Blazing-fast simulation of self-organized patterns in reaction-diffusion systems.
Project description
PySpecies
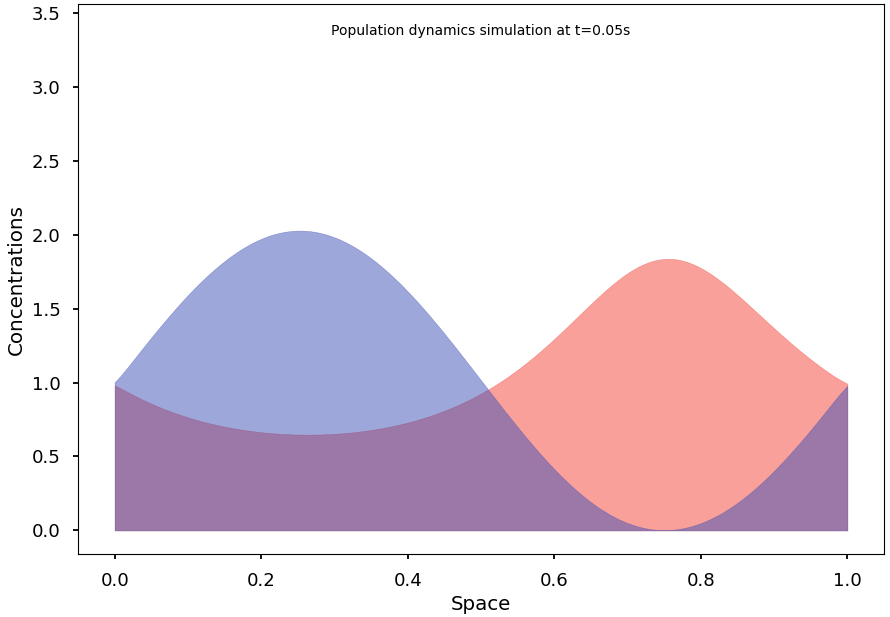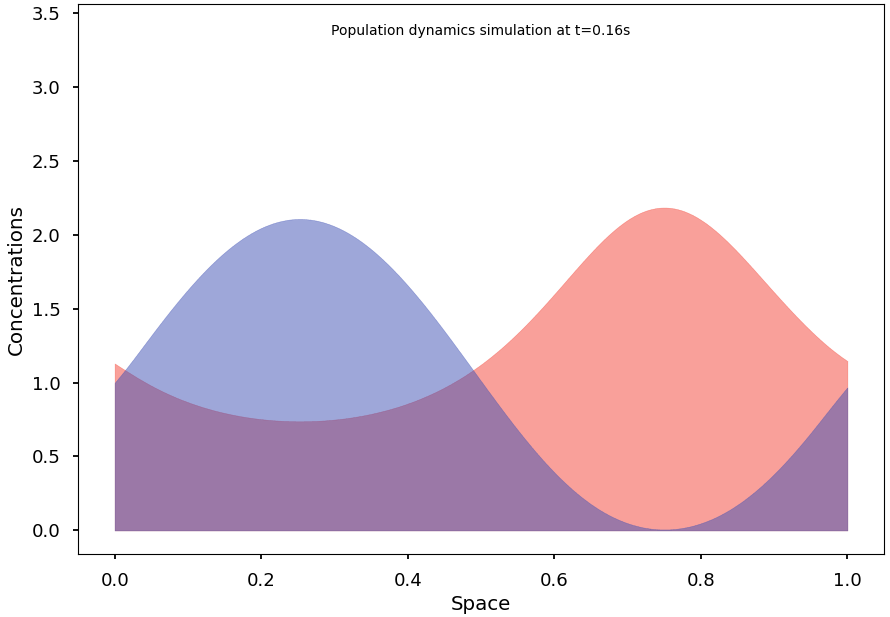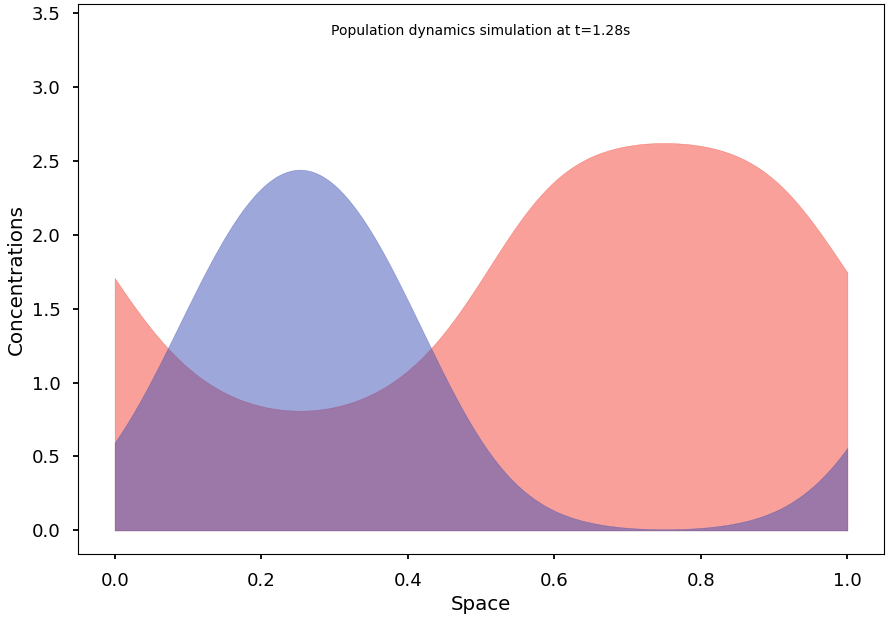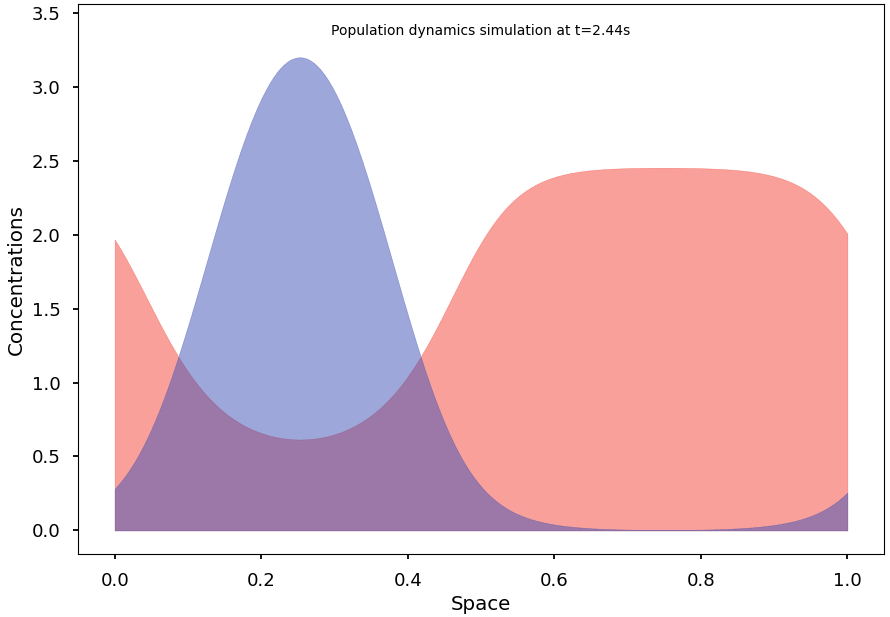
Blazing-fast simulation of population dynamics, based on the Shigesada-Kawasaki-Teramoto (SKT) reaction-diffusion model. [PubMed '79]
Installation
pip install pyspecies
Usage
For example, the following code computes a solution of the SKT model and converges to a non-homogeneous steady state:
import numpy as np
from pyspecies import models, pop
# Define population and interaction model
q = pop.Pop(
space=(0, 1, 200),
u0=lambda x: 1 + np.cos(2 * np.pi * x),
v0=lambda x: 1 + np.sin(2 * np.pi * x),
model=models.SKT(
D=np.array([[5e-3, 0, 3], [5e-3, 0, 0]]),
R=np.array([[5, 3, 1], [2, 1, 3]])
),
)
# Simulate with increasing speeds
for i in range(-2, 2):
q.sim(duration=2*10**i, N=100)
# Animate the result
q.anim()
# Show the evolution of the population over space and time
# q.heatmap()
# Show the final state of the population (100%)
# q.snapshot(1)
This code displays a cyclic, homogenous solution of the Lotka-Volterra equations:
p = pop.Pop(
space = (0, 1, 10),
u0 = lambda x: 1 + 0*x, # IC for prey
v0 = lambda x: 1 + 0*x, # IC for predator
model = models.LV(1.1, 0.4, 0.4, 0.1)
)
p.sim(duration=20, N=200)
p.sim(duration=100, N=200)
p.anim()
Theory
The calculations underlying this library are described (in French) in the following document.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file PySpecies-0.1.2.tar.gz.
File metadata
- Download URL: PySpecies-0.1.2.tar.gz
- Upload date:
- Size: 7.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.10.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c95c49aa6fbbcee77d053eaa27a3125400405c48b360dd56b68c044119a0878d
|
|
| MD5 |
ce0c96b951610fba52ba5577b35111e7
|
|
| BLAKE2b-256 |
500e11fd99d6cd1f85a7dc993c5d3e533c5d2ff7bce9636f32cdfd7fcc74f533
|
File details
Details for the file PySpecies-0.1.2-py3-none-any.whl.
File metadata
- Download URL: PySpecies-0.1.2-py3-none-any.whl
- Upload date:
- Size: 9.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.10.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2b12dc3d41de8848989931c8db538ba5086ca4db4835ecd283ab4d72cbb0dd62
|
|
| MD5 |
e67cf1a897f808d073b85d7cfe553e01
|
|
| BLAKE2b-256 |
0e17d2de642c1a811c2ffcf0656e65b5d92e508dc840795ada3ec86690d6f51e
|