CLEMENT
pip3 install . -
pip3 install git+https://github.com/Yonsei-TGIL/CLEMENT.git
Install from PyPi
- pip3 install CLEMENTDNA
Version update
1.0.11 (Jan 1st, 2024)
Input format
As now of 1.0.4, CLEMENT only supports standardized TSV input. Examples of input file is shown in "example" directory.
- 1st column: mutation ID (CHR_POS is recommended)
- 2nd column: label (answer), if possible. If user don't know the label (answer), just set 0
- 3rd column: Depth1,Alt1,Depth2,Alt2....,Depth_n,Alt_n * should be comma-separated, and no space permitted
- 4th column: BQ1,BQ2....,BQ_n * should be comma-separated, and no space permitted. If absent, CLEMENT set default BQ as 20.
Running
command line interface
CLEMENT [OPTIONS]
options
(Mandatory) These options are regarding User's input and output format
--INPUT_TSV Input data whether TSV. The tool automatically detects the number of samples
--CLEMENT_DIR Directory where the outputs of CLEMENT be saved
These options are regarding downsizing User's input or not
--RANDOM_PICK Set this variable to user want to downsize the sample. If user don't want to downsize, set -1. (default : -1).
These options are regarding the selection of likelihood model
--MODEL Model for TP, FN in E-step. (default: betabinomial)
--CONSTANT Constant multiplier for alpha and beta in beta-binomila distribution. (default:1)
These options are adjusting E-M algorithm parameter
--NUM_CLONE_TRIAL_START Minimum number of expected cluster_hards (initation of K) (default: 3)
--NUM_CLONE_TRIAL_END Maximum number of expected cluster_hards (termination of K) (default: 5)
--TRIAL_NO Trial number in each candidate cluster_hard number. DO NOT recommend over 15 (default: 5)
--FP_PRIOR FP_PRIOR Prior of false positive (FP). Recommendation : <= 0.1. (default : 0.01)
--TN_PRIOR TN_PRIOR Prior of true negative (TN). Recommendation : > 0.99. (default : 0.99)
--KMEANS_CLUSTERNO Number of initial K-means cluster. Recommendation : 5~8 for one-sample, 8-15 for larger-sample (default: 8)
--MIN_CLUSTER_SIZE The minimum cluster size that is acceptable. Recommendation : 1-3% of total variants number (default: 9)
Other options
--MODE Selection of clustering method. "Hard": hard clustering only, "Both": both hard and soft (fuzzy) clustering (default: "Both")
--MAKEONE_STRICT 1: strict, 2: lenient, 3: most lenient (default : 1)
--SCORING True : comparing with the answer set, False : just visualization (default: False)
Miscelleneous
--FONT_FAMILY Font family that displayed in the plots (default : "arial")
--VISUALIZATION Whether produce image in every E-M step (default: True)
--IMAGE_FORMAT Image format that displayed in the plots (default : jpg)
--VERBOSE 0: no record, 1: simplified record, 2: verbose record (default: 2)
output
${CLEMENT_DIR}"/result"
- CLEMENT_decision CLEMENT's best recommendation among hard and soft clustering.
- CLEMENT_hard_1st CLEMENT's best decomposition by hard clustering.
- CLEMENT_hard.gapstatistics.txt Selecting the optimal K in hard clustering based on gap* stastics.
- CLEMENT_soft_1st CLEMENT's best decomposition by soft (fuzzy) clustering.
- membership.txt Membership assignment of all variants to each clusters.
- membership_count.txt Count matrix of the membership assignment to each clusters.
- mixture.txt Centroid of each clusters
Example
DIR=[YOUR_DIRECTORY]
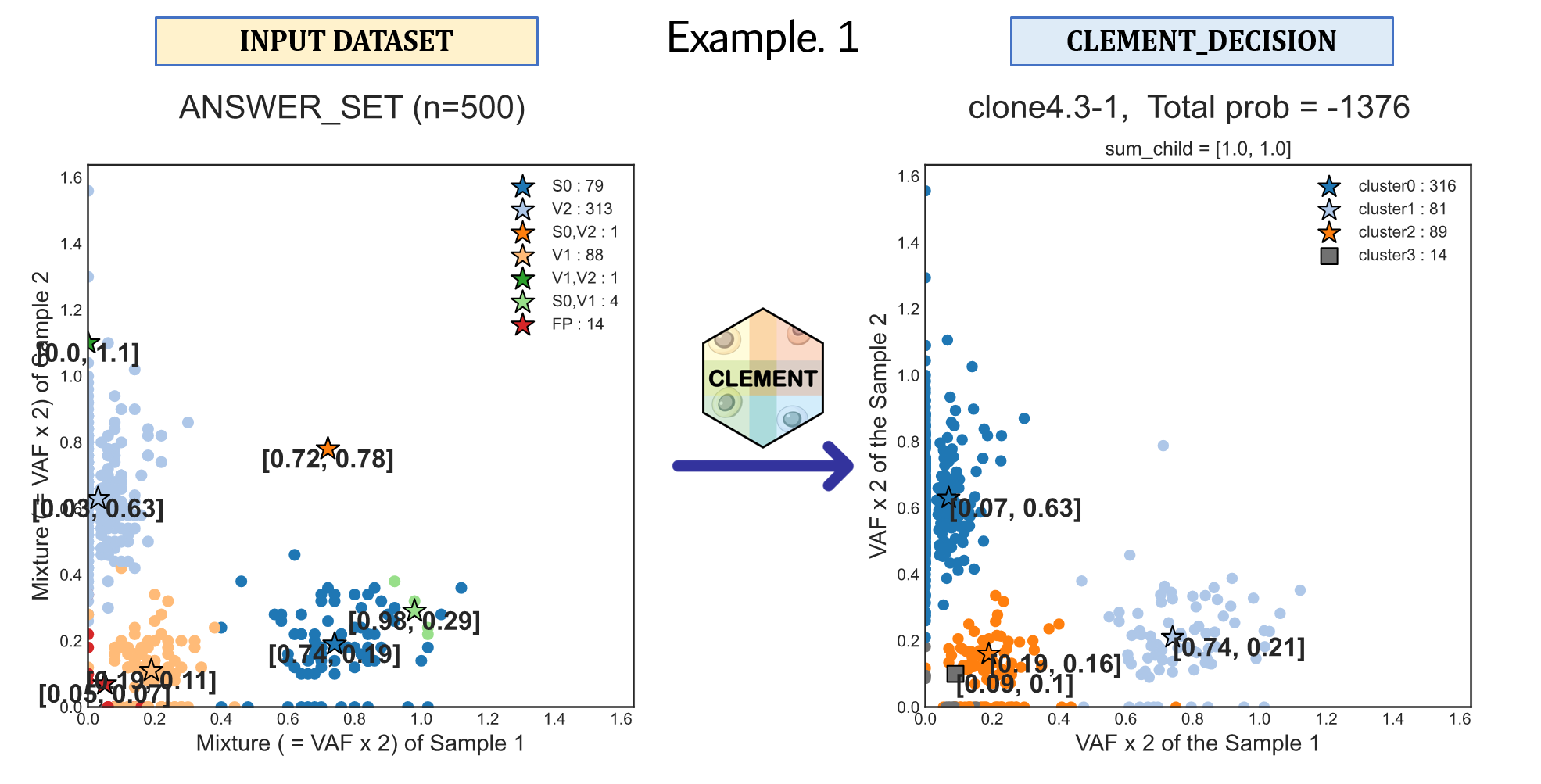
# Example 1
CLEMENT \
--INPUT_TSV ${DIR}"/example/1.SimData/SimData_1D/n500_125x/lump/0.0/clone_4/1/1.txt" \
--CLEMENT_DIR ${DIR}"/example/1.SimData/SimData_1D/n500_125x/lump/0.0/clone_4/1" \
--NUM_CLONE_TRIAL_START 1 \
--NUM_CLONE_TRIAL_END 5
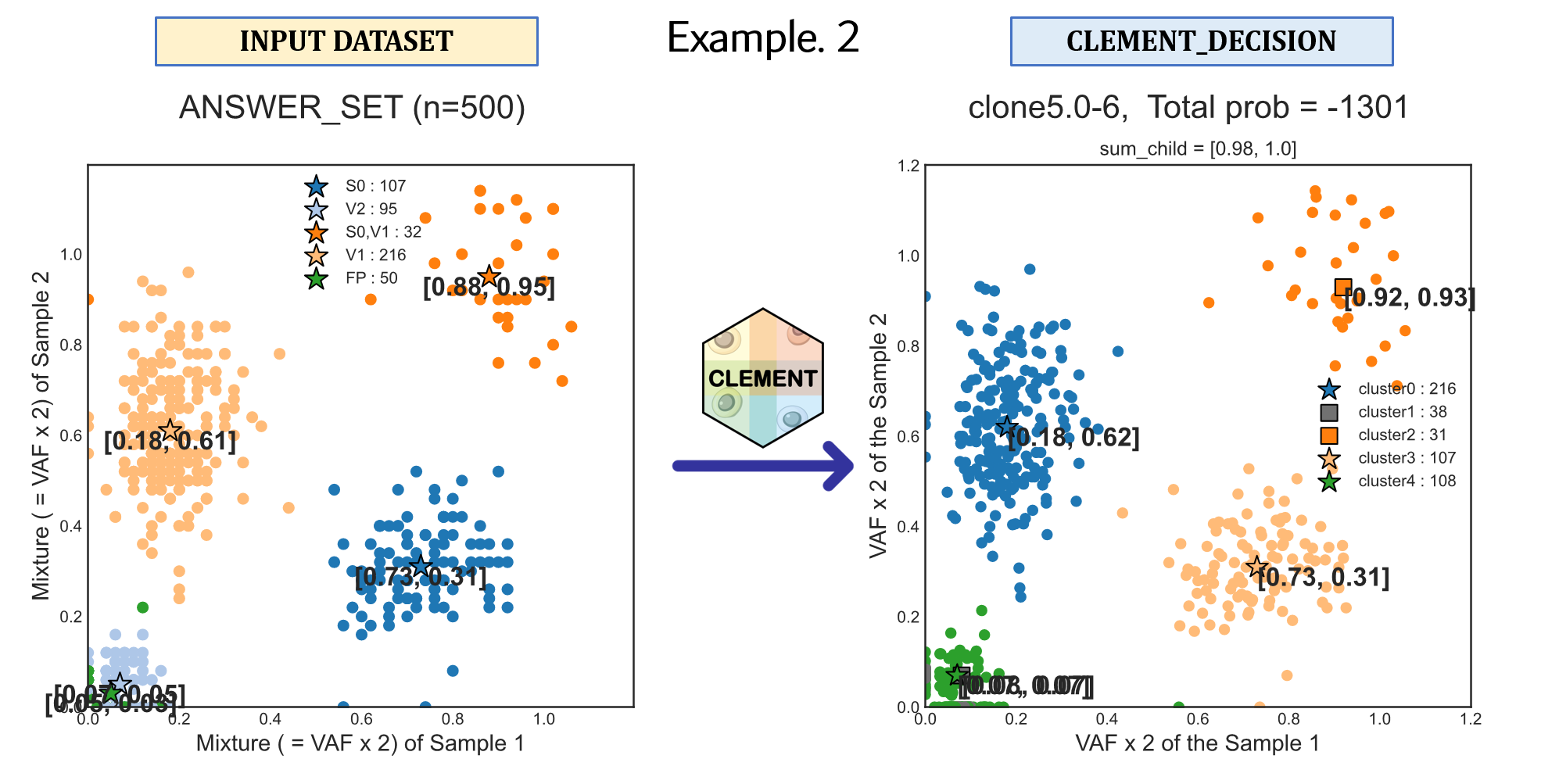
# Example 2
CLEMENT \
--INPUT_TSV ${DIR}"/example/2.CellData/MRS_2D/M1-8_M2-4/M1-8_M2-4_input.txt" \
--CLEMENT_DIR ${DIR}"/example/2.CellData/MRS_2D/M1-8_M2-4" \
--NUM_CLONE_TRIAL_START 2 \
--NUM_CLONE_TRIAL_END 6 \
--RANDOM_PICK 500
Contact
goldpm1@yuhs.ac
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file CLEMENTDNA-1.0.12.tar.gz.
File metadata
- Download URL: CLEMENTDNA-1.0.12.tar.gz
- Upload date:
- Size: 27.8 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.7.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4f48977805c6c681519ac60e178c709756b6d0d218b1a9559cc54a843c701e99
|
|
| MD5 |
b4ec038148b25409d000701994984c01
|
|
| BLAKE2b-256 |
48365aba1dd40aa5428dfe274a7b66b8822edc6846f9f885f77ed5a91f1a1a57
|
File details
Details for the file CLEMENTDNA-1.0.12-py3-none-any.whl.
File metadata
- Download URL: CLEMENTDNA-1.0.12-py3-none-any.whl
- Upload date:
- Size: 170.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.7.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d6a8825864bee8553160c171408997278a8bbab4dbfe61631cf9fc33f11eb0ff
|
|
| MD5 |
bebde17f040f65930d1ac2f1b520926b
|
|
| BLAKE2b-256 |
ecad1009fa99565a3444459a76454fe2897ab8f46a53e536bdfb9a7703bc5121
|











