Matplotlib/seaborn style presets matching scientific journal requirements, with validation, export safety, and preview capabilities.
Project description
plotstyle
Matplotlib figures formatted for journal submission, automatically.
PlotStyle makes it easy to produce Matplotlib figures that meet the exact typographic, dimensional, and export requirements of major academic journals. It also integrates with Seaborn, with more integrations planned. Pick a journal, create your figure, save it. PlotStyle handles the rest.
Table of Contents
- Installation
- Quick Start
- Examples
- Supported Journals
- CLI
- Documentation
- Contributing
- Citation
- License
Installation
Requires Python 3.10+ and Matplotlib >= 3.9.
pip install plotstyle
Optional extras:
pip install "plotstyle[color]" # colorblind / grayscale previews
pip install "plotstyle[seaborn]" # seaborn integration
pip install "plotstyle[all]" # everything
Quick Start
import numpy as np
import plotstyle
with plotstyle.use("nature") as style:
fig, ax = style.figure(columns=1) # sized to Nature's single-column width (89 mm)
x = np.linspace(0, 2 * np.pi, 200)
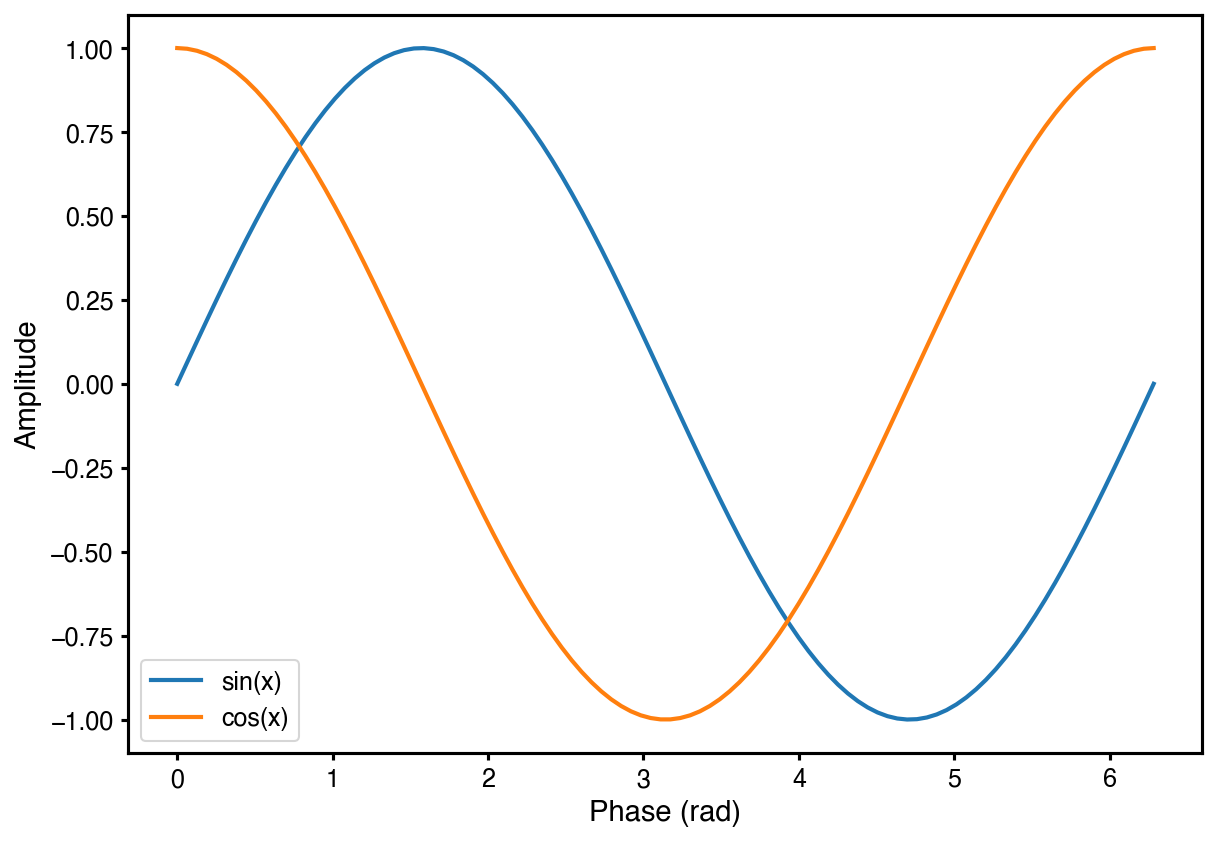
ax.plot(x, np.sin(x), label="sin(x)")
ax.plot(x, np.cos(x), label="cos(x)")
ax.set_xlabel("Phase (rad)")
ax.set_ylabel("Amplitude (a.u.)")
ax.legend()
style.savefig(fig, "figure.pdf") # 300 DPI minimum, TrueType fonts embedded
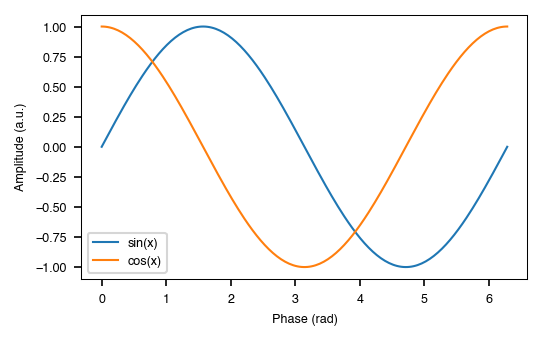
The with block is the recommended pattern. Matplotlib's rcParams are restored automatically when it exits, even if an exception occurs.
Examples
Multi-panel figures
style.subplots() works like plt.subplots() but sizes the figure to the journal spec and adds panel labels automatically. All built-in journal specs use bold lowercase labels (a, b, c, …). The label style is driven by each spec's panel_label_case field and can be lower, upper, parens_lower, parens_upper, sentence, or title.
import numpy as np
import plotstyle
rng = np.random.default_rng(42)
with plotstyle.use("science") as style:
fig, axes = style.subplots(nrows=2, ncols=2, columns=2)
x = np.linspace(0, 10, 100)
axes[0, 0].plot(x, np.sin(x), label="sin")
axes[0, 0].plot(x, np.cos(x), label="cos")
axes[0, 0].set_xlabel("x")
axes[0, 0].set_ylabel("f(x)")
axes[0, 0].legend()
xs = rng.normal(0, 1, 60)
ys = 0.7 * xs + rng.normal(0, 0.3, 60)
axes[0, 1].scatter(xs, ys, s=12, alpha=0.7)
axes[0, 1].set_xlabel("Variable X")
axes[0, 1].set_ylabel("Variable Y")
axes[1, 0].bar(["A", "B", "C", "D"], [3.2, 5.8, 4.1, 6.5])
axes[1, 0].set_xlabel("Category")
axes[1, 0].set_ylabel("Count")
axes[1, 1].hist(rng.normal(0, 1, 500), bins=25, edgecolor="white", linewidth=0.5)
axes[1, 1].set_xlabel("Value")
axes[1, 1].set_ylabel("Frequency")
style.savefig(fig, "multi_panel.pdf")
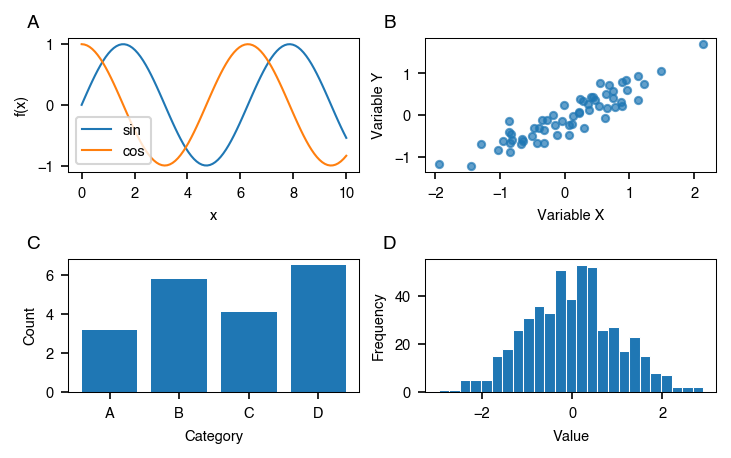
axesis always a 2-D NumPy array. Useaxes[0, 0]to access a single panel oraxes.flatto iterate. Passpanels=Falseto suppress the automatic labels.
Color palettes
Each journal has a recommended colorblind-safe palette. plotstyle.palette() returns hex color strings, cycling if you need more than the palette length.
import matplotlib.pyplot as plt
import plotstyle
journals = ["nature", "science", "ieee", "acs"]
fig, axes = plt.subplots(len(journals), 1, figsize=(6, 0.6 * len(journals)))
for ax, journal in zip(axes, journals, strict=False):
pal = plotstyle.palette(journal, n=8)
for i, color in enumerate(pal):
ax.barh(0, 1, left=i, color=color, edgecolor="none", height=0.8)
ax.set_xlim(0, 8)
ax.set_yticks([])
ax.set_ylabel(journal, rotation=0, ha="right", va="center")
ax.set_xticks([])
fig.suptitle("Journal Color Palettes")
fig.tight_layout()
fig.savefig("palette_comparison.png", dpi=150)
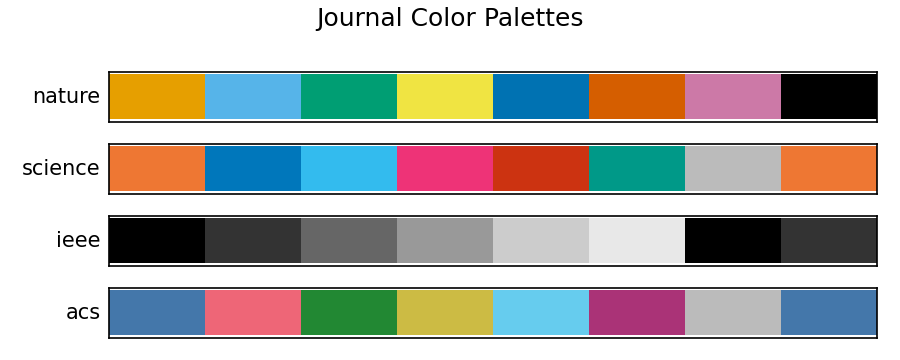
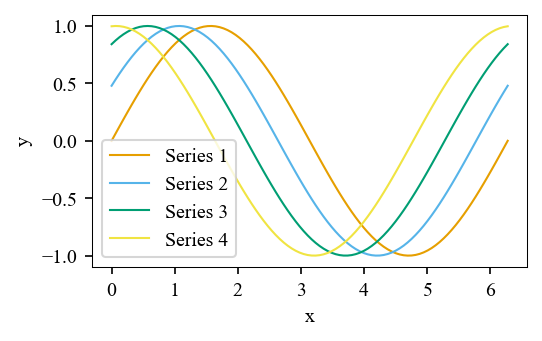
Pass with_markers=True to get (color, linestyle, marker) tuples, useful for journals like IEEE that print in grayscale:
import numpy as np
import plotstyle
x = np.linspace(0, 2 * np.pi, 100)
curves = [np.sin(x + i * 0.5) for i in range(4)]
with plotstyle.use("ieee") as style:
fig, ax = style.figure(columns=1)
styled = plotstyle.palette("ieee", n=4, with_markers=True)
for i, (color, ls, marker) in enumerate(styled):
ax.plot(x, curves[i], color=color, linestyle=ls, marker=marker,
markevery=20, label=f"Series {i + 1}")
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend()
style.savefig(fig, "ieee_markers.pdf")
# styled: one (color, linestyle, marker) tuple per series:
[('#000000', '-', 'o'), ('#333333', '--', 's'), ('#666666', '-.', '^'), ('#999999', ':', 'D')]
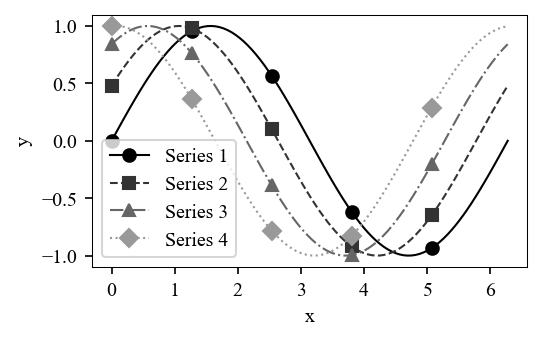
Overlays
Overlays are additive patches that layer on top of a journal preset. They let you adjust one aspect of a figure (the colour palette, the context, the chart type) without changing the base journal settings.
Pass overlay names in the same list as the journal key:
import matplotlib.pyplot as plt
import plotstyle
# Strip top/right spines for a clean editorial look
with plotstyle.use(["nature", "minimal"]) as style:
fig, ax = style.figure(columns=1)
ax.plot([1, 2, 3], label="data")
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend()
style.savefig(fig, "minimal_figure.pdf")
import matplotlib.pyplot as plt
import plotstyle
# Larger figure and fonts for Jupyter notebooks
with plotstyle.use(["nature", "notebook"]) as style:
fig, ax = plt.subplots() # plt.subplots() picks up the notebook figsize
ax.plot([1, 2, 3], label="data")
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend()
style.savefig(fig, "notebook_figure.pdf")
import matplotlib.pyplot as plt
import plotstyle
# Swap the colour cycle to a specific palette
with plotstyle.use(["ieee", "okabe-ito"]) as style:
fig, ax = style.figure(columns=1)
ax.plot([1, 2, 3], label="data")
ax.set_xlabel("x")
ax.set_ylabel("y")
ax.legend()
style.savefig(fig, "okabe_ito_figure.pdf")
| Category | Purpose | Examples |
|---|---|---|
color |
Swap the colour cycle | okabe-ito, tol-bright, safe-grayscale |
context |
Adjust scale for the medium | notebook, presentation, minimal, high-vis |
rendering |
Control LaTeX and grid rendering | no-latex, grid, latex-sans, pgf |
plot-type |
Optimise for a chart type | bar, scatter |
script |
Non-Latin font support | cjk-simplified, russian, turkish |
# List all available overlays
plotstyle.list_overlays()
plotstyle.list_overlays(category="context")
# plotstyle.list_overlays()
['bar', 'cjk-japanese', 'cjk-korean', 'cjk-simplified', 'cjk-traditional', 'grid',
'high-vis', 'latex-sans', 'minimal', 'no-latex', 'notebook', 'okabe-ito', 'pgf',
'presentation', 'russian', 'safe-grayscale', 'scatter', 'tol-bright',
'tol-high-contrast', 'tol-light', 'tol-muted', 'tol-rainbow-10', 'tol-rainbow-12',
'tol-rainbow-4', 'tol-rainbow-6', 'tol-rainbow-8', 'tol-vibrant', 'turkish']
# plotstyle.list_overlays(category="context")
['high-vis', 'minimal', 'notebook', 'presentation']
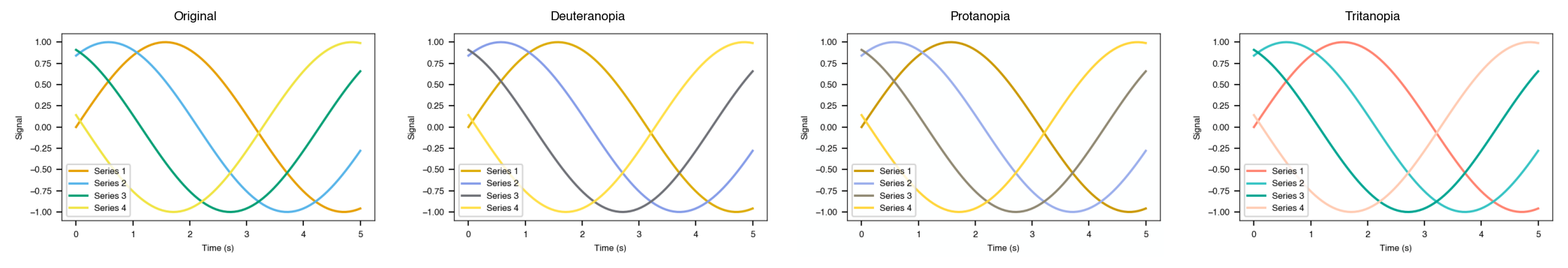
Colorblind and grayscale previews
Build a figure, then simulate how it looks under color vision deficiency or grayscale printing before you submit.
import numpy as np
import plotstyle
with plotstyle.use("nature") as style:
colors = style.palette(n=4)
fig, ax = style.figure(columns=1)
x = np.linspace(0, 5, 80)
for i, c in enumerate(colors):
ax.plot(x, np.sin(x + i), color=c, linewidth=1.5, label=f"Series {i + 1}")
ax.set_xlabel("Time (s)")
ax.set_ylabel("Signal")
ax.legend()
# Simulate colour vision deficiency (deuteranopia, protanopia, tritanopia)
cvd_fig = plotstyle.preview_colorblind(fig)
cvd_fig.savefig("accessibility_colorblind.png", dpi=150, bbox_inches="tight")
# Simulate grayscale print
gray_fig = plotstyle.preview_grayscale(fig)
gray_fig.savefig("accessibility_grayscale.png", dpi=150, bbox_inches="tight")
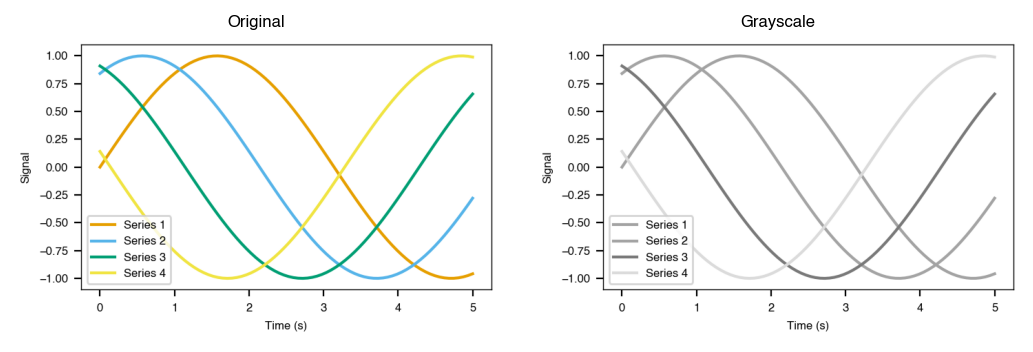
Validation and submission export
Validate a figure against the journal's requirements, then export in all required formats at once.
import numpy as np
import matplotlib.pyplot as plt
import plotstyle
x = np.linspace(0, 2 * np.pi, 100)
# Outside plotstyle.use() — some checks will fail
fig, ax = plt.subplots()
ax.plot(x, np.sin(x), label="sin(x)")
ax.set_xlabel("Phase (rad)")
ax.set_ylabel("Amplitude")
ax.legend()
report = plotstyle.validate(fig, journal="nature")
print(report) # formatted compliance table
print(report.passed) # False — rcParams not configured
for failure in report.failures:
print(failure.message) # what failed
print(failure.fix_suggestion) # how to fix it
plt.close(fig)
# Inside plotstyle.use() — all checks pass
with plotstyle.use("nature") as style:
fig, ax = style.figure(columns=1)
ax.plot(x, np.sin(x), label="sin(x)")
ax.set_xlabel("Phase (rad)")
ax.set_ylabel("Amplitude")
ax.legend()
report = plotstyle.validate(fig, journal="nature")
print(report)
print(report.passed) # True
┌──────────────────────────────────────────────────────┐
│ PlotStyle Validation Report: Nature │
├──────────┬───────────────────────────────────────────┤
│ ✓ PASS │ Figure width 89.0mm matches single colu...│
│ ✓ PASS │ Figure height 55.0mm is within the Natu...│
│ ✗ FAIL │ pdf.fonttype = 3; must be 42 for TrueTy...│
│ ✗ FAIL │ ps.fonttype = 3; must be 42 for TrueTyp...│
│ ⚠ WARN │ savefig.dpi = 'figure'; Nature requires...│
│ ✓ PASS │ All plotted lines and spines meet the N...│
│ ✗ FAIL │ 14 text element(s) outside the Nature r...│
└──────────┴───────────────────────────────────────────┘
3/7 checks passed, 1 warning(s), 3 failure(s)
passed: False
failure.message: pdf.fonttype = 3; must be 42 for TrueType font embedding.
failure.fix_suggestion: Call plotstyle.use() to apply all required rcParams, or set
mpl.rcParams['pdf.fonttype'] = 42 manually.
When called inside plotstyle.use(), all checks pass:
┌──────────────────────────────────────────────────────┐
│ PlotStyle Validation Report: Nature │
├──────────┬───────────────────────────────────────────┤
│ ✓ PASS │ Figure width 89.0mm matches single colu...│
│ ✓ PASS │ Figure height 55.0mm is within the Natu...│
│ ✓ PASS │ pdf.fonttype = 42 (TrueType fonts will ...│
│ ✓ PASS │ ps.fonttype = 42 (TrueType fonts will b...│
│ ✓ PASS │ savefig.dpi = 300.0 meets the Nature mi...│
│ ✓ PASS │ All plotted lines and spines meet the N...│
│ ✓ PASS │ All text elements are within the Nature...│
└──────────┴───────────────────────────────────────────┘
7/7 checks passed, 0 warning(s), 0 failure(s)
passed: True
import numpy as np
import os
import plotstyle
os.makedirs("submission", exist_ok=True)
with plotstyle.use("ieee") as style:
fig, ax = style.figure(columns=1)
x = np.linspace(0, 2 * np.pi, 100)
ax.plot(x, np.sin(x), label="sin(x)")
ax.set_xlabel("Phase (rad)")
ax.set_ylabel("Amplitude")
ax.legend()
paths = plotstyle.export_submission(
fig,
"figure1",
journal="ieee",
author_surname="Smith", # IEEE prepends the surname prefix to filenames
output_dir="submission/",
)
print(paths)
[PosixPath('submission/smith_figure1.tiff'),
PosixPath('submission/smith_figure1.eps'),
PosixPath('submission/smith_figure1.pdf'),
PosixPath('submission/smith_figure1.png')]
Supported Journals
| Key | Journal | Publisher |
|---|---|---|
acs |
ACS (JACS) | American Chemical Society |
cell |
Cell | Cell Press |
elsevier |
Elsevier | Elsevier |
ieee |
IEEE Transactions | IEEE |
nature |
Nature | Springer Nature |
plos |
PLOS ONE | Public Library of Science |
prl |
Physical Review Letters | American Physical Society |
science |
Science | AAAS |
springer |
Springer | Springer |
wiley |
Wiley | Wiley |
Need another journal? See CONTRIBUTING.md.
CLI
plotstyle list # list all journal presets
plotstyle info <journal> # show spec details
plotstyle diff <journal_a> <journal_b> # compare two journals
plotstyle fonts --journal <journal> # check font availability
plotstyle overlays [--category <category>] # list available overlays
plotstyle overlay-info <overlay> # show overlay details
plotstyle validate <file> --journal <journal> # validate a saved figure
plotstyle export <file> --journal <journal> # print snippet for re-exporting
plotstyle list
acs American Chemical Society
cell Cell Press
elsevier Elsevier
ieee IEEE
nature Springer Nature
plos Public Library of Science
prl American Physical Society
science AAAS
springer Springer Nature
wiley Wiley
plotstyle info nature
Journal: Nature
Publisher: Springer Nature
Source: https://www.nature.com/documents/nature-final-artwork.pdf
Last Verified: 2026-04-22
──────────────────────────
Dimensions:
Single column: 89.0mm (3.50in)
Double column: 183.0mm (7.20in)
Max height: 247.0mm
Typography:
Font: Helvetica, Arial (fallback: sans-serif)
Size range: 5.0-7.0pt
Panel labels: 5.0pt bold lower (a, b, c)
Export:
Formats: ai, eps, pdf
Min DPI: 300
Color: rgb
Accessibility:
Colorblind safe: Not required
Grayscale safe: Not required
plotstyle diff nature science
Nature → Science
──────────────────────────────────────────────────
Column Width (single): 89.0mm → 86.4mm
Column Width (double): 183.0mm → 177.8mm
Max Height: 247.0mm → -
Font Family: Helvetica, Arial → Minion Pro, Benton Sans Condensed
Min Font Size: 5.0pt → 7.5pt
Max Font Size: 7.0pt → 10.0pt
Panel Label Size: 5.0pt → 7.5pt
Preferred Formats: ai, eps, pdf → ai, eps, pdf, tiff
Colorblind Required: No → Yes
plotstyle fonts --journal nature
Font check for: Nature
Required: Helvetica, Arial
Available: Helvetica, Arial
Selected: Helvetica
Exact match: Yes
plotstyle overlays
bar [plot-type] Optimised rcParams for bar charts.
cjk-simplified [script] Font configuration for Simplified Chinese labels.
grid [rendering] Enable major grid lines with a subtle dashed style.
high-vis [context] Maximum contrast, bold lines, and oversized ticks.
latex-sans [rendering] Enable LaTeX rendering with a sans-serif font family.
minimal [context] Stripped-down axes with no top/right spines.
no-latex [rendering] Disable LaTeX text rendering; use Matplotlib MathText.
notebook [context] Enlarged figures and larger fonts for Jupyter.
okabe-ito [color] Colorblind-safe 8-color qualitative palette.
pgf [rendering] Use the PGF LaTeX backend for vector output.
presentation [context] Large text and thick lines for slide decks.
safe-grayscale [color] 6-step grayscale palette for black-and-white print.
scatter [plot-type] Optimised rcParams for scatter plots.
tol-bright [color] Paul Tol's bright 7-color qualitative palette.
...
plotstyle overlay-info minimal
Overlay: Minimal
Key: minimal
Category: context
Description: Stripped-down axes with no top/right spines for editorial and blog use.
──────────────────────────
rcParams:
axes.spines.top = False
axes.spines.right = False
xtick.top = False
ytick.right = False
axes.grid = False
axes.linewidth = 0.8
Documentation
Full documentation at plotstyle.readthedocs.io:
Working examples are in the examples/ directory.
Contributing
See CONTRIBUTING.md for development setup, adding journal specs, and pull request guidelines.
Citation
If PlotStyle helps your research, a citation or star is appreciated:
@misc{plotstyle,
author = {Kaushal, Rahul},
title = {PlotStyle: Publication-ready scientific figure presets for Matplotlib},
year = {2026},
url = {https://github.com/rahulkaushal04/plotstyle},
note = {Version 1.2.1},
}
License
MIT © 2026 Rahul Kaushal
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file plotstyle-1.2.1.tar.gz.
File metadata
- Download URL: plotstyle-1.2.1.tar.gz
- Upload date:
- Size: 218.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1c3cfdc89eb383b79acc5107f0df3095b2ca88d6a916412555632af6a3cc9aaf
|
|
| MD5 |
6b45ecc32d6c399279dc84c9bde3e37c
|
|
| BLAKE2b-256 |
be6bf2d2fddf4b7542c3a0a2c54fdfd38123c3162b3093f60b33453b799c0116
|
Provenance
The following attestation bundles were made for plotstyle-1.2.1.tar.gz:
Publisher:
release.yml on rahulkaushal04/plotstyle
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
plotstyle-1.2.1.tar.gz -
Subject digest:
1c3cfdc89eb383b79acc5107f0df3095b2ca88d6a916412555632af6a3cc9aaf - Sigstore transparency entry: 1389896780
- Sigstore integration time:
-
Permalink:
rahulkaushal04/plotstyle@dab72dd04b4df2a58cfd29b0c4dfdb7aec5666c5 -
Branch / Tag:
refs/tags/v1.2.1 - Owner: https://github.com/rahulkaushal04
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@dab72dd04b4df2a58cfd29b0c4dfdb7aec5666c5 -
Trigger Event:
push
-
Statement type:
File details
Details for the file plotstyle-1.2.1-py3-none-any.whl.
File metadata
- Download URL: plotstyle-1.2.1-py3-none-any.whl
- Upload date:
- Size: 116.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6b134a3ef4024332125938775fd3bb2164be472db1311d967d3337efbd4f6874
|
|
| MD5 |
20eb891ac9f18ba9755393ce4a6814d7
|
|
| BLAKE2b-256 |
eecdb0aff5cd4e0e1e4bac101aa7bfece4d3567a67268d1654bf8d55571c2675
|
Provenance
The following attestation bundles were made for plotstyle-1.2.1-py3-none-any.whl:
Publisher:
release.yml on rahulkaushal04/plotstyle
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
plotstyle-1.2.1-py3-none-any.whl -
Subject digest:
6b134a3ef4024332125938775fd3bb2164be472db1311d967d3337efbd4f6874 - Sigstore transparency entry: 1389896929
- Sigstore integration time:
-
Permalink:
rahulkaushal04/plotstyle@dab72dd04b4df2a58cfd29b0c4dfdb7aec5666c5 -
Branch / Tag:
refs/tags/v1.2.1 - Owner: https://github.com/rahulkaushal04
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yml@dab72dd04b4df2a58cfd29b0c4dfdb7aec5666c5 -
Trigger Event:
push
-
Statement type:
















