Discovering geroprotectors through the explainable artificial intelligence-based platform AgeXtend
Project description
AgeXtend
Introduction
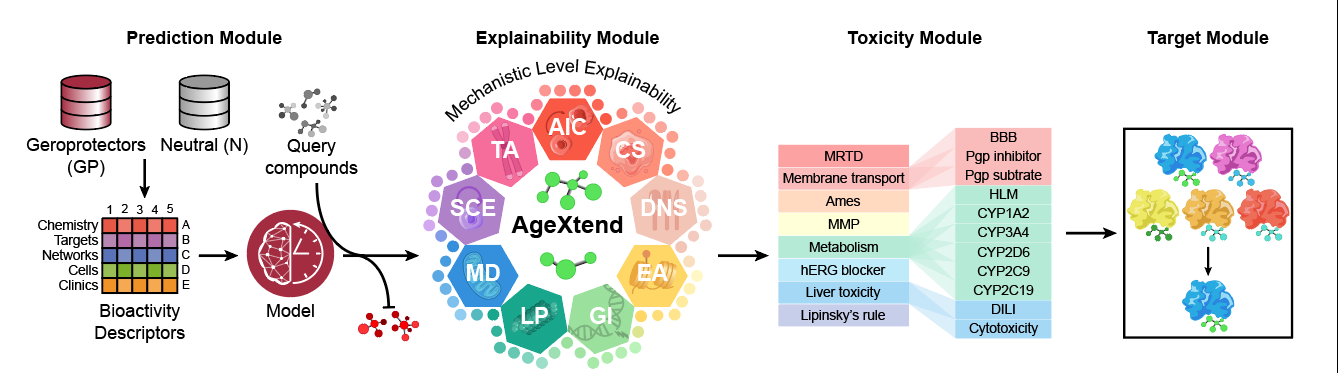
AgeXtend is a multimodal, bioactivity-based, and mechanism-backed Explainable AI Predictor for geroprotectors. AgeXtend supports 2 distinct work packages: Predictor and Browser. The first work package, Predictor, involves a bioactivity-based classifier for predicting geroprotective compounds, coupled with an explainability module providing mechanistic insights into biological processes behind the predictions, a toxicity module to evaluate their toxicity in the biological context, and finally, a target module that suggests protein targets of the putative geroprotective compounds. The second work package, Browser, allows the user to explore through the 20 pre-screened databases specifically curated for these functionalities.
License Key
AgeXtend is free for academic institutions, however, for commercial utilization a commercial license key is required. Users (academic/commercial) may apply for a valid "License Key" here.
You can also generate your own predictions using AgeXtend’s Colab notebook
Environment Setup (done using requirement.txt)
Major dependencies
Minor dependencies
- os
- scikit-learn (v1.2.1)
- xgboost (v1.5.1)
- pandas (v1.4.3)
- numpy (v>=1.20.3)
- tqdm
- joblib (v1.1.1)
- matplotlib (v>=3.2.2)
- seaborn (v0.11.2)
- importlib
- importlib-resources (v5.7.1)
Quick setup
The file requirement.txt will be used for the environment setup and by running the following command.
$ pip install -r requirement.txt
How to use AgeXtend?
Installation using pip
$ pip install -i https://pypi.org/simple/AgeXtend
License activation (One time)
To apply for the license click here
>>> from AgeXtend import Predictor
Activate AgeXtend license
>>> Predictor.license('license key') #Example: Predictor.license('KKKVFZ41111WF6RTQ')
Work Packages
AgeXtend supports 2 distinct work packages:
- Predictor
- Browser
Predictor
Prediction and Explainability Module
Predicts the anti-aging potential for the input SMILES:
>>> from AgeXtend import Predictor
Prepare a list of canonical SMILES (Open Babel generated) strings
>>> smiles = ['ClCC=C', 'C=CCOC(=O)CC(C)C']
Create an AgeXtend type object to featurize the query compounds (i.e. list of canonical SMILES)
>>> agex_obj = Predictor.featurize(smiles)
Use the AgeXtend object for predictions
>>> output = Predictor.predict(agex_obj)
Get the list of resulting data frames that are part of the rest of the modules
>>> output.keys()
dict_keys(['Anti_Aging_Prediction', 'Explainability_Status', 'Explainability_Probabilities'])
Note: Explainability_Probabilities will be empty by default unless selected otherwise by supplying additional arguments.
Get the result of a specific output data frame
>>> output['Explainability_Status']
Sub Explainability and Target Module
Additional arguments:
AgeXtend also supports the following modules along with the Prediction and Explainability modules, which can be chosen using the boolean option (True)
| Parameter Name | Description | Type | Default value | Output (If True) |
|---|---|---|---|---|
| probs | Probabilities of Explainability and/or Toxicity Module | boolean | False | Explainability_Probabilities / Explainability_Toxicity_Probabilities |
| HC | Run Toxicity Module (Health Check) | boolean | False | Explainability_Toxicity_Status |
| TS | Sub Explainability Level Tanimoto Similarity Test | boolean | False | Explainability_response |
| BDL | BindingDB Target Information and Druggability (Lipinski Rule) | boolean | False | Druggability_and_Potential_Targets |
Example
>>> output = Predictor.predict(agex_obj, TS=True)
>>> output.keys()
dict_keys(['Anti_Aging_Prediction', 'Explainability_Status', 'Explainability_Probabilities', 'Explainability_response'])
>>> output = Predictor.predict(agex_obj, HC=True, BDL=True)
>>> output.keys()
dict_keys(['Anti_Aging_Prediction', 'Explainability_Toxicity_Status', 'Explainability_Toxicity_Probabilities', 'Druggability_and_Potential_Targets'])
Efficient Bulk Prediction / Custom AgeXtend Database
Predictor module also provides functionalities for efficient bulk data prediction
>>> Predictor.bulk_predict(input=smiles_list)
Or user can also use pre-calculated AgeXtend type object as input
>>> agex_obj = Predictor.featurize(smiles_list)
>>> Predictor.bulk_predict(input=agex_obj)
Additional arguments:
| Parameter Name | Description | Type | Default value |
|---|---|---|---|
| outfolder | Output Database type folder name | string | AgeXtendDB |
| chunksize | Number of predictions input per iteration for faster job completion | Integer | 10000 |
Note The output folder of Bulk prediction function can be used as a Custom database for input in AgeXtend Browser module
Browser
To explore the AgeXtend pre-complied predictions of various databases:
>>> from AgeXtend import Browser
Use Open Babel to Generate Canonical SMILES format of the query compound as input
>>> Browser.search(query='OC(=O)CCCc1c[nH]c2c1cccc2', output='/path/to/output/folder/')
Unzip the AgeXtendBrowserOut.zip file to visualize/print the generated report (HTML format)
Note: the report file (AgeXtend_BrowserOut.html) must be in the same folder with the images folder
Additional arguments:
AgeXtend also supports the use of locally complied Predictor module outputs (Folder containing CSV format outputs)
| Parameter Name | Description | Default Database |
|---|---|---|
| path | Path to the AgeXtend pre-complied prediction database | gutMGene |
Example
>>> Browser.search(path='/path/to/Database/Folder/', query='OC(=O)CCCc1c[nH]c2c1cccc2', output='/path/to/output/folder/')
Pre-complied AgeXtend Prediction Databases
| DB Name | DB version | FTP size | FTP link |
|---|---|---|---|
| FOODB | Pre-release 1.0 | 63M | http://agextend.ahujalab.iiitd.edu.in:8080/FOODB/ |
| HMDB | v5.0 | 231M | http://agextend.ahujalab.iiitd.edu.in:8080/HMDB/ |
| IMPPAT | v2.0 | 4K | http://agextend.ahujalab.iiitd.edu.in:8080/IMPPAT/ |
| AfroDB | - | 756K | http://agextend.ahujalab.iiitd.edu.in:8080/AfroDB/ |
| AgingAtlas | v1.0 | 748K | http://agextend.ahujalab.iiitd.edu.in:8080/AgingAtlas/ |
| Chembridge | - | 788K | http://agextend.ahujalab.iiitd.edu.in:8080/Chembridge/ |
| ChemdivBBlocks | - | 39M | http://agextend.ahujalab.iiitd.edu.in:8080/ChemdivBBlocks/ |
| CMNPD | v1.0 | 20M | http://agextend.ahujalab.iiitd.edu.in:8080/CMNPD/ |
| DDPD | - | 4K | http://agextend.ahujalab.iiitd.edu.in:8080/DDPD/ |
| ECMDB | v2.0 | 4K | http://agextend.ahujalab.iiitd.edu.in:8080/ECMDB/ |
| RepoHub | release-3/24/2020 | 3.9M | http://agextend.ahujalab.iiitd.edu.in:8080/RepoHub/ |
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file agextend-0.0.2.tar.gz.
File metadata
- Download URL: agextend-0.0.2.tar.gz
- Upload date:
- Size: 87.2 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.8.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
376d2ac5b901aafd14c3e041c567569a215bbddb2660dec3072aba8c37688f6c
|
|
| MD5 |
7f7b0d80fc884562573a2cd0fde510da
|
|
| BLAKE2b-256 |
fad4f09b0bdfdfaa2dc415e61d2554c3d3840fde0544189eb8a6a96bf5457518
|
File details
Details for the file AgeXtend-0.0.2-py3-none-any.whl.
File metadata
- Download URL: AgeXtend-0.0.2-py3-none-any.whl
- Upload date:
- Size: 88.7 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.8.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
14b6100595289dfb7e552f0bb00a52131f93e0a29c08ac1645fe8196f3a2ef66
|
|
| MD5 |
e899b7b1b6ed7c50bd0ba7227ee52368
|
|
| BLAKE2b-256 |
304134a3d81283a30d1fd60079315669c60e353c563f7a11d1b3bb02e782f4b9
|