This package connects large language models with bioinformatics workflows.
Project description
BRAD: Bioinformatics Digital Assistant
Please see the projects main page available here!
Architecture
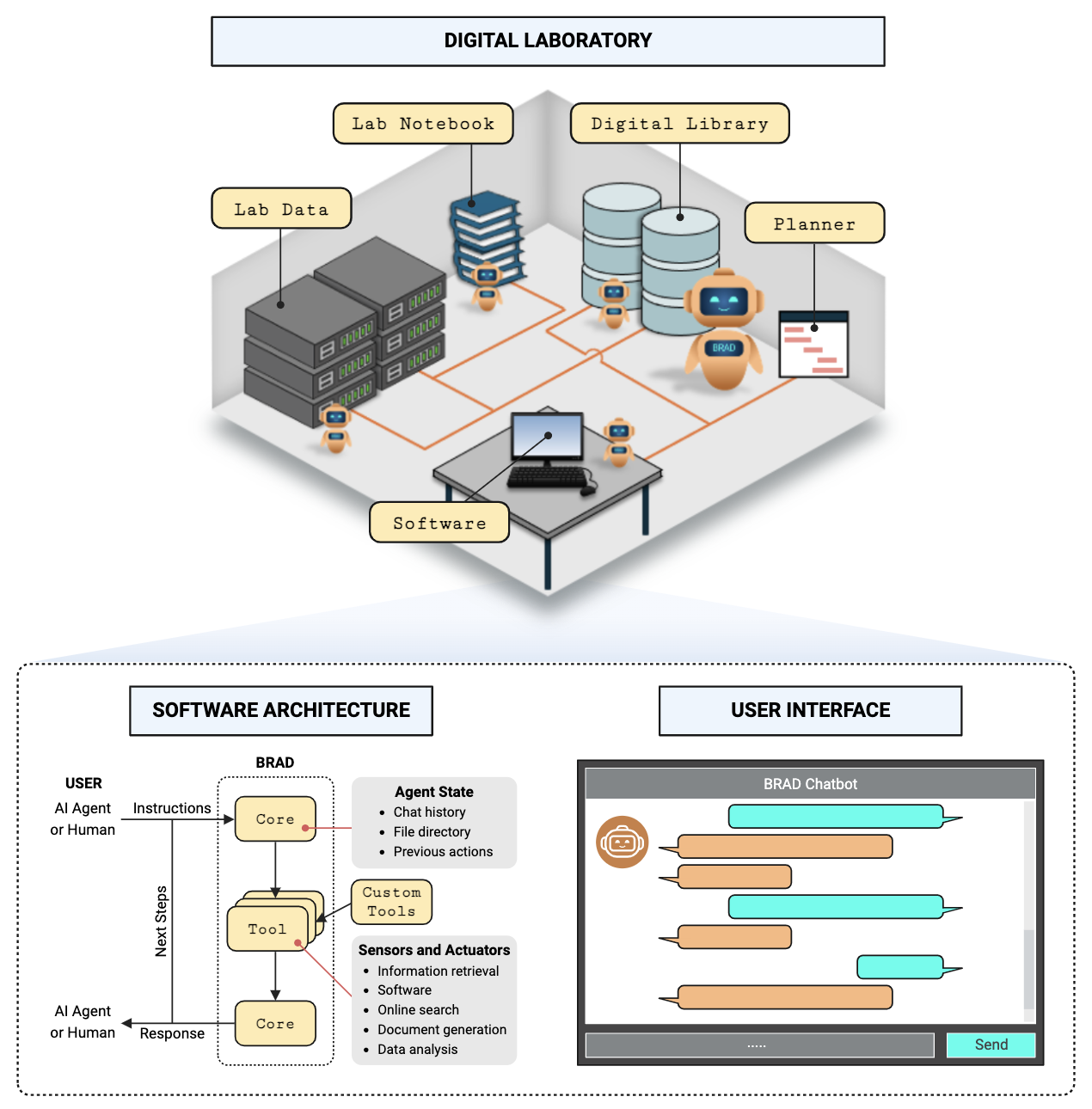
Installation
Brad can be installed directly from pip:
pip install -U BRAD-Chat
Web Server deployment
BRAD can be deployed as a web server to interact and experiment with its functionalities.
The BRAD web server is written with react.js for the frontend and flask backend.
The recommended method is to use docker.
Docker instructions
=======
BRAD-Examples
-
GUI Tutorial
A simple tutorial for how to use BRAD's Graphical User Interface. -
Hello World
A simple "Hello, World!" example to help you understand the basics of using the BRAD chatbot. -
Search and Retrieval-Augmented Generation (RAG)
Demonstrates how to use BRAD to scrape online data and integrate it into a Retrieval-Augmented Generation pipeline. -
Using the Scanpy Package with BRAD
Explores how BRAD can streamline workflows involving Scanpy, including preprocessing and visualization of single-cell data. -
Biomarker Selection Pipeline
Illustrates how BRAD can assist in selecting biomarkers from datasets using machine learning and bioinformatics tools.
Development
Docker
The Docker build can be used to deploy brad without having to install packages manually. after installing either docker desktop or docker engine docker intsallation, you can follow one of the following commands to install and run BRAD
- Use with docker compose:
cd deployment
docker compose up -d
- OR with just docker
docker run -e OPENAI_API_KEY=your_open_ai_key -e PYDANTIC_SKIP_VALIDATING_CORE_SCHEMAS='True' -p 5001:5000 -p 3000:3000 brad:full_frontend
- To build the dockerfile yourself:
docker build -t brad:local .
Then proceed to http://localhost:3000 to view the frontend
Development Environment
To contribute or make your own modifications to BRAD, A separate development environment needs to be set up. Follow the instructions below to set up your environment
Download the BRAD repository
git clone https://github.com/Jpickard1/BRAD.git
cd BRAD
Update ./BRAD/config/config.json to your preference
( specifically update the log_path key to point somewhere on your local system )
Make sure python is installed and then install the requirements
Our recommendation is to use conda and setup a separate conda environment to ensure package discrepencies
pip install -r requirements_frozen.txt
Install node=v20.18.0 and npm=10.8.2 from the (website)[https://nodejs.org/en/download/package-manager] or by using nvm
Our recommendation is to use nvm
https://raw.githubusercontent.com/creationix/nvm/master/install.sh | bash
source ~/.nvm/nvm.sh \
nvm install 20.18.0 \
nvm use 20.18.0
# install the necessary packages
npm install --prefix ./brad-chat
Start BRAD backend with
PYDANTIC_SKIP_VALIDATING_CORE_SCHEMAS=True OPENAI_API_KEY=<your_open_api_key> flask --app app run --host=0.0.0.0 --port=5000
Start BRAD frontend with
cd brad-chat
npm start
This will start BRAD Backend at - http://localhost:3000
And the BRAD frontend at - http://localhost:5000
Documentation
To build the projects documentation in the ReadTheDocs html formatting, in the docs/ directory, run the command make html. This will populate the docs/build/html directory with the webpages. The docs/build/ directory is excluded from git but will automatically be built when pushing to main.
During the build, the tutorials/ directory is automatically pulled in to the source by copying all jupyter notebooks. These are included in the user guide section of the documentation.
To remove the documentation from docs/build/ run make clean from the same directory where you built it.
Cite Us
@article{pickard2024bioinformatics,
title={Language Model Powered Digital Biology},
author={Pickard, Joshua and Choi, Marc Andrew and Oliven, Natalie and
Stansbury, Cooper and Cwycyshyn, Jillian and Galioto, Nicholas
and Gorodetsky, Alex and Velasquez, Alvaro and Rajapakse, Indika},
journal={arXiv preprint arXiv:2409.02864},
url={https://arxiv.org/abs/2409.02864},
year={2024}
}
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file brad_chat-0.0.4.tar.gz.
File metadata
- Download URL: brad_chat-0.0.4.tar.gz
- Upload date:
- Size: 250.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.11.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c23810c1c21f340e32942eebbf479f2938066bfc771507dde7c7a8e93b0b5ff0
|
|
| MD5 |
1d239815021f1e036ee7ee1b1382a038
|
|
| BLAKE2b-256 |
8adac567c45445eea9d533df38aa981c698e64b53110ff195797f6a6009c7097
|
File details
Details for the file brad_chat-0.0.4-py3-none-any.whl.
File metadata
- Download URL: brad_chat-0.0.4-py3-none-any.whl
- Upload date:
- Size: 261.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.11.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
afc3432aca9a4fce92917361cf1da9aced2b8138e615175584e9179ac45196c3
|
|
| MD5 |
5e54ecdbcb8c50085d19157e0cc51950
|
|
| BLAKE2b-256 |
1f19199be2b0707d6b5e5083bfdf9b4cf9b26a97de9b5ddbe71a6584b0edffa9
|











