RBP Activity Inference from Splicing Events
Project description
RAISE
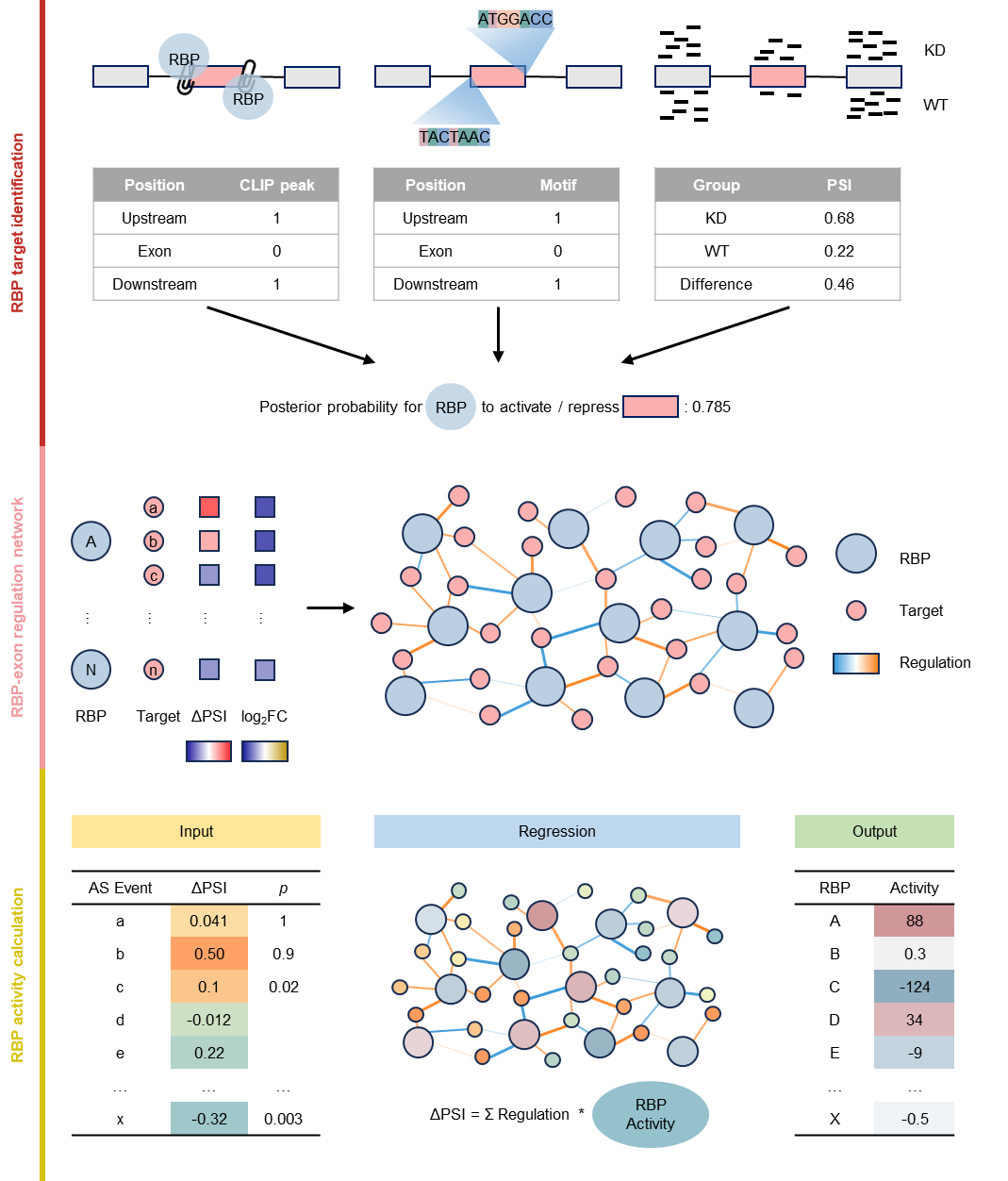
RBP Activity Inference from Splicing Events
RAISE is a computational pipeline for identifying the activity of RNA-binding proteins (RBPs). It integrates CLIP-seq peaks, motifs, and alternative splicing (AS) data to construct a splicing regulatory network, and infers RBP activities using regression modeling.
Installation
Option 1. Install RAISE through pip [recommended]
conda create -n RAISE python=3.8
pip install RAISE
Option 2. Local installation
conda create -n RAISE python=3.7
git clone https://github.com/liuyilei8969/RAISE.git
cd RAISE
pip install .
Usage
1. Identify targets of an RBP
usage: find_target.py [-h] --rmats RMATS --clip_peaks CLIP_PEAKS --ref_genome REF_GENOME --rbp_motif RBP_MOTIF --cell_line CELL_LINE --rbp RBP --output
OUTPUT [--max_iter MAX_ITER] [--tol TOL]
EM algorithm for inferring RBP targets using motif, CLIP peaks, and PSI changes.
options:
-h, --help show this help message and exit
--rmats RMATS Input rMATS SE.MATS.JC.txt file.
--clip_peaks CLIP_PEAKS Input CLIP peaks BED file.
--ref_genome REF_GENOME Reference genome in FASTA format.
--rbp_motif RBP_MOTIF RBP motif file with two columns: RBP and motif.
--cell_line CELL_LINE Cell line name, used to label the output file.
--rbp RBP Target RBP name.
--output OUTPUT Output directory.
--max_iter MAX_ITER Maximum number of EM iterations.
--tol TOL Convergence threshold for EM.
--use_motif Use motif features in EM.
--use_clip Use clip features in EM.
2. Construct RBP-AS network
usage: construct_network.py [-h] --Target_dir TARGET_DIR [--threshold THRESHOLD] --DE_dir DE_DIR --output OUTPUT
Build a splicing regulatory network from target predictions and RBP expression changes.
options:
-h, --help show this help message and exit
--Target_dir TARGET_DIR Directory containing RBP target result folders
--threshold THRESHOLD Minimum conditional probability P(T|S,M,C) to include interaction (default: 0.6)
--DE_dir DE_DIR Directory containing RBP expression change files
--output OUTPUT Path to output GEXF file for the constructed network
3. Infer RBP activity
usage: calculate_activity.py [-h] --diffAS DIFFAS --network NETWORK --output OUTPUT
Infer RBP activity from a splicing regulatory network using ridge regression.
options:
-h, --help show this help message and exit
--diffAS DIFFAS Path to the rMATS differential splicing results file
--network NETWORK Path to the splicing regulatory network
--output OUTPUT Output file for inferred RBP activity scores
Example & Test
Examples are provided in the test/ directory: https://github.com/liuyilei8969/RAISE/tree/main/test
Data are provided in the data/ directory for users' convenience: https://github.com/liuyilei8969/RAISE/tree/main/data
Note: All differential splicing results should be provided in the rMATS format. For users' convenience, we also provide scripts to either convert data into this format or perform a simple differential splicing analysis using a limma test.
Requirements
Python >= 3.8
Packages: pandas, numpy, networkx, scikit-learn, argparse
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file raise_rbp-0.1.6.tar.gz.
File metadata
- Download URL: raise_rbp-0.1.6.tar.gz
- Upload date:
- Size: 3.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
453d7065b1a2cef3722e7abfa113db764771bcd9c8b43831c4841784dc4ec655
|
|
| MD5 |
baf308723fcf2330ad8bf0ab8475dd80
|
|
| BLAKE2b-256 |
6d1f6868ead9fc070d061116c6b0b5380617fb23ea4d1ea85903a4ee9e119ba8
|
File details
Details for the file raise_rbp-0.1.6-py3-none-any.whl.
File metadata
- Download URL: raise_rbp-0.1.6-py3-none-any.whl
- Upload date:
- Size: 27.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b7e1e001eac149f88c34135b6eb50f02a6eab3a75d094f254f7da56343a1d7c7
|
|
| MD5 |
6828f4d3bf6bd323807ffe937ad0fb94
|
|
| BLAKE2b-256 |
ce6d044863f5c27f4916ef12ad0bd967ca806e488d169cf081380b76ac510348
|