Deep Velocity
Project description
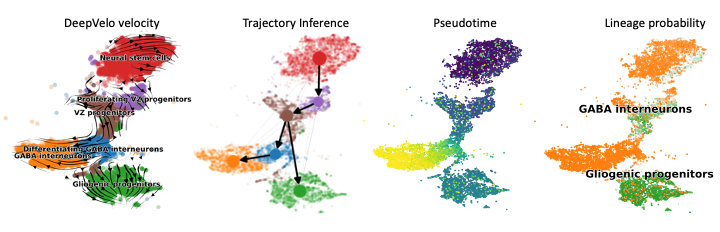
DeepVelo - A Deep Learning-based velocity estimation tool with cell-specific kinetic rates
This is the official implementation of the DeepVelo method. DeepVelo employs cell-specific kinetic rates and provides more accurate RNA velocity estimates for complex differentiation and lineage decision events in heterogeneous scRNA-seq data. Please check out the paper for more details.
Installation
pip install deepvelo
The dgl package is required, the cpu version is installed by default. Feel free to install the dgl cuda version for GPU acceleration.
pip install dgl-cu101>=0.4.3 # an example for CUDA 10.1
Install the development version
We use poetry to manage dependencies.
poetry install
This will install the exact versions in the provided poetry.lock file. If you want to install the latest version for all dependencies, use the following command.
poetry update
Usage
We provide a number of notebooks in the exmaples folder to help you get started. DeepVelo fullly integrates with scanpy and scVelo. The basic usage is as follows:
import deepvelo as dv
import scvelo as scv
adata = ... # load your data in AnnData format
# preprocess the data
scv.pp.filter_and_normalize(adata, min_shared_counts=20, n_top_genes=2000)
scv.pp.moments(adata, n_neighbors=30, n_pcs=30)
# run DeepVelo using the default configs
trainer = dv.train(adata, dv.Constants.default_configs)
# this will train the model and predict the velocity vectore. The result is stored in adata.layers['velocity']. You can use trainer.model to access the model.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file deepvelo-0.2.4.tar.gz.
File metadata
- Download URL: deepvelo-0.2.4.tar.gz
- Upload date:
- Size: 58.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.7.3 Linux/4.4.0-210-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a337148d6666a8bfcb4d3500dcb3b511e4c31279a509994877475d3a37aa7aa8
|
|
| MD5 |
5205cd0c75c0511763e085f5cb72bf39
|
|
| BLAKE2b-256 |
5e53f763cfc3eb3bb9f2447810f86d46351608989c92037d8e5e4ccf996a2b8c
|
File details
Details for the file deepvelo-0.2.4-py3-none-any.whl.
File metadata
- Download URL: deepvelo-0.2.4-py3-none-any.whl
- Upload date:
- Size: 66.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.1.13 CPython/3.7.3 Linux/4.4.0-210-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
61d2fc6b537195b74fb064f6f4b6be5cf467b8f6fecd21fda25a8259b5347ced
|
|
| MD5 |
4e8253ec5c191517b18c77864dc1544e
|
|
| BLAKE2b-256 |
7eba11d98db0b617b02d7756bd541bef17a006ffe2c171c41430c977c209e879
|