nDTomo software suite
Project description
nDTomo Software Suite
nDTomo is a Python-based software suite for the simulation, visualization, pre-processing, reconstruction, and analysis of chemical imaging and X-ray tomography data, with a focus on hyperspectral datasets such as X-ray powder diffraction computed tomography or XRD-CT.
It includes:
- A suite of notebooks and scripts for advanced processing, sinogram correction, CT reconstruction, peak fitting, and machine learning-based analysis
- A PyQt-based graphical user interface (GUI) for interactive exploration and analysis of hyperspectral tomography data
- A growing collection of simulation tools for generating phantoms and synthetic datasets
The software is designed to be accessible to both researchers and students working in chemical imaging, materials science, catalysis, battery research, and synchrotron radiation applications.
📘 Official documentation: https://ndtomo.readthedocs.io
Key Capabilities
nDTomo provides tools for:
- Interactive visualization of chemical tomography data via the
nDTomoGUI - Generation of multi-dimensional synthetic phantoms
- Simulation of pencil beam CT acquisition strategies
- Pre-processing and correction of sinograms
- CT image reconstruction using algorithms like filtered back-projection and SIRT
- Dimensionality reduction and clustering for unsupervised chemical phase analysis
- Pixel-wise peak fitting using Gaussian, Lorentzian, and Pseudo-Voigt models
- Peak fitting using the self-supervised PeakFitCNN
- Simultaneous peak fitting and tomographic reconstruction using the DLSR approach with PyTorch GPU acceleration
- Registration 2D, 3D and point cloud with PyTorch
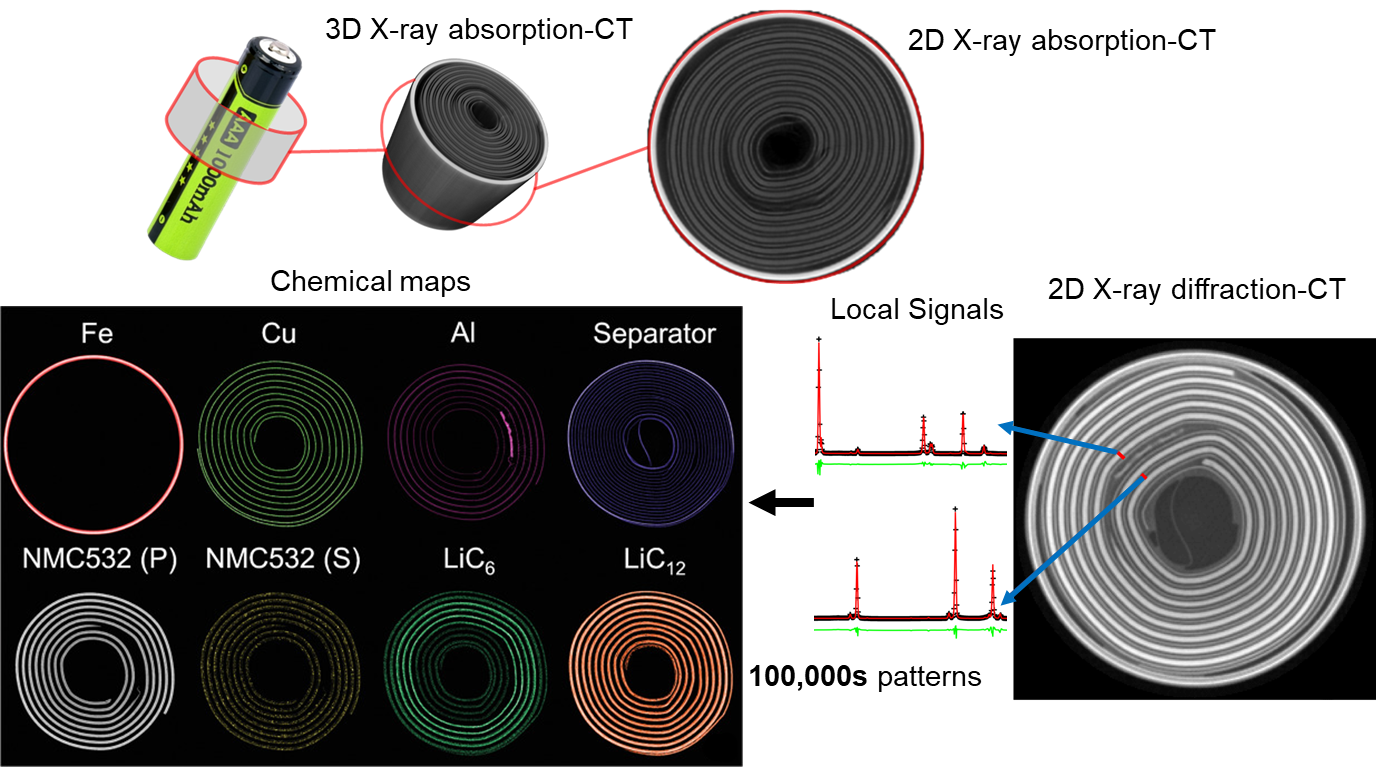
Figure: Comparison between X-ray absorption-contrast CT (or microCT) and X-ray diffraction CT (XRD-CT or Powder diffraction CT) data acquired from a cylindrical Li-ion battery. For more details regarding these XRD-CT studies using cylindrical Li-ion batteries see [1,2].
Included Tutorials
The repository includes several example notebooks to help users learn the API and workflows:
| Notebook Filename | Topic |
|---|---|
tutorial_phantoms.ipynb |
Generating and visualizing 2D/3D phantoms |
tutorial_pencil_beam.ipynb |
Simulating pencil beam CT data with different acquisition schemes |
tutorial_detector_calibration.ipynb |
Calibrating detectors and integrating diffraction patterns using pyFAI |
tutorial_texture_2D_diffraction_patterns.ipynb |
Investigating the effects of texture on 2D powder patterns |
tutorial_sinogram_handling.ipynb |
Pre-processing, normalization, and correction of sinograms |
tutorial_ct_recon_demo.ipynb |
CT image reconstruction from sinograms using analytical and iterative methods |
tutorial_dimensionality_reduction.ipynb |
Unsupervised learning for phase identification in tomography |
tutorial_peak_fitting.ipynb |
Peak fitting in synthetic XRD-CT datasets |
tutorial_peak_fit_cnn.ipynb |
Peak fitting in GPU using a self-supervised PeakFitCNN |
tutorial_DLSR.ipynb |
Simultaneous peak fitting and CT reconstruction in GPU using the DLSR method |
tutorial_registration.ipynb |
Registration 2D, 3D and point cloud with PyTorch |
Each notebook is designed to be standalone and executable, with detailed inline comments and example outputs.
Note:
-
Binder is built with CPU-only support (including
torch) and can be used to run all notebooks. However, some notebooks may take longer to execute due to the lack of GPU acceleration. -
Google Colab provides GPU support and
torchis preinstalled. You will also need to installnDTomoat the beginning of each notebook session.
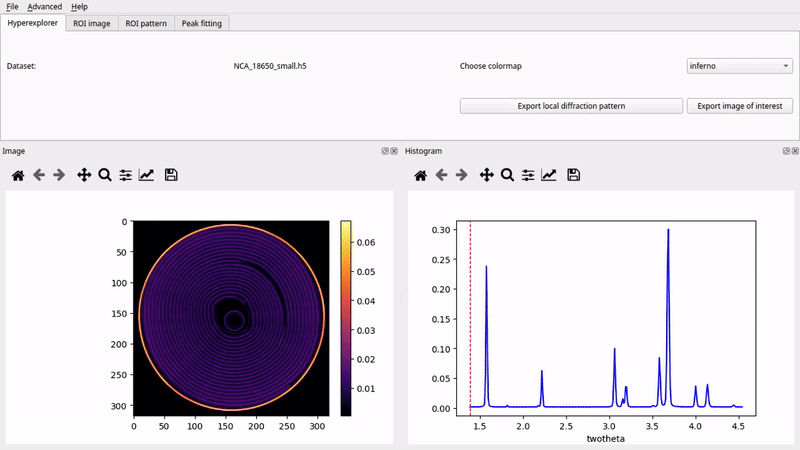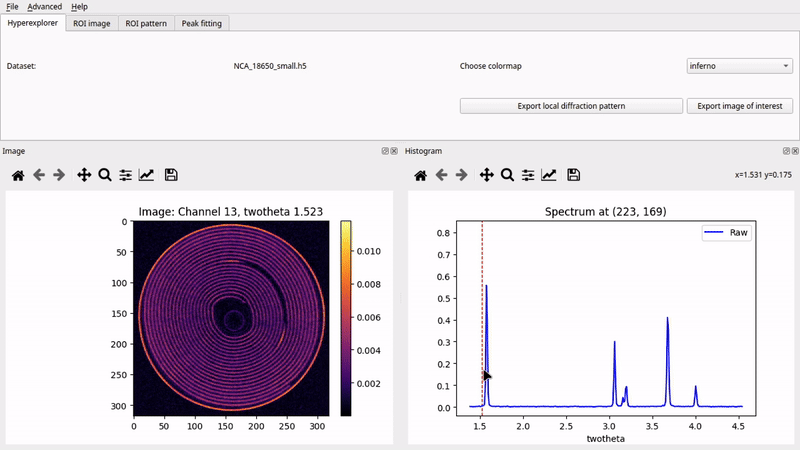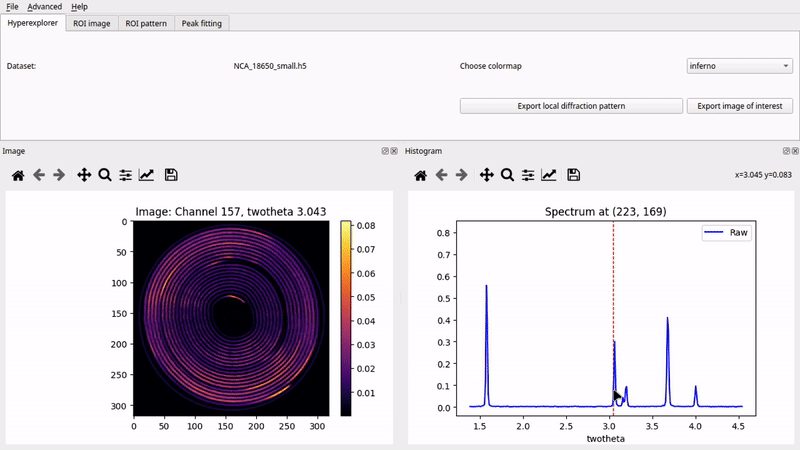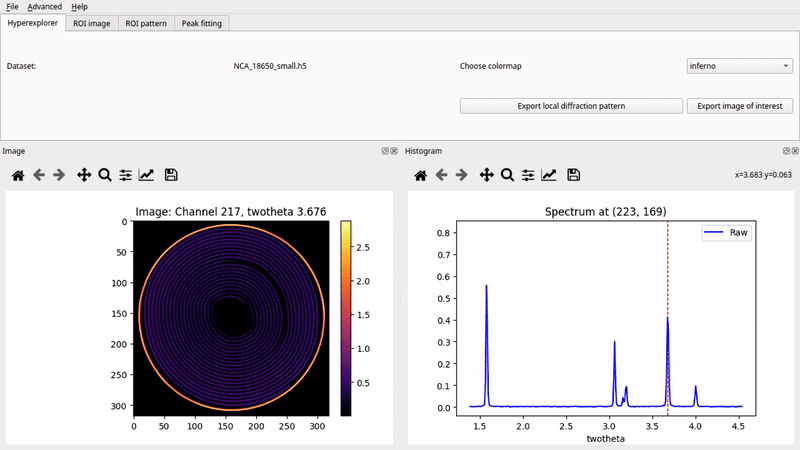
Graphical User Interface (nDTomoGUI)
The nDTomoGUI provides a complete graphical environment for:
- Loading
.h5/.hdf5chemical imaging datasets - Visualizing 2D slices and 1D spectra interactively
- Segmenting datasets using channel selection and thresholding
- Extracting and exporting local diffraction patterns
- Performing single-peak batch fitting across regions of interest
- Generating a synthetic XRD-CT phantoms for development tests
- Using an embedded IPython console for advanced control and debugging
The GUI is described in more detail in the online documentation and supports both novice and expert workflows.
Launch with:
conda activate ndtomo
python -m nDTomo.gui.nDTomoGUI
Installation Instructions
To make your life easier, please install Anaconda. The nDTomo library and all associated ode can be installed by following the next three steps:
1. Install astra-toolbox
An important part of the code is based on astra-toolbox, which is currently available through conda.
It is possible to install astra-toolbox from sources (i.e., if one wants to avoid using conda), but it is not a trivial task. We recommend creating a new conda environment for nDTomo.
Create a new environment and first install astra-toolbox:
conda create --name ndtomo python=3.11
conda activate ndtomo
conda install -c astra-toolbox -c nvidia astra-toolbox
2. Install nDTomo from GitHub
You can choose one of the following options to install the nDTomo library:
a. To install using pip:
pip install nDTomo
b. To install using Git:
pip install git+https://github.com/antonyvam/nDTomo.git
For development work (editable install):
git clone https://github.com/antonyvam/nDTomo.git && cd nDTomo
pip install -e .
c. For local installation after downloading the repo:
Navigate to where the setup.py file is located and run:
pip install --user .
or:
python3 setup.py install --user
3. Install PyTorch
The neural networks, as well as any GPU-based code, used in nDTomo require Pytorch which can be installed through pip.
For example, for Windows/Linux with CUDA 11.8:
pip install torch torchvision torchaudio --index-url https://download.pytorch.org/whl/cu118
Launching the GUI
After installing nDTomo, the graphical user interface can be launched directly from the terminal:
conda activate ndtomo
python -m nDTomo.gui.nDTomoGUI
Diamond Light Source
As a user at the Diamond Light Source, you can install nDTomo by doing:
git clone https://github.com/antonyvam/nDTomo.git && cd nDTomo
module load python/3
python setup.py install --user
Citation
If you use parts of the code, please cite the work using the following paper:
nDTomo: A Modular Python Toolkit for X-ray Chemical Imaging and Tomography, A. Vamvakeros, E. Papoutsellis, H. Dong, R. Docherty, A.M. Beale, S.J. Cooper, S.D.M. Jacques, Digital Discovery, 4, 2579-2592, 2025, https://doi.org/10.1039/D5DD00252D
References
[1] "Cycling Rate-Induced Spatially-Resolved Heterogeneities in Commercial Cylindrical Li-Ion Batteries", A. Vamvakeros, D. Matras, T.E. Ashton, A.A. Coelho, H. Dong, D. Bauer, Y. Odarchenko, S.W.T. Price, K.T. Butler, O. Gutowski, A.-C. Dippel, M. von Zimmerman, J.A. Darr, S.D.M. Jacques, A.M. Beale, Small Methods, 2100512, 2021. https://doi.org/10.1002/smtd.202100512
[2] "Emerging chemical heterogeneities in a commercial 18650 NCA Li-ion battery during early cycling revealed by synchrotron X-ray diffraction tomography", D. Matras, T.E. Ashton, H. Dong, M. Mirolo, I. Martens, J. Drnec, J.A. Darr, P.D. Quinn, S.D.M. Jacques, A.M. Beale, A. Vamvakeros, Journal of Power Sources 539, 231589, 2022, https://doi.org/10.1016/j.jpowsour.2022.231589
Previous technical work (reverse chronological order)
[1] "Upsampling DINOv2 features for unsupervised vision tasks and weakly supervised materials segmentation", R. Docherty, A. Vamvakeros, S.J. Cooper, Advanced Intelligence Systems, 2026, accepted, preprint: https://doi.org/10.48550/arXiv.2410.19836
[2] "Maybe you don't need a U-Net: convolutional feature upsampling for materials micrograph segmentation", R. Docherty, A. Vamvakeros, S.J. Cooper, 2025, preprint: https://doi.org/10.48550/arXiv.2508.21529
[3] "Obtaining parallax-free X-ray powder diffraction computed tomography data with a self-supervised neural network", H. Dong, S.D.M. Jacques, K.T. Butler, O. Gutowski, A.-C. Dippel, M. von Zimmerman, A.M. Beale, A. Vamvakeros, npj Computational Materials 10 (1), 201, 2024, https://doi.org/10.1038/s41524-024-01389-1
[4] "SAMBA: A Trainable Segmentation Web-App with Smart Labelling", R. Docherty, I. Squires, A. Vamvakeros, S.J. Cooper, Journal of Open Source Software 9 (98), 6159, 2024, https://doi.org/10.21105/joss.06159
[5] "A scalable neural network architecture for self-supervised tomographic image reconstruction", H. Dong, S.D.M. Jacques, W. Kockelmann, S.W.T. Price, R. Emberson, D. Matras, Y. Odarchenko, V. Middelkoop, A. Giokaris, O. Gutowski, A.-C. Dippel, M. von Zimmermann, A.M. Beale, K.T. Butler, A. Vamvakeros, Digital Discovery, 2 (4), 967-980, 2023, https://doi.org/10.1039/D2DD00105E
[6] "A deep convolutional neural network for real-time full profile analysis of big powder diffraction data", H. Dong, K.T. Butler, D. Matras, S.W.T. Price, Y. Odarchenko, R. Khatry, A. Thompson, V. Middelkoop, S.D.M. Jacques, A.M. Beale, A. Vamvakeros, npj Computational Materials 7 (1), 74, 2021, https://doi.org/10.1038/s41524-021-00542-4
[7] "DLSR: a solution to the parallax artefact in X-ray diffraction computed tomography data", A. Vamvakeros, A.A. Coelho, D. Matras, H. Dong, Y. Odarchenko, S.W.T. Price, K.T. Butler, O. Gutowski, A.-C. Dippel, M. von Zimmermann, I. Martens, J. Drnec, A.M. Beale, S.D.M. Jacques, Journal of Applied Crystallography 53 (6), 1531-1541, https://doi.org/10.1107/S1600576720013576
[8] "5D operando tomographic diffraction imaging of a catalyst bed", A. Vamvakeros, S.D.M. Jacques, M. Di Michiel, D. Matras, V. Middelkoop, I.Z. Ismagilov, E.V. Matus, V.V. Kuznetsov, J. Drnec, P. Senecal, A.M. Beale, Nature communications 9 (1), 4751, 2018, https://doi.org/10.1038/s41467-018-07046-8
[9] "Interlaced X-ray diffraction computed tomography", A. Vamvakeros, S.D.M. Jacques, M. Di Michiel, P. Senecal, V. Middelkoop, R.J. Cernik and A.M. Beale, Journal of Applied Crystallography 49 (2), 485-496, 2016, https://doi.org/10.1107/S160057671600131X
[10] "Removing multiple outliers and single-crystal artefacts from X-ray diffraction computed tomography data", A. Vamvakeros, S.D.M. Jacques, M. Di Michiel, V. Middelkoop, C.K. Egan, R. J. Cernik, A. M Beale, Journal of Applied Crystallography 48 (6), 1943-1955, 2015, Jacques, Journal of Applied Crystallography 48 (6), 1943-1955, 2015, https://doi.org/10.1107/S1600576715020701
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file ndtomo-2026.1.tar.gz.
File metadata
- Download URL: ndtomo-2026.1.tar.gz
- Upload date:
- Size: 58.9 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d6dc63a81a32d71c9a01ac651da077741b6ad2ed172397c257b3d19766e459e7
|
|
| MD5 |
e2f4c89a427ed1ec8b9118165fa260d8
|
|
| BLAKE2b-256 |
0744aa9ab7f7e820a27f6ef8086df6f059789617e16fc18898ecff62393150eb
|
File details
Details for the file ndtomo-2026.1-py3-none-any.whl.
File metadata
- Download URL: ndtomo-2026.1-py3-none-any.whl
- Upload date:
- Size: 59.0 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
03b06af6c7591060fad015b773dd0973d99c009ea38b7c350ebc05a3850890b3
|
|
| MD5 |
e058ecc1c15d53e832f6e0f88c1a6d23
|
|
| BLAKE2b-256 |
44320be6fb27e1923bc7fb8f080aed30f532f39ce91e8fb6f5abd0334df7546e
|