A novel spatially aware multi-omics dimension reduction method
Project description
Spatial Multi-Omics PCA
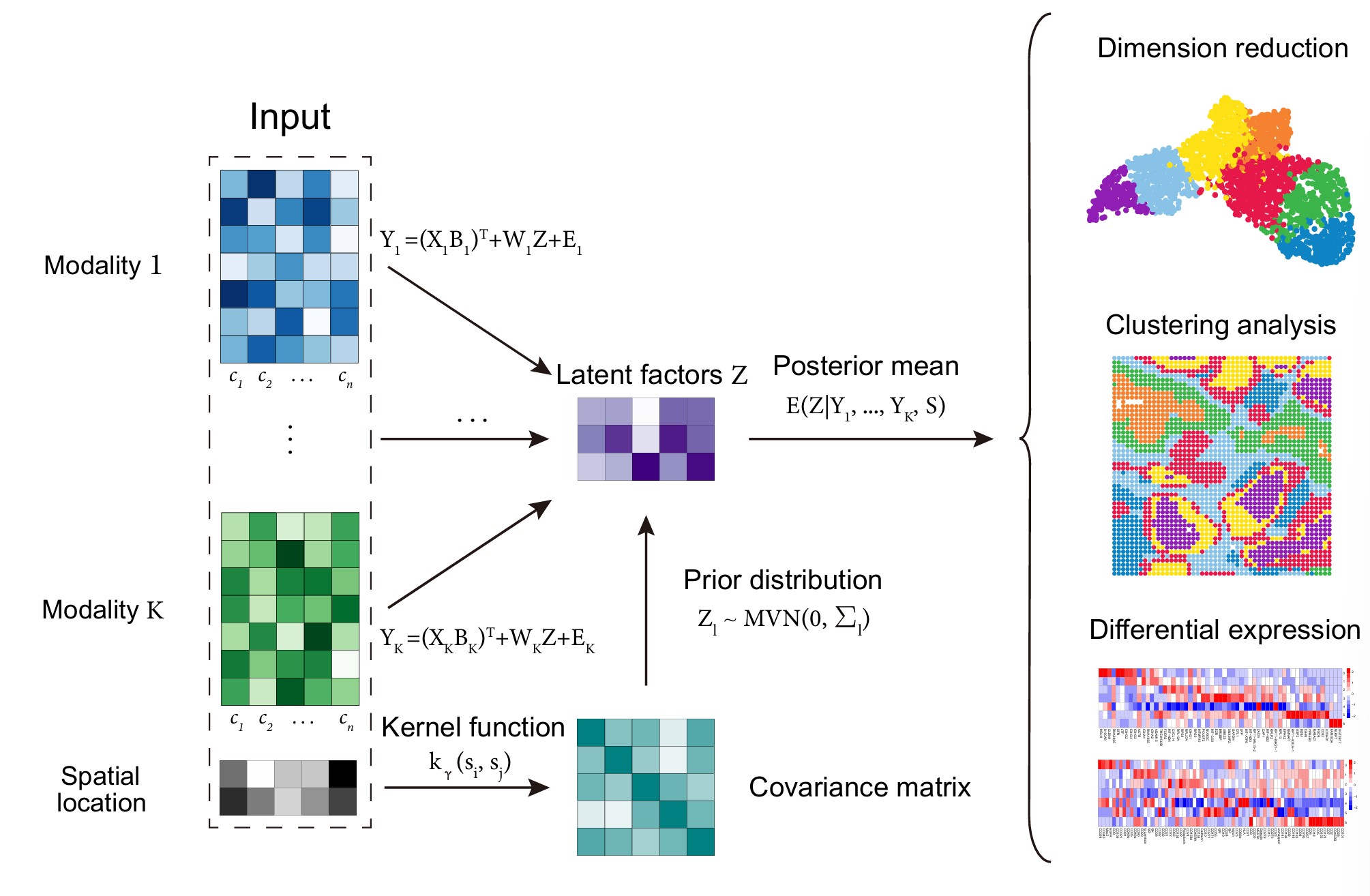
Spatial Multi-Omics PCA (SMOPCA) is a novel dimension reduction method to integrate multi-modal data and extract low-dimensional representations with preserved spatial dependencies among spots.
Installation
SMOPCA can be installed directly from PyPI using the following command:
pip install SMOPCA
Run SMOPCA
- Prepare input data
- SMOPCA accepts gene expression and protein/atac data matrices. Each modality of the data is preprocessed and normalized separately. Initially, genes and proteins with zero counts were filtered out. Subsequently, the count matrix was normalized based on library size, followed by log-transformation and scaling to achieve unit variance and zero mean. ATAC reads are mapped to gene regions and the peak matrix is collapsed into a gene activity matrix, adhering to the established protocol from the Satija lab. The gene activity matrix was preprocessed and normalized using the same method as applied to mRNA data. Finally, we recommend to save the data into a hdf5 file.
- Note that SMOPCA takes input matrices with columns corresponding to cells or spots.
- Specify model hyperparameters and Model training
- the dimensionality of the latent factors (default 20)
- the kernel type (default matern kernel)
- the length_scale parameter (We set length_scale=0.25 for UMAP coordinates and length_scale=5 for simulated and real spatial coordinates.)
- For the rest of the parameters, see more in tutorials.
- Downstream analysis
- Visualization
- Clustering analysis
- Differential expression analysis
- and many other tasks
Datasets
Sample datasets are provided in ./data folder. The rest of the datasets used in this study are available at https://drive.google.com/drive/folders/11RfeF_yrdSGRtXZnzPlfxojM5V4ezMVr?usp=drive_link
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file smopca-0.1.1.tar.gz.
File metadata
- Download URL: smopca-0.1.1.tar.gz
- Upload date:
- Size: 8.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.10.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9d2bccc1b56f56e36117088dacdd06111488069c76af1925159db29483e5a455
|
|
| MD5 |
11ce9eee50bc2b005d20b1e19ac03d03
|
|
| BLAKE2b-256 |
43756e528d295233f825ab43e97214241df70441c49c1fa8d3ef12a3bb031abe
|
File details
Details for the file SMOPCA-0.1.1-py3-none-any.whl.
File metadata
- Download URL: SMOPCA-0.1.1-py3-none-any.whl
- Upload date:
- Size: 8.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.10.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
84d98c44069053f0f88f566cc2b89afd053b6330d92f2c9ecc6cab37fd945e57
|
|
| MD5 |
6f878f0fda83cc0539434abd1bfb772f
|
|
| BLAKE2b-256 |
ae93f177b828f85ece7f62e94bf259a0c6b941828978b7d24ee012585814715a
|