scDREAMER is a single-cell data integration framework that employs a novel adversarial variational autoencoder for learning lower-dimensional cellular embeddings and a batch classifier neural network for the removal of batch effects.
Project description
scDREAMER
Overview
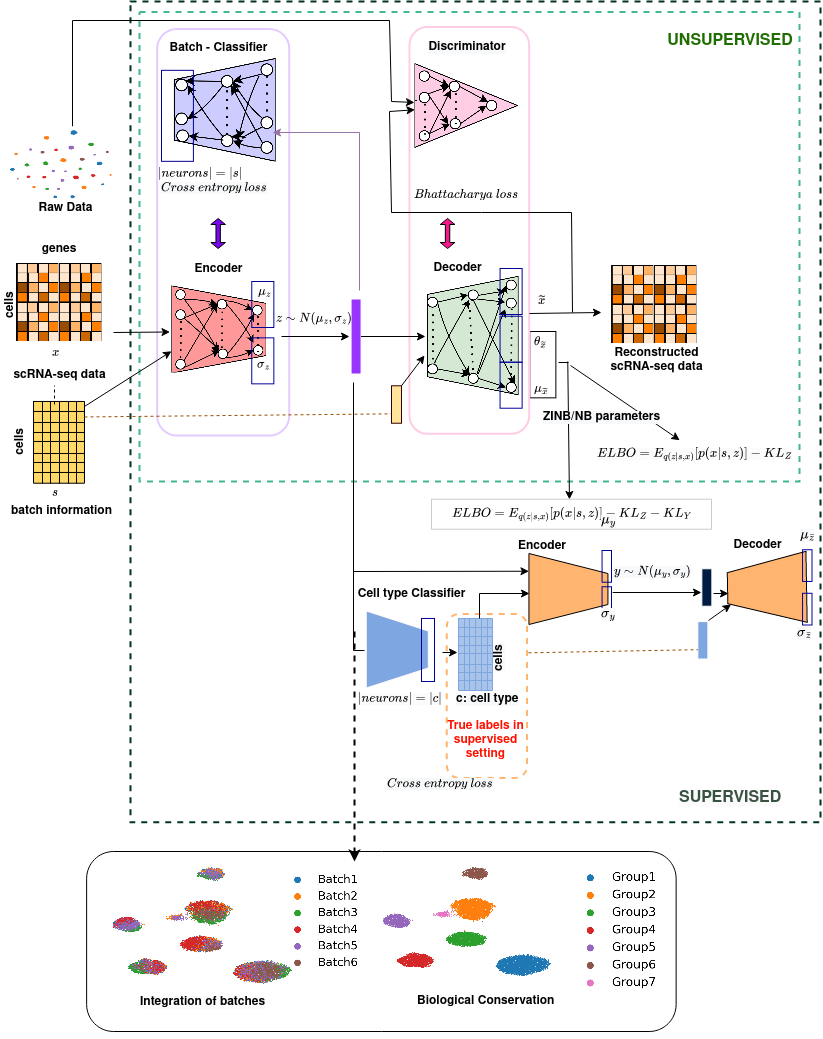
scDREAMER is a single-cell data integration framework that employs a novel adversarial variational autoencoder for learning lower-dimensional cellular embeddings and a batch classifier neural network for the removal of batch effects. The jupyter notebooks for reproducibility of results in the manuscript are available at https://github.com/Zafar-Lab/scDREAMER-reproducibility. See our paper below for more details. DOI: https://doi.org/10.1038/s41467-023-43590-8
Installation
A stable pip installation release for scDREAMER package will be made available shortly. For now, we recommend users to directly clone our stable main branch and set scDREAMER as the working directory. Creating conda environment using ./ENVIRONMENTS/scDREAMER.yml will install all the dependent packages and libraries. scDREAMER can be set up as follows
git clone https://github.com/Zafar-Lab/scDREAMER.git
cd scDREAMER/Environments
conda env create -f scDREAMER.yml
conda activate scdreamer
What Computational tasks can scDREAMER be used for?
scDREAMER suite can be used for:
- scDREAMER for an unsupervised integration of multiple batches
- scDREAMER-SUP for a supervised integration across multiple batches
- scDREAMER-SUP can also be when cell type annotations are missing in the datasets i.e., 10%, 20%, 50%
- Atlas level and cross-species integration
- Large datasets with ~1 million cells
Tutorials
Check out the following Colab notebook to get a flavor of a typical workflow for data integration using scDREAMER and scDREAMER-SUP (Link to Datasets) below.
- scDREAMER applied to human immune integration task
- scDREMER-SUP applied to human immune cells integration task
- scDREAMER-SUP applied to human immune cells integration task under 50% missing cell labels setting
Documentation
Read the docs: https://scdreamer.readthedocs.io/en/latest/
Contributing
In case of any bug reports, enhancement requests, general questions, and other contributions, please create an issue. For more substantial contributions, please fork this repo, push your changes to your fork, and submit a pull request with a good commit message.
Cite this article
Ajita Shree*, Musale Krushna Pavan*, Hamim Zafar. scDREAMER: atlas-level integration of single-cell datasets using deep generative model paired with adversarial classifier. bioRxiv 2022.07.12.499846; doi: https://doi.org/10.1038/s41467-023-43590-8
* equally contributed
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file scdreamer-0.0.0rc1.tar.gz.
File metadata
- Download URL: scdreamer-0.0.0rc1.tar.gz
- Upload date:
- Size: 4.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.9.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0a6455c1ff51140a4c8f78e93f3974572e61419e02c8e2e661db1913eac1d920
|
|
| MD5 |
41b801512189c41b14ff05b72a11feb5
|
|
| BLAKE2b-256 |
dfb5f51dd3dc214f26b7c24e3ce062545317756cf5934dad11d3382c547d685a
|
File details
Details for the file scdreamer-0.0.0rc1-py3-none-any.whl.
File metadata
- Download URL: scdreamer-0.0.0rc1-py3-none-any.whl
- Upload date:
- Size: 19.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.9.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d085a691124d737abb99e231f50182514facfe7928310da40550a9b33c7be898
|
|
| MD5 |
c6491b0c8afeacb70ee0adee4e4f4d0e
|
|
| BLAKE2b-256 |
662bf221d395c45698b45c75b181183e3b4219ea79617c70f7ad37adffd4bba0
|












