B cell repertoire analysis and lineage tracking in spatial omics
Project description

A Python package for B cell repertoire analysis and lineage tracking in spatial omics.
Background
In contrast to previously reported bulk or single-cell immune receptor sequencing technologies, the simultaneous acquisition of spatially resolved high-dimensional gene expression profiles and complex immune receptor sequences from the same tissue section presents unique challenges for bioinformatic analysis.
To systematically investigate the spatial organization and clonal dynamics of B cells in the tumor microenvironment, we developed BAITS (B cell repertoire Analysis and lIneage Tracking in Spatial omics), a comprehensive and adaptable computational framework for analyzing spatially resolved BCR sequencing data.
Introduction
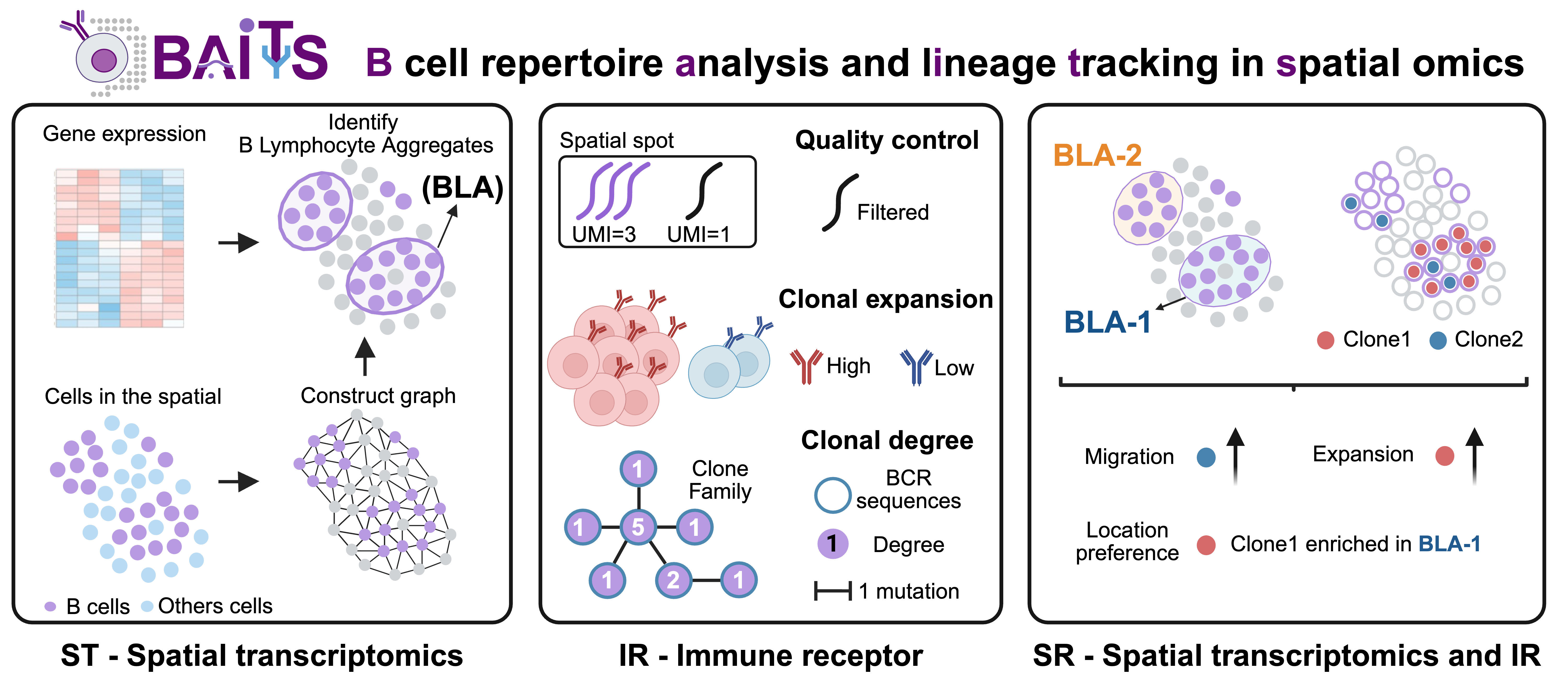
BAITS comprises three core modules:
-
Spatial Transcriptomics (ST) module: identify B lymphocyte aggregates based solely on spatial transcriptomic data
-
Immune Repertoire (IR) module: quantify clonal expansion, clonal degree centrality, and other repertoire features using spatial BCR sequencing data
-
SR module: reveal patterns of clonal migration, expansion, and niche restriction by integrating spatial transcriptomic and BCR data
Installation
1. Create a conda environment and then install Python >= 3.10,<3.13
conda create --name BAITS python=3.10
2. Pip install BAITS
conda activate BAITS
pip install BAITS
This example is based on a Linux CentOS 7 system
Contribution
If you found a bug or you want to propose a new feature, please use the issue tracker.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file baits-0.1.1.tar.gz.
File metadata
- Download URL: baits-0.1.1.tar.gz
- Upload date:
- Size: 4.7 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7482def54cbb3d5100c0f41f8c6a9c5dcb09ab6e102eebde3d34b5b5bf59a239
|
|
| MD5 |
f8c8c7bf230371a6122722a5abd79c97
|
|
| BLAKE2b-256 |
bdc64dcbaa6c1dd08febf6b85ad93feb7fb27bdc7154261faf52efc431068d11
|
File details
Details for the file baits-0.1.1-py3-none-any.whl.
File metadata
- Download URL: baits-0.1.1-py3-none-any.whl
- Upload date:
- Size: 38.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e686e80a179321f260ba154b4f93b6d5878d8ab7bd4797f6e355687a50a8e198
|
|
| MD5 |
c108660dfca3477247fe405086d61d8c
|
|
| BLAKE2b-256 |
d7bd5a73e560f52161173dc236a5a019d55c731d5d9bfbc224fb6a8e5fcc687a
|










